Cryptochrome refSeqs: Difference between revisions
Tomemerald (talk | contribs) |
(Updated genome-test.cse links to .gi, one link broken) |
||
| (21 intermediate revisions by one other user not shown) | |||
| Line 1: | Line 1: | ||
See also: [[Cryptochrome_evolution|Comparative genomics analysis of cryptochromes and photolyases]] | See also: [[Cryptochrome_evolution|Comparative genomics analysis of cryptochromes and photolyases]] | ||
'''Updates''': fixes and additions become difficult to locate in a long article; these are provided below in reverse chronological order linked to their approximate spot in the article. | |||
20 May 12: added [[#DASH|9 new DASH sequences]], fixed various sequence discrepancies and provided a groomed sequence alignment. | |||
18 May 12: scoured [[#Cryptochromes_from_cnidarians.2C_ctenophore.2C_trichoplax.2C_sponge.2C_choanoflagellate_and_diatom|ctenophore genome]] for photolyases and cryptochromes, finding only CRY64 and CPD. | |||
10 Oct 13: intensive search of new amniote genomes for complete gene family sets | |||
== Introduction: curated metazoan cryptochrome and photolyase sequences == | == Introduction: curated metazoan cryptochrome and photolyase sequences == | ||
The | The 307 metazoan cryptochrome and photolyase reference sequences provided here are hand-curated for greater accuracy. This is necessary in part because many full length genes are implicit in genome projects but not provided as such at GenBank. When predicted by unsupervised bioinformatic algorithms, the resulting gene models too often are unacceptably inaccurate -- erroneous start codons, missing short and diverged exons, bogus splice junctions in gappy regions, uncorrected homopolymer run frameshift errors, read-through of penultimate exons to the next random stop codon, and making no use of massive and synergistic homological data. | ||
Comparative genomic analysis of a gene family cannot be built upon a rotten foundation of faulty sequences -- this can only lead to downstream rubbish. The method of curation used here is complex but necessary. It begins with a seed set of thoroughly studied experimental sequences in model species where there is no doubt about completeness of the sequence nor its accuracy. Using the four main divisions of GenBank (nr, ESTs, transcriptome projects, whole genome assemblies), the seed set of each orthology class is slowly expanded with additional representatives in closely related species that share homology, syntenic location and exon pattern. | |||
The build-out of the reference sequence collection grows recursively in accuracy because of four independent tools: an ever-growing [http://www.proweb.org/proweb/Tools/WU-blast.html blast classifier], a [http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_multalin.html phylogenetically aware] protein multi-aligner, a pre-computed best-blast [http://www.dyogen.ens.fr/genomicus-66.01/cgi-bin/search.pl phylogenetic overview] of neighboring genes, and a BROKEN [http://genome-test.gi.ucsc.edu/cgi-bin/hgPal?g=knownGene&c=chr12&l=107385143&r=107487598&i=uc001tmi.3&hgsid=52863&db=hg19 46-species] whole genome alignment based gene predictions. | |||
The blast classifier allows homologs extracted from raw dna contigs and 46-way preliminary models to be assigned to the closest available reference sequence class. The multi-aligner highlights anomalies in the sequence collection that required additional curational focus (such as incorrect exon boundaries and regions temporarily out of reading phase). The synteny browser is often correct even when its underlying gene models are somewhat flawed and is imperative to determining orthology. However synteny dissipates fairly rapidly with phylogenetic distance, so the much deeper conservation of intron position and phase becomes critical to refining gene models and orthologous classification. | |||
Despite these improvements, a set of sequences is never perfect because of errors in GenBank data (sequencing lab contamination, systemic errors in read technology, mis-assembled contigs, gaps in coverage, premature truncation of contigs, high levels of polymorphism, unsuspected hybrids, parasites and commensals, erroneous taxonomy, variations of the single animal sequenced unrepresentative of its species, lineage sorting during a messy speciation, horizontal gene transfer, and inevitable data handling errors. | |||
Contaminated DNA is a significant problem in GenBank entries. This can arise somewhat excusably from embedded viruses, endosymbionts, commensals or parasites not being cleanly separated from target organism, but less excusably from DNA contamination by species previously sequenced in the same lab (eg, DNA from the Lottia genome project contaminated sponge, the subsequent project at JGI). It is not unusual to find tuna sandwich, chicken salad and lab staff DNA in older mammalian genome projects. Horizontal gene transfer must also be considered when a sequence does not fit into its phylogenetic niche, though that is a non-issue in vertebrates. Some clades, such as tunicates and nematodes, just evolve so rapidly that their sequences seem implausible; even when confirmable, these are not informative to the comparative genomics endeavor. | |||
The | The focus here is entirely on metazoan sequences -- sponge to human -- supplemented slightly with homologs for intensively studied model species such as Arabidopsis, single-celled eukaryotes when an outgroup is helpful, and prokaryotic species where the fold has been determined. The instructive class of 4Fe-4S photolyases occurs only in prokaryotes so an exception was made there; a few primases are also included because of deep fold homology to photolyases. | ||
Because of multiple events of gene duplication, followed by accelerated asymmetric divergence and sometimes much later loss in various lineages, orthologous correspondences are not so straightforward in this gene family. Because it is not feasible to experimentally study every protein in every species, it is imperative to correctly establish orthologous families to optimize the accuracy of annotation transfer. That gives far more accurate outcomes than across paralogs -- and when it doesn't, leads to discovery of unsuspected developments in protein evolution. | |||
By definition, two genes in two species are orthologous only when they are vertically descended from a single gene '''in their last common ancestor'''. Orthology is strictly genetic -- its definition makes no reference to function but instead describes evolutionary descent. It cannot be reliably determined by reciprocal best blast. Classification by non-heritable and possibly revertible attributes such as enzyme vs signaling, presence or absence of antenna molecule, ssDNA or dsDNA substrate, peak wavelength adsorption and so forth are not productive. | |||
Orthologs can and do drift in exon number and position, cofactors, post-translational modifications, tissue and intra-cellular location of expression, regulation of that transcription, protein binding partners, enzymatic substrates and products, and relative catalytic efficiency. However, mostly they don't -- genes can persist for trillions of years of cumulative branch length time with very few changes in any attribute. Annotation transfer to orthologs provides a default hypothesis that can only be refined by actual experiment. | |||
The | The five main tools for determining orthology are primary and tertiary sequence relatedness, location and size of indels (insertions and deletions), positions and phases of introns, and retention of syntenic chromosomal position. These tools are independent of each other and so synergistic; any model of orthology relationships must be compatible with and fully utilize the information of all five. Lazy methods such as uncurated downloads, best reciprocal blast and evolution-modeling software can only provide crude provisional outcomes. | ||
Previous efforts in the comparative genomics of cryptochromes are not reviewed here. These have floundered on the twin shoals of overly sparse phylogenetic sampling relative to gene duplication event rate and over-reliance on modeling tools based on simplifying but unwarranted assumptions about clade-specific variations in the tempo and mode of nucleotide evolution over a vast evolutionary time span involved. Unpretentious blast clustering in the classifier accomplishes the same thing with far greater speed and transparency. | |||
Since today the phylogenetic tree is largely [http://www.sciencemag.org/content/334/6059/1091.full geologically dated and topologically fixed], individual gene trees -- which are commonly discrepant -- can be clamped (subordinated) to the species tree even if that is less parsimonious. After a gene duplication, if one copy retains parental location and main role, the other copy may re-functionalize and evolve rapidly as it adapts better to its newly specialized role. This scenario can account for anomalies in gene trees and is likely applicable here to the shift from repair enzyme to light sensor. | |||
In the view of [http://en.wikipedia.org/wiki/Protein_moonlighting Piatigorsky], parental genes are usually multi-functional to begin with, so the duplicated copy already has a pre-existing niche available to take over -- it does not need to scramble desperately for a function before pseudogenization from lack of selective pressure support. All fixed gene duplications of cryptochromes within Bilateria have been segmental, meaning upstream regulatory and enhancer regions were likely carried along, which is not the case with retropositioned gene copies which duplicate only a matured mRNA transcript and not its promoter regions. | |||
The | The sequence collection has been split from a [[Cryptochrome_evolution|companion analysis page]] to separate data from analysis. Not every conceivable cryptochrome gene fragment at GenBank is represented below; the intensity of curational effort increases in proportion to that needed to resolve specific evolutionary issues. For example, vertebrate CRY1 genes evolve very slowly in their main region but acquired additional terminal exons; the issue there is where those originated, what parts of them are conserved, and what they are doing. Here a terminal gene fragment from a phylogenetically critical position is more informative than infilling static regions of the tree with intermediate species. | ||
The sequences are provided in [[Opsin_evolution:_ancestral_introns#Intron_location_and_phase_for_dummies|advanced fasta format]] -- parsed into exons, preceded by splice acceptor overhang, followed by splice donor coding phase overhang, with lower case amino acids indicating uncertainty, spaces indicating gaps in coverage, and stop codons shown when explicitly in the data. Frameshifts in homopolymer runs -- a common form of sequencing error -- have been edited to restore reading frame. | |||
The sequences are grouped as orthologs, re-named as necessary to avoid confusion with historical use, and presented in phylogenetic order relative to human. This does not determine order in subtrees (eg rodents) but there the order can be set by assembly quality (eg mouse is the best murid assembly. | |||
The fasta header lines are simple space-delimited databases showing first gene name, then genus, species, common name, accession number if not a simple genomic blat or whole genome alignment output, PubMed accession if specifically studied in a journal article, followed by an unstructured comment field. The headers and exons are easily reformatted into a spreadsheet single lines by replacing spaces and paragraph returns with tabs. | |||
Here gene names are designed for consistency across all Eukaryota, with special attention to phylogenetically localized gene duplications. Following [http://www.genenames.org/guidelines.html international agreement], no greek letters, roman numerals, hyphenation, lower case letters, subscripts, or superscripts can be used in gene names. It is completely unworkable with 10,000 vertebrate genomes to use narrowly conceived lab conventions such as mCRY1 for mouse -- even 6 letters for genusSpecies can provide no more than a mnemonic. Since annotation today is done by gene family, the gene name comes first to preserve orthologous clustering when listed alphabetically, which also provides clarity in alignment tool output. | |||
The | The availability of some orthology classes is quite limited due to recent origin and restricted phylogenetic persistence. Sequencing effort is unfortunately spread very unevenly across the phylogenetic tree of metazoans. For species with good assemblies, the entire repertoire of cryptochromes and photolyases can be extracted. For large genes with numerous exons, absence from the assembly usually means genuine absence from the genome. When only an exon or two gene fragment is available, the classifier can still usually assign the correct orthology class. However it is risky to assemble an entire gene from small unlinked contigs and that was not done here with the exception of critical clades such as cartilaginous fish that lack coherent assemblies and comprehensive transcript collections. It sometimes suffices just to establish the presence/absence of a given gene at a given time of divergence. | ||
A remarkable amount of the data below surfaced at GenBank during the last six months, implying much better phylogenetic coverage will surface during 2012-14. This promises to resolve the timing of gene duplications to the extent that extant species can determine it. The mysterious role of 4Fe-4S clusters in photolyases, primases, helicases and other DNA repair enzymes may also be clarified experimentally, providing a better foundation to understanding the deeper evolutionary origin of photolyases and cryptochromes. | |||
In terms of massive gene losses in this family supporting the [http://rspb.royalsocietypublishing.org/content/280/1765/20130508 recently revisited] 1942 Wall hypothesis concerning a deep nocternal epoch for mammals, the record is still somewhat spotty in the critical region of early tetrapods. Amphibians other than frogs remain very poorly represented at GenBank. Only four photolyases/cryptochromes can be confirmed as of Oct 13, all transcripts from a single species of salamader. Only only one of these -- DASH -- is full length. | |||
Thus it remains difficult to determine the full crytochrome/photolyase repertoire at the various amniote ancestral nodes. The Wall hypothesis has narrower versions, such as predicting losses in say, coelocanths, which live year-round in an extremely dark deep-sea environment and so might also be expected to have lost most genes in this family. Recently the genome of a cave fish has become available -- it too will likely have lost its crytochrome/photolyases as well as most opsins (to the extent these have no light-independent functions), depending on how many millions of years it has been out of genetic contact with surface fish. | |||
An exonic pseudogene must be fairly new (several million years) to still be recognizable so indicates not only contemporary gene loss but that the species possessed an operational gene in the recent past. Actually, the same is true for a processed (intronless) pseudogene because it implies the existence (possibly continuing but just not in the current assembly) of a parent gene that gave rise to it. The difference is that processed pseudp\ogenes are far more persistent in terms of Blast dectectability because the longer length of their contiguous (formerly) coding region. | |||
For example, DASH consists of 524 amino acids located in 14 widely separated exons with lengths varying from 20 to 59 residues (average of 37). As frameshifts from indels and substitutions from nucleotide changes accrue over time, these exons become increasingly difficult for Blast to confidently identify, a situation exacerbated by the best probe itself (necessarily from another species, potentially only ~85% identical) and potential mis-attribution to other members of the gene family and their pseudogenes. | |||
=== Vertebrate CRY1 reference sequences === | === Vertebrate CRY1 reference sequences === | ||
| Line 241: | Line 255: | ||
2 GRSSMGTSLSSGKRPSQEEETRSVDPKVQRQSSN* 0 | 2 GRSSMGTSLSSGKRPSQEEETRSVDPKVQRQSSN* 0 | ||
>CRY1_spaJud Spalax judaei ( | >CRY1_spaJud Spalax judaei (blind_molerat) AJ606298 | ||
0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRW 2 | 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRW 2 | ||
1 RFLLQCLEDLDANLRKLNSRLLVIRGQPADVFPRLFKEWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 | 1 RFLLQCLEDLDANLRKLNSRLLVIRGQPADVFPRLFKEWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 | ||
| Line 268: | Line 282: | ||
2 GRSSMGTGLSSGKRPSQEEDSQSIGPKVQRQSTN* 0 | 2 GRSSMGTGLSSGKRPSQEEDSQSIGPKVQRQSTN* 0 | ||
>CRY1_hetGla Heterocephalus glaber ( | >CRY1_hetGla Heterocephalus glaber (naked_molerat) AHKG01030319 revised hystricognath | ||
0 | 0 MAVNAVHWFRKGLRLHDNPALKECTRGADTIRCVYILDPWFAGSSNVGINRWR 2 | ||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | ||
0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 | 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 | ||
| Line 278: | Line 292: | ||
0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 | 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 | ||
1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 | 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 | ||
2 | 2 GLLASVPSNPNGNGGLMGYAPGESIPGGSSSG 1 | ||
2 SCAHGSGILPCAHTDGQQAHLLKP 1 | 2 SCAHGSGILPCAHTDGQQAHLLKP 1 | ||
2 GRNCVGPVLSSGKRPSQEEDAQSIGPKLQRQSTD* 0 | 2 GRNCVGPVLSSGKRPSQEEDAQSIGPKLQRQSTD* 0 | ||
>CRY1_cavPor Cavia porcellus ( | >CRY1_cavPor Cavia porcellus (guinea_pig) AAKN02054126 last two exons diverged, only 69 bp separation | ||
0 MGVNAVHWFRKGLRLHDNPALKECIRGADTIRCVYILDPWFAGSSNVGINRWR 2 | 0 MGVNAVHWFRKGLRLHDNPALKECIRGADTIRCVYILDPWFAGSSNVGINRWR 2 | ||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | ||
| Line 296: | Line 310: | ||
2 GRDPAGPGLGGGKRPSQEEDAQSTGHKIQRQSPD* 0 | 2 GRDPAGPGLGGGKRPSQEEDAQSTGHKIQRQSPD* 0 | ||
>CRY1_speTri Spermophilus tridecemlineatus (squirrel) Ictidomys | >CRY1_octDeg Octodon degus (degu) AJSA01012260 AJSA01012259 hystricognath | ||
0 | 0 MGVNAVHWFRKGLRLHDNPALKECIRGADTIRCVYILDPWFAGSSNVGINRWR 2 | ||
1 | 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | ||
0 | 0 EWNITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 | ||
1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCMTPLSDDHDEKYGVPSLEEL 1 | |||
2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 | |||
0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 | |||
0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWEEGMK 0 | |||
0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 | |||
1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQLYQQLSRYRGL 1 | |||
2 GLLASVPSNPNGNGGLLGYAAGETLPGGGGGG 1 | |||
2 SCTPGSSILHCAHQDGQQAPLLKP 1 | |||
2 GRSSGGPAPGSGKRPSQEEDTQSIGHKVQRQSTT* 0 | |||
>CRY1_speTri Spermophilus tridecemlineatus (squirrel) AGTP01024540 AGTP01024543 Ictidomys | |||
0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 | |||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | |||
0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 | |||
1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 | 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 | ||
2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 | 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 | ||
| Line 625: | Line 653: | ||
2 GRTSLSSGISAGKRPNPEEETQSIGPKVQRQSTN* 0 | 2 GRTSLSSGISAGKRPNPEEETQSIGPKVQRQSTN* 0 | ||
> | >CRY1_croAda Crotalus adamanteus (rattlesnake) JU174244 TSA | ||
0 | 0 MGVNSVHWFRKGLRLHDNPALREVIEGADTVRCVYILDPWFAGSSNVGINRWR 2 | ||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | ||
0 | 0 EWNISKLSIEYDSEPFGKERDAAIKKLATEAGLEVIVRISHTLYDLDK 2 | ||
1 | 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMSTCITPVSDDHDEKYGVPSLEEL 1 | ||
2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 | 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 | ||
0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 | 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 | ||
0 | 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGVK 0 | ||
0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 | 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 | ||
1 | 1 RYLPILRGFPAKYIYDPWNAPEGVQKAAKCTIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 | ||
2 | 2 GLLASVPSNGNGNGNGGLMGYSPSENLPGCVGTN 1 | ||
2 | 2 GTQMGTSEGHTGNVQACTLGETHTGTSGIHQQ 1 | ||
2 | 2 GYCQGNSGILHYAHGDSKKTHLMQQ 1 | ||
2 | 2 GRTSLSIGGCTGKRPNPEEGIQSIGPKVQRQSSN* 0 | ||
>CRY1_pytMol Python molurus (python) AEQU010547455 | |||
0 MGVNSVHWFRKGLRLHDNPALREVIQGADTVRCVYILDPWFAGSSNVGINRWR 2 | |||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFk 0 | |||
0 EWNISKLSIEYDSEPFGKERDAAIKKLATEAGLEVIVRISHTLYDLDK 2 | |||
>CRY1_pytMol Python molurus (python) AEQU010547455 | |||
0 MGVNSVHWFRKGLRLHDNPALREVIQGADTVRCVYILDPWFAGSSNVGINRWR 2 | |||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFk 0 | |||
0 EWNISKLSIEYDSEPFGKERDAAIKKLATEAGLEVIVRISHTLYDLDK 2 | |||
1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMSKCTTPVSDDHDEKYGVPSLEEL 1 | 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMSKCTTPVSDDHDEKYGVPSLEEL 1 | ||
2 GFDTDGLPSAVWPGGETEALTRLERHLER 0 | 2 GFDTDGLPSAVWPGGETEALTRLERHLER 0 | ||
| Line 685: | Line 698: | ||
2 GRTVSTGISTGKRPNPEKETQSIGPKVQRQSTN* 0 | 2 GRTVSTGISTGKRPNPEKETQSIGPKVQRQSTN* 0 | ||
> | >CRY1_anoCar Anolis carolinensis (lizard) XM_003220923 AAWZ02014443 | ||
0 | 0 MGVNAVHWFRKGLRLHDNPALREVIQGADTVRCVYILDPWFAGSSNVGINRWR 2 | ||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | ||
0 EWKITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 | 0 EWNITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYELDK 2 | ||
1 IIELNGGQPPLTYKRFQTLISKMDPLEIPVETITAEVMEKCTTPVSDDHDEKYGVPSLEEL 1 | 1 IIELNGGQPPLTYKRFQTLISRMEPLEIPVETITAEVMSKCTTPVSDDHDEKYGVPSLEEL 1 | ||
2 GFDTEGLPSAVWPGGETEALTRLERHLERK 0 | 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 | ||
0 AWVANFERPRMNANSLLASTTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 | 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 | ||
0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMDGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 | 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 | ||
0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGKRTDPNGDYIR 2 | 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 | ||
1 RYLPILKGFPPKYIYDPWNAPETVQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 | 1 RYLPVLRGFPAKYIYDPWNAPDGVQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQMYQQLSRYRGL 1 | ||
2 GLLASVPSNPNGNGNGGLMSYSPGESMSGCSNNG 1 | 2 GLLASVPSNGNGNGNGGLMGYSTGENIPGCTNTN 1 | ||
2 GGQMGVNEGSSASNPNANKGEVHPGTSGLQ 1 | 2 GSQMGMNEGHIGNVQACTMGESHTGTSGIQQQ 1 | ||
2 GYSQGSGILLYSHGDNQKTHSAQKGRISLGT 1 | |||
2 GVCTGKRPSPEVETQSVGPKVQRQSSN* 0 | |||
>CRY1_podSic Podarcis siculus (wall_lizard) DQ376040 16809482 | |||
0 MGVNAVHWVRQGLRLPDNRALREVIQGADTARCVYILDPSFAGSSNVGINRWR 2 | |||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | |||
0 EWNITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 | |||
1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMSKCTTPVSDDHDEKCGVPSLEEL 1 | |||
2 GFDTGGLPSAVWPGGETEALTRLERHLERK 0 | |||
0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 | |||
0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARRAVACFLTRGDLWISWEEGMK 0 | |||
0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 | |||
1 RYLPVLRGFPAKYIYDPWNAPEGVQKVAKCVIGVNYPKPMVNHTEASRLNIERMKQIYQQLSRYRGL 1 | |||
2 GLLASVPLMEWNVIVLMGYHQGTLLVYHAS 1 | |||
2 GSQMGTNEAHTGSVQTCTLGESHTGTSGIQQQ 1 | |||
2 GYPQGSDILHYAHGEGQKTHLIQQGRASLVA 1 | |||
2 GVCTGKRPNPEEETQSIGPKVQRQSSK* 0 | |||
>CRY1_xenTro Xenopus tropicalis (frog) NM_001087660 11533577 final four exons confirmed by many ESTs | |||
0 MGVNAVHWFRKGLRLHDNPALRECIQGADTVRCVYILDPWFAGSSNVGINRWR 2 | |||
1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 | |||
0 EWKITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 | |||
1 IIELNGGQPPLTYKRFQTLISKMDPLEIPVETITAEVMEKCTTPVSDDHDEKYGVPSLEEL 1 | |||
2 GFDTEGLPSAVWPGGETEALTRLERHLERK 0 | |||
0 AWVANFERPRMNANSLLASTTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 | |||
0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMDGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 | |||
0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGKRTDPNGDYIR 2 | |||
1 RYLPILKGFPPKYIYDPWNAPETVQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 | |||
2 GLLASVPSNPNGNGNGGLMSYSPGESMSGCSNNG 1 | |||
2 GGQMGVNEGSSASNPNANKGEVHPGTSGLQ 1 | |||
2 GYWQGSSILHYSHSDSQQSYLMQ 1 | 2 GYWQGSSILHYSHSDSQQSYLMQ 1 | ||
2 ARNPLHSVVSSGKRPNPEEETQSIGPKVQRQSSH* 0 | 2 ARNPLHSVVSSGKRPNPEEETQSIGPKVQRQSSH* 0 | ||
>CRY1_ambMex Ambystoma mexicanum (axolotl) JV201332 fragment 154-312 | |||
QTLISRMEPLEMPVETIGPEVMEKCTTPVLDDHDDKYGVPSLEELGFDTEGLPSAVWPGGE | |||
TEALTRLERHLERKAWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK | |||
VKKNSLPPLSLYGQLLWREFFYTAATNNPRFDKMEGN | |||
>CRY1A_latCha Latimeria chalumnae (coelocanth) AFYH01018055 AFYH01018053 AFYH01018050 | >CRY1A_latCha Latimeria chalumnae (coelocanth) AFYH01018055 AFYH01018053 AFYH01018050 | ||
| Line 928: | Line 976: | ||
1 RYLPKLKAFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 | 1 RYLPKLKAFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 | ||
2 CLLASVPSCVEDLSHPVAEPSSSQAGSMSSA 1 | 2 CLLASVPSCVEDLSHPVAEPSSSQAGSMSSA 1 | ||
2 | 2 GPRPLPSGPAS<font color=magenta>PKRK</font>LEAAEEPPGEELSKRARVAELPTPELPSKDA* 0 | ||
>CRY2_panTro Pan troglodytes (chimp) | >CRY2_panTro Pan troglodytes (chimp) | ||
| Line 1,060: | Line 1,108: | ||
2 GSRPLSSGPASPKRKLEAAEEPPGEELSKRARVTEMPAQEPQTKDA* 0 | 2 GSRPLSSGPASPKRKLEAAEEPPGEELSKRARVTEMPAQEPQTKDA* 0 | ||
>CRY2_spaJud Spalax judaei ( | >CRY2_spaJud Spalax judaei (blind_molerat) AJ606300 | ||
0 MAAASVVVATSAAPAMAVDGGSSVHWFRKGLRLHDNPSLLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 | 0 MAAASVVVATSAAPAMAVDGGSSVHWFRKGLRLHDNPSLLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 | ||
1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 | 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 | ||
| Line 1,085: | Line 1,133: | ||
2 CLLASVPSCVEDLSHPVAEPSSSQAGSISST 1 | 2 CLLASVPSCVEDLSHPVAEPSSSQAGSISST 1 | ||
2 GPRPLSSGPASPKRKLEAAEEPPGEELSKRARVTEMPAQEPPSKDS* 0 | 2 GPRPLSSGPASPKRKLEAAEEPPGEELSKRARVTEMPAQEPPSKDS* 0 | ||
> | >CRY2_hetGla Heterocephalus glaber (naked_molerat) EHA99865 hystricognath | ||
0 | 0 MAAAVGTGTGAAPTPATGAEGACSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 | ||
1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 | 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 | ||
0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 | 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 | ||
1 | 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGAVSSQQMKSCRAEIQENHDDTYGVPSLEEL 1 | ||
2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 | 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 | ||
0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 | 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 | ||
0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 | 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 | ||
0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 | 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 | ||
1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 | 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 | ||
2 CLLASVPSCVEDLSHPVAEPTLSQAGSSSST 1 | |||
2 GPRSLPDGPASPKRKLEAAEEPPGEELSKRARVAELPAPEPTSKDA* 0 | |||
>CRY2_cavPor Cavia porcellus (guinea_pig) hystricognath | |||
0 MAAAVGTGTAAAPTPVTGAEGACSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 | |||
1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 | |||
0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 | |||
1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGAVSSQQMESCRAEIQENHDDTYGVPSLEEL 1 | |||
2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 | |||
0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 | |||
0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 | |||
0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 | |||
1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 | |||
2 CLLASVPSCVEDLSHPVAEPSLSQAGSSTST 1 | 2 CLLASVPSCVEDLSHPVAEPSLSQAGSSTST 1 | ||
2 GPRPLPGGPASPKRKLEAAEEPPGEELSKRARVAELPTPEPPSKDA* 0 | 2 GPRPLPGGPASPKRKLEAAEEPPGEELSKRARVAELPTPEPPSKDA* 0 | ||
> | >CRY2_octDeg Octodon degus (degu) AJSA01043118 hystricognath | ||
0 | 0 MAAAVGTGTSAAPTPAAGTEGACSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 | ||
1 | 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPAEVFPRLFK 0 | ||
0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 | 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 | ||
1 | 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGVVSSQQMEGCRAEIPENHDDTYGVPSLEEL 1 | ||
2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 | 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 | ||
0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 | 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 | ||
0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 | 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 | ||
0 | 0 VFDELLLDADFSVNSGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 | ||
1 | 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIVGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 | ||
2 | 2 CLLASVPSCVEDLSHPVAEPSSSQAGSSTST 1 | ||
2 | 2 GPRPLPSGSASPKRKLEAAEEPPGDELSKRARVAELPTPEPSSKDA* 0 | ||
>CRY2_speTri Spermophilus tridecemlineatus (squirrel) | >CRY2_speTri Spermophilus tridecemlineatus (squirrel) | ||
| Line 1,371: | Line 1,432: | ||
2 GLLASVPSCAEDLSGPVTDPASVQGCSTST 1 | 2 GLLASVPSCAEDLSGPVTDPASVQGCSTST 1 | ||
2 ALKPSQSGQASPKRKHEGIEEMCTEDLYKRAKVTGLHGPEIPSKSL* 0 | 2 ALKPSQSGQASPKRKHEGIEEMCTEDLYKRAKVTGLHGPEIPSKSL* 0 | ||
>CRY2_croAda Crotalus adamanteus (rattlesnake) JU174243 TSA | |||
0 MAALPGPVPRPRSGGGCCSVHWFRRGLRLHDNPALQAAIREATSVRCIYILDPWFAASSAVGINRWR 2 | |||
1 FLLQSLEDLDNSLRKLNSRLFVVRGQPTDVFPRLFK 0 | |||
0 EWGVTHLTFEYDSEPFGEERDAAIVKLAKEAGVKVTTENSHTLYDLDR 2 | |||
1 IIELNGHKPPLTYKRFQAIISRMDLPKKPVSTITSQQMEMCQTKIQENHDETYGVPSLEEL 1 | |||
2 GFFTEGLAPAVWQGGETEALTRLDKHLERK 0 | |||
0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 | |||
0 VKRNSTPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAAACFLTRGDLWISWESGVR 0 | |||
0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 | |||
1 RYLPQLKGFPSRYIYEPWNASESVQKAAKCIIGVDYPKPMVNHAETSRLNIERMKQIYQQLSRYRGL 1 | |||
2 CLLASVPSCIEDFSNITTDLSSGSCSNTTL 1 | |||
2 RQSHCSQTSPKRKCDAEEDNNEELHKRVRMMVHQVSKKPQQLSQPEFGN* 0 | |||
>CRY2_anoCar Anolis carolinensis (lizard) XM_003214641 | >CRY2_anoCar Anolis carolinensis (lizard) XM_003214641 | ||
| Line 1,385: | Line 1,459: | ||
2 ASRQTHCGQTSPKRKHDVVQEYKEELYKRAKVVASQFAENPRQELVHHEMGN* 0 | 2 ASRQTHCGQTSPKRKHDVVQEYKEELYKRAKVVASQFAENPRQELVHHEMGN* 0 | ||
>CRY2_xenTro Xenopus tropicalis (frog) NM_001088670 AY049035 CX389867 11533577 discrepancies | >CRY2_xenTro Xenopus tropicalis (frog) NM_001088670 AY049035 CX389867 CX841030 11533577 discrepancies | ||
0 MEGKPSVSSVHWFRKGLRLHDNPALLSALRGANSVRCVYILDPWFAASSSGGVNRWR 2 | 0 MEGKPSVSSVHWFRKGLRLHDNPALLSALRGANSVRCVYILDPWFAASSSGGVNRWR 2 | ||
1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 | 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 | ||
| Line 1,395: | Line 1,469: | ||
0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVR 2 | 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVR 2 | ||
1 RYLPVLKAFPSRYIYEPWSAPESVQKEAKCIIGIDYPKPIVNHAEASRMNIERMKQTYQQLSRYRGL 1 | 1 RYLPVLKAFPSRYIYEPWSAPESVQKEAKCIIGIDYPKPIVNHAEASRMNIERMKQTYQQLSRYRGL 1 | ||
2 | 2 CILASVPSCMEDLGGPMADSSQNISEA 1 | ||
2 | 2 APKQSLCNVESPKRKRKVSEEEPHMKVRVLCAPETERPAKDF* 0 | ||
>CRY2_ranCat Rana catesbeiana (bullfrog) GO458565 AY256684 extra SS removed | >CRY2_ranCat Rana catesbeiana (bullfrog) GO458565 AY256684 extra SS removed | ||
| Line 1,410: | Line 1,484: | ||
2 CILASVPSSVEDLSGPMADPASSQQGSS 1 | 2 CILASVPSSVEDLSGPMADPASSQQGSS 1 | ||
2 DPAPRLCAVDSPKRKHEDLDGELCKKARLQCVQEMERAAKDFL* 0 | 2 DPAPRLCAVDSPKRKHEDLDGELCKKARLQCVQEMERAAKDFL* 0 | ||
>CRY2_latCha Latimeria chalumnae (coelocanth) AFYH01005158 AFYH01005161 AFYH01005164 | |||
0 MVVNSVHWFRKGLRLHDNPALQEALKGADTVRCVYMLDPRFAGSSNLGINRWR 2 | |||
1 0 | |||
0 EWNVTRLTFEYDSEPFGKERDAVIIKVARESGVEVIMRDSHTLYNLNR 2 | |||
1 IIELNGHKPPLTYKRFQAIISRMELPRRPVAAVTQQQMETCKTDIGNDHDEQYGVPSLAEL 1 | |||
2 GFQVENDPAIWQGGETEALARLEKHLDRK 0 | |||
0 AQVANFERPWINANSLLANPTSLSPYLRFGCLSCRLFYYRLLELYKK 0 | |||
0 IKGNSSPPLSLYGQLLWREFFYTAATNNPRFDRMEGNPICVQIPWDSNPKALAKWAEGQTGFPWIDAIMTQLRQEGWIHHLARHSVACFLTRGDLWISWECGMK 0 | |||
0 VFEELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDFVR 2 | |||
1 1 | |||
2 1 | |||
2 * 0 | |||
>CRY2_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01038797 | >CRY2_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01038797 | ||
| Line 1,423: | Line 1,510: | ||
2 GLLASVPSVPDDARSPVTDDPHGHSS 1 | 2 GLLASVPSVPDDARSPVTDDPHGHSS 1 | ||
2* 0 | 2* 0 | ||
>CRY2_danRer Danio rerio (zebrafish) aka CRY3 NM_131786 | >CRY2_danRer Danio rerio (zebrafish) aka CRY3 NM_131786 | ||
| Line 1,507: | Line 1,581: | ||
=== Invertebrate cryptochromes === | === Invertebrate cryptochromes === | ||
>CRY1B_strPur Strongylocentrotus purpuratus (sea_urchin) XM_001183029 echinoderm lacks final 2 exons | >CRY1B_strPur Strongylocentrotus purpuratus (sea_urchin) XM_001183029 echinoderm lacks final 2 exons | ||
0 MPGGACIHWFRHGLRLHDNPALLEGMTLGKEFYPVFIFDNEVA 1 | 0 MPGGACIHWFRHGLRLHDNPALLEGMTLGKEFYPVFIFDNEVA 1 | ||
| Line 2,053: | Line 2,128: | ||
2 * 0 | 2 * 0 | ||
=== Cryptochrome | === Cryptochrome 6-4 photolyases === | ||
>CRY64_anoCar Anolis carolinensis (lizard) XM_003225714 6-4 photolyase synteny: DCPS TIRAP CRY64 SRPR FOXRED1 | >CRY64_anoCar Anolis carolinensis (lizard) XM_003225714 6-4 photolyase synteny: DCPS TIRAP CRY64 SRPR FOXRED1 | ||
| Line 2,064: | Line 2,139: | ||
0 GQTGYPFIDAIMTQLRTEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 | 0 GQTGYPFIDAIMTQLRTEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 | ||
0 VFEELLLDADWSLNAANWQWLSASAFFHQFFRVYSPVTFGKKTDKNGEYIK 2 | 0 VFEELLLDADWSLNAANWQWLSASAFFHQFFRVYSPVTFGKKTDKNGEYIK 2 | ||
1 | 1 KYLPFLRKFSNDYIYEPWKAPRSLQERAGCIIGQDYPKPIVEHEKVYKRNLERMKAAYARRSPNLVIQAKDKVSQKK 1 | ||
2 | 2 GVNRKRPEAPTKAKVQAKKVKTKSS* 0 | ||
>CRY64_chrPic Chrysemys picta (turtle) AHGY01135270 AHGY01135271 no synteny | >CRY64_chrPic Chrysemys picta (turtle) AHGY01135270 AHGY01135271 no synteny | ||
| Line 2,114: | Line 2,189: | ||
1 KYLPILKKFPAEYIYEPWKAPRSLQERAGCIIGKDYPKPIVEHDVASKQNIQRMKAAYARRSGSTAEVDKDSGQSNKN 1 | 1 KYLPILKKFPAEYIYEPWKAPRSLQERAGCIIGKDYPKPIVEHDVASKQNIQRMKAAYARRSGSTAEVDKDSGQSNKN 1 | ||
2 GAKRKVAGGPSVAELFKKNKSKKD* 0 | 2 GAKRKVAGGPSVAELFKKNKSKKD* 0 | ||
>CRY64_ambMex Ambystoma mexicanum (axolotl) CO788545 fragment 224-370 | |||
TVRNPGMFEMHMKRTAWVCNFKKPDTEPNSLTASTTVLSPYVKFGCMSVRTFWWRLAEIY | |||
QGKKHSDPPVSLHGQLLWREFFYTTGAGIPNFNKMEGNSVCVQVDWDNNQENLRAWSEGR | |||
TGYPFIDAIMTQLRTRRLDPPLGASCCGVLPYERIHHLARHAVACFLTRGDLWKL | |||
>CRY64_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01024141 AHAT01024142 | >CRY64_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01024141 AHAT01024142 | ||
| Line 2,371: | Line 2,451: | ||
2 ILGQNYPHRIVLDLEERREQSLKDVVEVRKKHLEYLDEVSGSDMIPIPDQLLALTLGRPNGDDEVVRNRTSSFLLPVITRKEFKYKTLQPEAKDNPYSTVLKGYV | 2 ILGQNYPHRIVLDLEERREQSLKDVVEVRKKHLEYLDEVSGSDMIPIPDQLLALTLGRPNGDDEVVRNRTSSFLLPVITRKEFKYKTLQPEAKDNPYSTVLKGYV | ||
GVSRKRDETIAYMNEKHFTASTIHEGAQRHERTERTSRLMEGLPAPSEPKNKSRRTPRKDPFSIIPPSYLHLAN* 0 | GVSRKRDETIAYMNEKHFTASTIHEGAQRHERTERTSRLMEGLPAPSEPKNKSRRTPRKDPFSIIPPSYLHLAN* 0 | ||
>CRY7_ranCla Rana clamitans (green frog) GAEG01010186 GAEG01009609 fragments 91% frog | |||
DLLSETLKKHGVTFTRYHSYCLYEPYSVSTEGVGLRGIGSVSHFMGCCERNSSSPIGDPL | |||
EAPTSLPLPSCWPDSLELDRLDLAKMPRRKDGTVIDWAATIKKSWDFSEDGAYICLGNFL | |||
EDGIKHYDKESGRADKPYTSHISPYLHFGQISPRTVLHEAYFTKK | |||
SVPKFLRKLAWRDLSYWLLVLFPDMHVEPVRPAYKSQRWSSDRSHLRAWQKGMTGYPL | |||
VDAAMRELWLTGWMCNYSRHVVASFLVAYLHLHWIHGYRWFQDTLVDADVAINAMM | |||
wQNGGMSGLDHWNFVMHPVDAAMTCDPYGSYVRKWCPELAGLPDEYIHKPWKCPPSQLRRA | |||
GIVLGQNYPHRIVVDLEERREQSLRDVVEVRQKHSEYVDKVSGSDMVSVPDQLLAFTLGC | |||
ADGDDDVIKSNTCQFLLPVITRKEFKYKTLQPDTKDNPYNTVLKGYVSRKRDETIAYM | |||
NERHFTASTINEGAQLYERRERTARIMEGLPQTRDPKNKSRRTPTSDPFSIVPPAYFTI | |||
>CRY7_pseReg Pseudacris regilla (Pacific treefrog) GAEI01034610 fragment | |||
WSNDKNNLRAWQKGMTGYPLVDAAMRELWLTGWMCNYSRHIVASFLVAYLHLHWIHGYRW | |||
FQDTLVDADVAINAMMWQNGGMSGLDHWNFVMHPVDAALTCDPDGSYVKKWCPELAGLPD | |||
EYIHKPWKCAPSQLRRAGM | |||
>CRY7_latCha Latimeria chalumnae (coelocanth) AFYH01265207 pseudogene | >CRY7_latCha Latimeria chalumnae (coelocanth) AFYH01265207 pseudogene | ||
| Line 2,523: | Line 2,619: | ||
EIAALPNDFIHQPWKCPPSILRRCGIKLGETYPYRVISDLEGAREQSLTDVVNVRKKHPEFVDRRTGNDLMPLPDGLCVPVITRKEFKYKLHHPEAKDNPHTAVLRGYRSRKRDEAIAFANERDFMASATNESVKLTERRLKATQYEAL* 0 | EIAALPNDFIHQPWKCPPSILRRCGIKLGETYPYRVISDLEGAREQSLTDVVNVRKKHPEFVDRRTGNDLMPLPDGLCVPVITRKEFKYKLHHPEAKDNPHTAVLRGYRSRKRDEAIAFANERDFMASATNESVKLTERRLKATQYEAL* 0 | ||
=== DASH cryptochrome | === DASH cryptochrome sequences === | ||
> | >DASH_galGal Gallus gallus (chicken) pseudogene in syntenic position with numerous indels, frameshifts, internal stops | ||
0 | 0 0 | ||
0 | 0 2 | ||
1 | 1 MVVRKGKLEDMVHDVITSLGCVGVVAFHEE 0 | ||
0 | 0 2 | ||
1 | 1 IQPDVYTNF*KAVESEVKTWPALQRADQLKPLAGG.EEGCIPTTEDLGQK 1 | ||
2 | 2 DPGTDPQTAF-LGFNETAAL 0 | ||
0 | 0 LLFASYNETWSGLIGVD*STKFAPW 2 | ||
1 | 1 LVLGCLSP*YIFSQIGKYVKERAANHST*W 2 | ||
1 | 1 AVFELLWWDCF*FVALKYGRRILSLK 1 | ||
2 | 2 0 | ||
0 | 0 LGLDWRMGAEWFEYLL 0 | ||
0 | 0 VDYNVFSN*STWLYSAGIGNHPKDNRKFNVIKQGPDYDGN 0 | ||
0 | 0 GDCIQLRVTELQGMKEADIHSPWALNSATLSQVGVTLGKTYPQVQSDSP^EWSRQIRQRA 0 | ||
0 | 0 GRSPHPKGRRGPAHSPLQHKERGTDCFSLKK* 0 | ||
> | >DASH_melGal Meleagris gallopavo (turkey) ADDD01036185 pseudogene in syntenic position | ||
0 0 | 0 0 | ||
0 | 0 2 | ||
1 TLVVRKGKPEDVVRDLITQLGCVGVVAFHEE 0 | 1 0 | ||
0 2 | |||
1 DVYTNL*KAVESEVKVWPALQRADQLKPLA-GSGRKMYPYNKGFG 1 | |||
2 0 | |||
0 LFASYKEIWSGLTGADNSTKFAPW 2 | |||
1 LVLDCLSP*CIFSQVQKYIKERAANQSTFW 2 | |||
1 VFELLWWDCF*FVPLKYGRRIFSL 1 | |||
2 0 | |||
0 LKVDYNVPNN*VT*LYSADIGNHPKENRKFNMIKEGPNDDG 0 | |||
0 VPELQGIKGPDIHSPWALNSASVFQVGGTLVETYPQPAV 0 | |||
0* 0 | |||
>DASH_anaPla Anas platyrhynchos (duck) scaffold1769 | |||
0 0 | |||
0 VLHWAASNADYLIPLYCFDPRHYAGTHCYGFPKTGPHRLRFLLESVKDLRETLKKKGS 2 | |||
1 TLVVRKGKPEDVVRDLITQLGCVGVVAFHEE 0 | |||
0 ATQEELDVEKELCQVCEQHGVKVQPFWAATLYHRDDLPFGPIAR 2 | 0 ATQEELDVEKELCQVCEQHGVKVQPFWAATLYHRDDLPFGPIAR 2 | ||
1 LPDVYTQFRKAVEAEAKVRPALQLPDQLKPLAPGVEEGCIPTMEDLGQK 1 | 1 LPDVYTQFRKAVEAEAKVRPALQLPDQLKPLAPGVEEGCIPTMEDLGQK 1 | ||
| Line 2,557: | Line 2,668: | ||
0 GRSPHPRGRRGPADAPAQRKDRGIDFYFSRKKDE* 0 | 0 GRSPHPRGRRGPADAPAQRKDRGIDFYFSRKKDE* 0 | ||
>DASH_melUnd Melopsittacus undulatus (budgerigar) AGAI01061648 | >DASH_taeGut Taeniopygia guttata (finch) ABQF01044665 ABQF01044669 ABQF01044671 synteny: ACAA1 DASH MYD66 OXSR1 | ||
0 MSGTAGTAICLLRCDLRAHDNQ 0 | |||
0 VLHWAQHNADFVIPLYCFDPRHYLGTHCYRLPKTGPHRLRFLLESVKDLRETLKKKGS 2 | |||
1 TLVVRKGKPEDVVCDLITQLGSVTAVVFHEE 0 | |||
0 ATQEELDVEKGLCQVCRQHGVKIQTFWGSTLYHRDDLPFRPIDR 2 | |||
1 LPDVYTHFPKGLESGAKVRPTLRMADQLKPLAPGLEEGSIPTMEDFGQK 1 | |||
2 DPVADPRTAFPCSGGETQALMRLQYYFWDT 0 | |||
0 NLVASYKETRNGLVGMDYSTKFAPW 2 | |||
1 LALGCISPRYIYEQIQKYERERTANESTYW 2 | |||
1 VLFELLWRDYFRFVALKYGRRIFSLR 1 | |||
2 GLQSKDIPWKKDLQLFSCWQ 0 | |||
0 EGKTGVPFVDANMRELSATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0 | |||
0 VDYDVCSNYGNWLYSAGIGNDPRDNRKFNMIKQGLDYDGN 0 | |||
0 GDYVRLWVPELQGIKGADIHTPWALSSAALSQAGVTLGETYPQPVVTAPEWSRHIHRRP 0 | |||
0 GGSPHPRGRRGPAQRKDRGIDFYFSRKKDAC* 0 | |||
>DASH_melUnd Melopsittacus undulatus (budgerigar) AGAI01061648 homopolymer frameshift exon 6 | |||
0 msgtSARTAICLLRCDLRAHDNQ 0 | 0 msgtSARTAICLLRCDLRAHDNQ 0 | ||
0 | 0 VLHWAQSNAEFVVPLYCFDPRHYLRTHWYGFPKTGPYRLRFLLESVKDLRETLKKKGS 2 | ||
1 TLVVRKGKPEDVVRDLITQLGSVSAVVFHEE 0 | 1 TLVVRKGKPEDVVRDLITQLGSVSAVVFHEE 0 | ||
0 ATQEELDVENELCQVCSQHGVKVYTFWASTLYHRDDLPFRPIAR 2 | 0 ATQEELDVENELCQVCSQHGVKVYTFWASTLYHRDDLPFRPIAR 2 | ||
1 LPDVYTHFRKAVESEAKVRPTLRMADQLQPLAPGVEEGCIPTMEDFGQK 1 | 1 LPDVYTHFRKAVESEAKVRPTLRMADQLQPLAPGVEEGCIPTMEDFGQK 1 | ||
2 | 2 DPDTDPRTAFPGSGGETQALMRLQYYFWDT 0 | ||
0 NLVASYKEVRNGLVGMDYSTKFAPW 2 | 0 NLVASYKEVRNGLVGMDYSTKFAPW 2 | ||
1 LALGCISPRYIYEQIKKYERERTANQSTYW 2 | 1 LALGCISPRYIYEQIKKYERERTANQSTYW 2 | ||
| Line 2,573: | Line 2,700: | ||
0 GRSLHPRGSRRPAHSPVQHKDRGIDFYFSRKKDA* 0 | 0 GRSLHPRGSRRPAHSPVQHKDRGIDFYFSRKKDA* 0 | ||
> | >DASH_chrPic Chrysemys picta (turtle) AHGY01416292 AHGY01416294 | ||
0 | 0 MSGAAPRTVICLLRSDLRYHDSE 0 | ||
0 | 0 VLRWAQNNADYVIPLYCFDPRQYLRTYYYGFPKTGPHRLKFLLESVKDLRQTLKKKGS 2 | ||
1 | 1 NLVVRKGNPEDVVHDLITQLGSVTAVAFHEE 0 | ||
0 | 0 ATKEELDVEKALQRVCVQHGVKIQTFWASTLYHRDDLPFRHISR 2 | ||
1 | 1 LPDVYTQFRKAVEAQAKVRPTVQMPDQLKPLAPGVEDGCIPVPEEFGQK 1 | ||
2 | 2 DPLTDPRTAFPCCGGETQALMRLQHYFWDT 0 | ||
0 NLVASYKETRNGLIGIDYSTKFAPW 2 | |||
1 LALGCISPRYIYEQIRKYEKERTANQSTYW 2 | |||
1 VIFELLWRDYFRFVALKYGTRIFLLK 1 | |||
2 GLQDKEVPWKKDPKLFDAWK 0 | |||
0 EGRTGVPFVDANMRELAATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0 | |||
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDSN 0 | |||
0 GDYVRLWVPEIQCVKGGEIHTLWALSNAALSQAGIALGETYPLPVVLAREWSRHINQKP 0 | |||
0 NKRDHPPRGRKDPSDTPRQHKDRGMDFYFSRNKDLS* 0 | |||
0 | |||
1 | |||
1 | |||
2 | |||
0 | |||
0 | |||
0 | |||
0* 0 | |||
>DASH_anoCar Anolis carolinensis (lizard) XM_003221869 14 exons | >DASH_anoCar Anolis carolinensis (lizard) XM_003221869 14 exons | ||
| Line 2,617: | Line 2,729: | ||
0 EGRTGVPFVDANMRELATTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFESLL 0 | 0 EGRTGVPFVDANMRELATTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFESLL 0 | ||
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQALDYDGN 0 | 0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQALDYDGN 0 | ||
0 | 0 GDYARLWVPELQCIRGGDVHTPWALSNGALSQAGVSLGSSYPVPIIKAPEWGRHINQKP 0 | ||
0 SNQGPKGRKGPSHTPKQHKNRGVDFYFSHKKDLS* 0 | 0 SNQGPKGRKGPSHTPKQHKNRGVDFYFSHKKDLS* 0 | ||
>DASH_xenTro Xenopus tropicalis (frog) XM_002938001 BQ392047 CX471210 Pmid: 15147276 synteny: ACAA1 DASH MYD66 transcripts AL790297 CR419606 etc | |||
0 MAARARVIICLLRNDLRLHDNE 0 | |||
0 VLHWAHRNADQIVPLYCFDPRHYGGTHYFNFPKTGPHRLKFLLESVQDLRNTLKERGS 2 | |||
1 NLLLRRGKPEEIIAGLVKQLGNVSAVTLHEE 0 | |||
0 ATKEETDVESAVRRVCTQLGVRYQTFWGSTLYHREDLPFRHISS 2 | |||
1 LPDVYTQFRKAAETQGKVRSTFQMPDRLKPLPSGLEEGSVPTHQDFDQQ 1 | |||
>DASH_xenTro Xenopus tropicalis (frog) XM_002938001 | |||
0 MAARARVIICLLRNDLRLHDNE 0 | |||
0 VLHWAHRNADQIVPLYCFDPRHYGGTHYFNFPKTGPHRLKFLLESVQDLRNTLKERGS 2 | |||
1 NLLLRRGKPEEIIAGLVKQLGNVSAVTLHEE 0 | |||
0 ATKEETDVESAVRRVCTQLGVRYQTFWGSTLYHREDLPFRHISS 2 | |||
1 LPDVYTQFRKAAETQGKVRSTFQMPDRLKPLPSGLEEGSVPTHQDFDQQ 1 | |||
2 DPLTDPRSAFPCCGGETQALQRLHHYFWET 0 | 2 DPLTDPRSAFPCCGGETQALQRLHHYFWET 0 | ||
0 NLVASYKDTRNGLIGIDYSTKFAPW 2 | 0 NLVASYKDTRNGLIGIDYSTKFAPW 2 | ||
| Line 2,647: | Line 2,743: | ||
1 VIFELLWRDYFRFVALKYGRRIFFLR 1 | 1 VIFELLWRDYFRFVALKYGRRIFFLR 1 | ||
2 GLQDKDVPWKKDPKLFDAWK 0 | 2 GLQDKDVPWKKDPKLFDAWK 0 | ||
0 | 0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGIDWRLGAEWFEYLL 0 | ||
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDAG 0 | 0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDAG 0 | ||
0 | 0 GDYIRLWVPELQQIKGGDAHTPWALSTASLAHSNVSLGETYPYPIVMAPEWSRHINQKPS 0 | ||
0 | 0 GSWETSARRGKGPSHTPKQHRNRGIDFYFSRNKNV* 0 | ||
>DASH_xenLae Xenopus laevis (frog) NM_001090969 BU916216 BX850012 PMID: 15147276 match to xenTro: 483/524 (92%) | |||
0 MCVPSRVIICLLRNDLRLHDNE 0 | |||
0 VLHWAHRNADQIVPLYCFDPRHYVGTHYFNFPKTGPHRLKFLLESVRDLRITLKKKGS 2 | |||
1 NLLLRRGKPEEVIEDLVKQLGNVSAVTLHEE 0 | |||
0 ATKEETDVESAVKQACTRLGIKYQTFWGSTLYHREDLPFRHISS 2 | |||
1 LPDVYTQFRKAVETQGKVRPTFQMPDKLKPLPSGLEEGSVPSHEDFDQQ 1 | |||
2 DPLTDPRTAFPCSGGESQALQRLEHYFWET 0 | |||
0 NLVASYKDTRNGLIGLDYSTKFAPW 2 | |||
1 LALGCVSPRYIYEQIGKYEKERTANQSTYW 2 | |||
1 VIFELLWRDYFRFVALKYGRRIFFLR 1 | |||
2 GLQDKDIPWKRDPKLFDAWK 0 | |||
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGIDWRMGAEWFEYLL 0 | |||
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDSG 0 | |||
0 GDYIRLWVPELQQIKGGDAHTPWALSNASLAHANLSLGETYPYPIVMAPEWSRHINQKPA 0 | |||
0 GSWEKSARRGKGPSHTPKQHKNRGIDFYFSRNKDV* 0 | |||
>DASH_hymCut Hymenochirus curtipes (frog) fragment | >DASH_hymCut Hymenochirus curtipes (frog) fragment | ||
| Line 2,660: | Line 2,772: | ||
2 GLAHTNV* 0 | 2 GLAHTNV* 0 | ||
>DASH_ambMex Ambystoma mexicanum (axolotl) | >DASH_ambMex Ambystoma mexicanum (axolotl) JV207726 TSA 73% duck | ||
0 MSVQARTIICLLRNDLRFHDNE 0 | 0 MSVQARTIICLLRNDLRFHDNE 0 | ||
0 VLLWALKNAERIVPLYCFDPRHYVGTHNYNFPKTGPHRLKFLLESVKDLRETLKKKGS 2 | 0 VLLWALKNAERIVPLYCFDPRHYVGTHNYNFPKTGPHRLKFLLESVKDLRETLKKKGS 2 | ||
1 NLLVRKGKPEDVIQELLKQLGSVSAVAFHNE 0 | 1 NLLVRKGKPEDVIQELLKQLGSVSAVAFHNE 0 | ||
0 | 0 VTKEELDVEAAVKQVCLGPGIKIQTFWGSTLYHRDDLPFRHISR 2 | ||
1 | 1 LPDVYTQFRKGVESQGTVRPTLQMPQKVPPLPSGLEEGTIPTASDFGQE 1 | ||
2 GPLADPRSAFPCSGGETQARSRLQHYFWDT 0 | |||
0 NLVASYKDTRNGLIGLDYSTKFAPW 2 | |||
1 LALGCISPRYIYEQIQKYEKERTANQSTYW 2 | |||
1 VIFELLWRDYFRFVALKYGRKIFFLN 1 | |||
2 GLQDKEVPWKKDIQLFDAWK 0 | |||
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0 | |||
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDCN 0 | |||
0 GEYIRLWVPELQGVSGGDAHTPWALSSASLANANVSLGETYPLPIITAPEWGRHINQKP 0 | |||
0 NDRGSTGRRTRGQPYTPKQHKNRGVDFYFSHNKDLH* 0 | |||
>DASH_latCha Latimeria chalumnae (coelocanth) AFYH01055296 AFYH01281932 probable pseudogene | >DASH_latCha Latimeria chalumnae (coelocanth) AFYH01055296 AFYH01281932 probable pseudogene | ||
| Line 2,683: | Line 2,804: | ||
0 * 0 | 0 * 0 | ||
>DASH_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01010414 | >DASH_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01010414 AHAT01010415 | ||
0 MSTIRTIICLLRNDLRFHDNE 0 | 0 MSTIRTIICLLRNDLRFHDNE 0 | ||
0 | 0 vfHWAQGNAEQIVPLYCFDPRHYLGTHCYNFPKTGPFRLRFLLESVKDLRDTLKKNGS 2 | ||
1 NLLVKRGKPENVVSDLIKQLGSVTAVAFHEE 0 | 1 NLLVKRGKPENVVSDLIKQLGSVTAVAFHEE 0 | ||
0 VTKEEQDVETEVTRVCAQFKVRVHTCWGSTLYHREDLPFNHIAR 2 | 0 VTKEEQDVETEVTRVCAQFKVRVHTCWGSTLYHREDLPFNHIAR 2 | ||
| Line 2,699: | Line 2,820: | ||
0 SGGGLSQKGKRGPSHTPKQHRDRGIDFYFSRNNKKLQ* 0 | 0 SGGGLSQKGKRGPSHTPKQHRDRGIDFYFSRNNKKLQ* 0 | ||
>DASH_danRer Danio rerio (zebrafish) NM_205686 | >DASH_danRer Danio rerio (zebrafish) NM_205686 chr24 | ||
0 MSASRTVICLLRNDLRLHDNE 0 | 0 MSASRTVICLLRNDLRLHDNE 0 | ||
0 VFHWAQRNAEHIIPLYCFDPRHYQGTYHYNFPKTGPFRLRFLLDSVKDLRALLKKHGS 2 | 0 VFHWAQRNAEHIIPLYCFDPRHYQGTYHYNFPKTGPFRLRFLLDSVKDLRALLKKHGS 2 | ||
| Line 2,713: | Line 2,834: | ||
0 VDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0 | 0 VDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0 | ||
0 GDYVRQWVPELRGIKGGDVHTPWTLSNSALSHAQVSLNQTYPCPIITAPEWSRHVNNKS 0 | 0 GDYVRQWVPELRGIKGGDVHTPWTLSNSALSHAQVSLNQTYPCPIITAPEWSRHVNNKS 0 | ||
0 | 0 SGSSSSKGRKGSSYTARQHKDRGIDFYFSKNKHF* 0 | ||
>DASH_oreNil Oreochromis niloticus (tilapa) | >DASH_oreNil Oreochromis niloticus (tilapa) GL831138 | ||
0 MSSSRTVICLLRNDLRLHDNE 0 | 0 MSSSRTVICLLRNDLRLHDNE 0 | ||
0 LFHWAQRNAEHIVPLYCFDPTHYVGTYNYSLPKTGPFRLRFLLEGIRDLRNTLINKGS 2 | 0 LFHWAQRNAEHIVPLYCFDPTHYVGTYNYSLPKTGPFRLRFLLEGIRDLRNTLINKGS 2 | ||
| Line 2,728: | Line 2,849: | ||
0 EGRTGVPFVDANMRELAATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFESLL 0 | 0 EGRTGVPFVDANMRELAATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFESLL 0 | ||
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0 | 0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0 | ||
0 | 0 GDYVRGWVPELQNIRGGDVHTPWALSTAVLAHAHVTLGDTYPAPIVTAPEWSRHVNKKA 0 | ||
0 | 0 GGTGPSPRGKKGPSHTPKQHRDRGIDFYFSRSKNL* 0 | ||
>DASH_anoFim Anoplopoma fimbria (sablefish) JO682130 JO685461 | |||
0 MSASRTVICLLRNDLRLHDNE 0 | |||
0 LFHWAQRNAEYIVPLYCFDPRHYVGTYNYNLPKTGPFRLRFLLESIRDLRNTMLTKGS 2 | |||
1 NLVVRRGKPEEVVADLIKQLGSVSTVAFHEE 0 | |||
0 VTSEELNVEKRVKDVCAQMNVKVHTCWGSTLYHRDDLPFHHMSR 2 | |||
1 LPDVYTQFRKAVETQSRVRPLFPTPEQLKPLPKGLEEGAIPTAEDLEQT 1 | |||
2 EPLADPRSAFPCgGGEsqALARLKHYFWDT 0 | |||
0 DAVATYKETRNGLIGADYSTKFAPW 2 | |||
1 LAMGCISPRYIYHQIQQYERERTANQSTYW 2 | |||
1 VIFELLWRDYFKFVGVKYGNRLFQIK 1 | |||
2 GLQEKSVPWKKDMRLFNSWK 0 | |||
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAELFEYLL 0 | |||
0 VDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDGN 0 | |||
0 GDYVRQWVPELQGIKGADVHTPWTLSTASLSHAHVSLGETYPTPIVMAPEWSRHANKKP 0 | |||
0 SGTGPSPRGKKGASHTPKQHRDRGIDFYFSKSKNL* 0 | |||
>DASH_takRub Takifugu rubripes (fugu) CAAB02011066 | |||
0 MSGPRTVICLLRNDLRLFDNE 0 | |||
0 LFHWAQRNADHIVPLYCFDPRHYMGTYHYNLPKTGPFRLRFLLESIKDLRNTLLNKGS 2 | |||
1 NLIVRRGKPEEVVASLIKQLGSVSTVAFHEE 0 | |||
0 VTSEELDVEKRVKDVCAQMKVNVHTCWGSTLYHRDDLPFHHISR 2 | |||
1 LPDVYTQFRKAVESQCRVRPVFPPPEHLKPLPQGLEEGTILTAEDLEQK 1 | |||
2 EPVADPRSAFPCSGGESQALARLKHYFWDT 0 | |||
0 DAVAVYKETRNGLIGVDYSTKFSPW 2 | |||
1 LALGCISPRYIYHQIKQYESERTANQSTYW 2 | |||
1 VIFELLWRDYFRFVAVKYGTKLFQVN 1 | |||
2 GLQDKSVSWRKDMKLFNAWK 0 | |||
0 EGKTGVPFVDANMRELATTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0 | |||
0 VDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0 | |||
0 GEYVRLWVPELQRIMGADVHTPWTLSSAILSHAHLSLGETYPTPIIVAPEWSRHFNKKM 0 | |||
0 SGAGPSPRGRKGPSHTPKQHRDRGIDFYFSRSKNL* 0 | |||
>DASH_gasAcu Gasterosteus aculeatus (stickleback) chrXXI DN658701 | |||
0 MSTSRTVICLLRNDLRLHDNE 0 | |||
0 LFHWAQRNAEYIVPLYCFDPRHYVGTYNYNLPKTGPFRLHFLLEGVKDLRNTMLSKGS 2 | |||
1 NLVVRRGKPEEVVADLIKRLGSVSTVAFHEE 0 | |||
0 VTSEELSVEKRVKDVCARMKVKVHTCWGSTLYHRDDVPFHHISR 2 | |||
1 LPDVYTQFRKAVETQSRVRPLFPTPEQLKPLPEGLEEGAIPTAEDLEQT 1 | |||
2 GPVTDPRSAFPCSGGESRALARLKHYFWDT 0 | |||
0 DAVATYKETRNGLIGMDYSTKFAPW 2 | |||
1 LAMGCISPRYIYHQIQQYEKERTANQSTYW 2 | |||
1 VIFELLWRDYFKFVAVKYGNRLFQIN 1 | |||
2 GLQEKSVPWKKDMTLFNAWK 0 | |||
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAELFEYLL 0 | |||
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDGN 0 | |||
0 GDYVRRWVPELQGIRGADVHAPWTLSTASLSHAGVSLGETYPNPVVVAPEWSRHVNKKP 0 | |||
0 SGARPSPRGKKGPSHTPKQHGDRGIDFYFSRSKNL* 0 | |||
>DASH_oryLat Oryzias latipes (medaka) FS553506 | |||
0 MASSRIVICLLRNDLRLHDNE 0 | |||
0 LFFWAQKNADHIVPLYCFDPRHYVGTYNFNFPKTGPFRLRFLLDSVRDLRNTLLSKGS 2 | |||
1 NLVVRRGKPEEVVADLIKQLGSVSSVAFHEE 0 | |||
0 VASEELNVEKKVKEVCAQMEVKVHTCWGSTLFHRDDLPFPHMAR 2 | |||
1 LPDVYTEFRKAVESKSRVRPVFPTPDRLNSLPPGLEGGAIPTAEDLEQT 1 | |||
2 EPETDPRSAFPCSGGESQALARLKHYFWDT 0 | |||
0 DAVATYKETRNGLIGVDYSTKFSPW 2 | |||
1 LAMGCISPRYIYHQIKKYEQERTANQSTYW 2 | |||
1 VIFELLWRDYFKFVGVKYGNKMFFIK 1 | |||
2 GLQDKSLPWKRDTKLFDAWK 0 | |||
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0 | |||
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQALDYDSN 0 | |||
0 GDYVRQWIPELTAVRGGDVHTPWNLSSAALSRARVSLGETYPTPIVTAPEWGRHFNKKP 0 | |||
0 SGPGPSQKGKKGPSHTPKQHRDRGIDFYFSKSKNL* 0 | |||
>DASH_dicLab Dicentrarchus labrax (seabass) CBN81995 | |||
0 MSTFRTIICLLRNDLRFQDNE 0 | |||
0 LFHWAQRNAEYIVPLYCFDPRHYVGTYNYNLPKTGPFRLRFLLDSIRDLRNTLLSKGS 2 | |||
1 NLVVRQGKPEEVVADLIKQLGSVSAVAFHEE 0 | |||
0 VTSEELNVEKGVKDVCAQMKVKVHTCWGSTLYHRDDLPFHHISR 2 | |||
1 LPDVYTQFRKAVETQSRVRPVFPTPEQLKPLPSGLEEGAIPTAEDLQQT 1 | |||
2 EPLTDPRSAFPCSGGESQVLARLKHYFWDT 0 | |||
0 DAVATYKETRNGLIGVDYSTKFAPW 2 | |||
1 LAMGCISPRYIYHQIKQYEKERTANQSTYW 2 | |||
1 VIFELLWRDYFKFVGVKYGNRLFQVK 1 | |||
2 GLQDKSIPWKKDMKLFNAWK 0 | |||
0 EGQTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0 | |||
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDSN 0 | |||
0 GDYVRQWVPELQGIKGADVHTPWTLSTAALSHAHVSLGETYPNPIVIAPEWSRHVNKKP 0 | |||
0 SGTGPSQRGKRGPPHTPKQHRDRGIDFYFSRSKNL* 0 | |||
>DASH_patPec Patiria pectinifera (starfish) HP101597 | >DASH_patPec Patiria pectinifera (starfish) HP101597 | ||
| Line 2,747: | Line 2,948: | ||
0 QERGKSGKGPGQRQQTRGIDFYFSSGKQGKFKK* 0 | 0 QERGKSGKGPGQRQQTRGIDFYFSSGKQGKFKK* 0 | ||
>DASH_strPur Strongylocentrotus purpuratus (urchin) | >DASH_strPur Strongylocentrotus purpuratus (urchin) SPU_027694 JT098657 last exon compositionally simple expansion | ||
0 MAGRMKIIICLLRNDLRYRDNE 0 | 0 MAGRMKIIICLLRNDLRYRDNE 0 | ||
0 VLFWAHKNATNVIPLYCFDPRHYKGTHQFGFPKTGPHRLKFLLESVRDLRTTLQSVGS 2 | 0 VLFWAHKNATNVIPLYCFDPRHYKGTHQFGFPKTGPHRLKFLLESVRDLRTTLQSVGS 2 | ||
| Line 2,761: | Line 2,962: | ||
0 VDHDVTSNYGNWLYSAGVGNDPRQDRKFNMIKQGLDYDPN 0 | 0 VDHDVTSNYGNWLYSAGVGNDPRQDRKFNMIKQGLDYDPN 0 | ||
0 GDYIRLWVPELSGIKGGSIHMPWTLSSAQLNQAGVSLGETYPSPIVTAPEWSRHSKRP 0 | 0 GDYIRLWVPELSGIKGGSIHMPWTLSSAQLNQAGVSLGETYPSPIVTAPEWSRHSKRP 0 | ||
0 | 0 QSGRGGAGGSQGGGSQGGGRNQSGKGQRRNQGPPRGQQRGLDFYFSNPSQK* 0 | ||
>DASH_aplCal Aplysia californica (sea_hare) scaffold_151:75,790-145,485 | >DASH_aplCal Aplysia californica (sea_hare) scaffold_151:75,790-145,485 mollusk | ||
0 MSSFSSSPVNTVIYLFRNDLRVHDNE 0 | 0 MSSFSSSPVNTVIYLFRNDLRVHDNE 0 | ||
0 ALLLANQKGTRLLPLYCFEPRHYSGTYHFGFPKTGNHRLSFLLDSVKDLQKNLKSRGS 2 | 0 ALLLANQKGTRLLPLYCFEPRHYSGTYHFGFPKTGNHRLSFLLDSVKDLQKNLKSRGS 2 | ||
| Line 2,769: | Line 2,970: | ||
0 VMEEEVKVEKAIEAHVNVPVNTTWGHTLYHVEDLPFQPI 2 | 0 VMEEEVKVEKAIEAHVNVPVNTTWGHTLYHVEDLPFQPI 2 | ||
1 LPDVYTQFRKKVEDNTTIRKCVHVPDALKPLPEGIEVGSLPSYEDLGVS 1 | 1 LPDVYTQFRKKVEDNTTIRKCVHVPDALKPLPEGIEVGSLPSYEDLGVS 1 | ||
2 | 2 EPEKDSRAAFPFLGGETTALERLHSYLWGTDNVSTYKETRNGMIGSDYSTKFSPW 2 | ||
1 | 1 LAHGCLSPRKIYWEIKKYEKERTSNQSTYWVIFELIWRDYFRFVGLKYGNKLFHSG 1 | ||
2 GIKGDRVEWKVNKEQFKAWQ | 2 GIKGDRVEWKVNKEQFKAWQ 1 | ||
2 EGRTGVPYVDANMRELMATGFMSNRGRQ 0 | |||
0 | 0 NVASFLTKDLHLDWRLGAEWFESML 0 | ||
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNVVKQGLDYDAE 0 | 0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNVVKQGLDYDAE 0 | ||
0 GDYVRLWVPELAGIKTGSVHCVWTLSPAVLESGDVSLGQTYPLPLVKAPEWSRHVNRP 0 | 0 GDYVRLWVPELAGIKTGSVHCVWTLSPAVLESGDVSLGQTYPLPLVKAPEWSRHVNRP 0 | ||
0 GSSGGGRDKGPGSARGRGQNKGSHRGQGHQSQQAGQKRGLDFYFSSSKPH* 0 | 0 GSSGGGRDKGPGSARGRGQNKGSHRGQGHQSQQAGQKRGLDFYFSSSKPH* 0 | ||
>DASH_vilLie Villosa lienosa (mussel) JR504188 transcript assembly | >DASH_vilLie Villosa lienosa (mussel) JR504188 transcript assembly mollusk | ||
0 MSKTLTVICLLRNDLRIHDNE 0 | 0 MSKTLTVICLLRNDLRIHDNE 0 | ||
0 VLYWANKYADYVLPVYCFDPRHFKGTHHFNFPKTGPHRLRFLLESIKDLRKNLQSCGS 2 | 0 VLYWANKYADYVLPVYCFDPRHFKGTHHFNFPKTGPHRLRFLLESIKDLRKNLQSCGS 2 | ||
| Line 2,784: | Line 2,985: | ||
0 ATKEELDVEETLRAKCGVKIQTFWGHTLFHRDDLPFGVGQ 2 | 0 ATKEELDVEETLRAKCGVKIQTFWGHTLFHRDDLPFGVGQ 2 | ||
1 LPDVYTQFRKQVEGECVVRDTIDMPRTFKPLPPGIVLDEIPTAEALGVK 1 | 1 LPDVYTQFRKQVEGECVVRDTIDMPRTFKPLPPGIVLDEIPTAEALGVK 1 | ||
2 | 2 EAVSDPRSVFPWSGGETSALERLNQYLWNTDKVATYKETRNGMIGADYSTKFSSW 2 | ||
1 | 1 LALGCLSPRVIYWEIKRYEKERASNQSTYWVVFELLWRDYFRFVALKYGNRIFYLS 1 | ||
2 | 2 GIQGKYIEWKQDTKLFDAWR 0 | ||
0 | 0 DGKTGVPYVDANMRELATTGFMSNRGRQ 0 | ||
0 NVASFLTKDLKLDWRLGAEWFESML 0 | 0 NVASFLTKDLKLDWRLGAEWFESML 0 | ||
0 IDHDVCSNYGNWLYSAGIGNDPREDRKFNMVKQGLDYDPN 0 | 0 IDHDVCSNYGNWLYSAGIGNDPREDRKFNMVKQGLDYDPN 0 | ||
0 GDYVRLWIPELAGVKDGSIHTVWTLNSGALSRAGVSLGKTYPHPILIAPEWNRHMGRT 0 | 0 GDYVRLWIPELAGVKDGSIHTVWTLNSGALSRAGVSLGKTYPHPILIAPEWNRHMGRT 0 | ||
0 KPGMGRGTGSQPNKLKGIDFYFSSGTQRQ* 0 | 0 KPGMGRGTGSQPNKLKGIDFYFSSGTQRQ* 0 | ||
>DASH_rudPhi Ruditapes philippinarum (clam) JO106851 mollusk fragment | |||
1 YYEIKRYEQERVANNSTYWVIFELLWRDYFRYVALKYGNRLFYLS 1 | |||
2 GIQGKQVPWKQDRELFRAWK 0 | |||
0 EGRTGVPYVDANMRELAATGFMSNRGRQ 0 | |||
0 NVASFLTKDLKLDWRLGAEWFESML 0 | |||
0 IDHDVCSNYGNWLYSAGLGNDPREDRKFNMIKQGLDYDPE 0 | |||
0 GDYVRLWVPELQNVSGGNVHTVWTMSNNALAKLGISLGETYPNPITVAPEWSRHYGKS 0 | |||
0 GGASARGGGGGRGRGGYSGGGQSHGHSGRKESAGQKRGIDFYFKGS* 0 | |||
>DASH_dapPul Daphnia pulex (water_flea) ACJG01000530 FE360630 FE400207 FE361270 crustacean mostly re-intronated | |||
0 MSNRVAICLFRNDLRYHDNE 0 | |||
0 VlALAHKSADFVLPLYCFDPRHFEGTHHYKFPKTGIFRTQFLLESVEDFRQTLVKRGSNLMIVHSKPEEALLKIFKSLTGLKVTLILQTEVTKEETDVEKcLQKICQEIKASYINCWG | |||
STLYHKGDLPFQINHVPDSYTGFRKDVEEKLRIRPEISMPDKMKPVPTFAHEIPWGNLPTIEALNSTKPIPNSSSAFPFNGGETAALLRLKSYLWDTNAVAQYKETRNGLIGSDYSTKFSS | |||
WLSHGCLSPRRIHWELEKYELQRTKNQSTYWVRFELLWRDYFKFVSMKYGDRIFYPNGMKGRRQQWKKDMELFKAWQ 1 | |||
2 MGKTGVPFVDANMRELLATGWMSNRGRQNVASFLVKDLLLDWRLGAEWFESLLLDHDVCSNYGNWNYVA 1 | |||
2 GIGNDPRENRKFNMIKQSMDYDLEGNYIRMWVPELREIPGSKIHSPWMLSSGALSAAKIRLGDNYPNPVVVAPEWSRHQKGGK 0 | |||
0 DFGQGNPKGGTQRGIDFYFKNPGGQK* 0 | |||
>DASH_celPug Celuca pugilator (fiddler_crab) JO491098 distal fragment | |||
0 WVLFEMIWRDYFKFVCMKFGDRVFYPSGIMGKKTVWKQNHELFKKWK 1 | |||
2 EGRTGVPFVDANMRELRETGWMSNRGRQNVASFLIKDMGLDWRLGAEWFESQLVDHDVCSNYGNWNYSA 1 | |||
2 GIGNDPRENRKFNMIKQAFDYDPEVTCAVLVLSWLG | |||
>DASH_bemTab Bemisia tabaci (whitefly) EZ942653 HP660316 Hemiptera but phylogenetic isolation and best-blast suggest fungal contamination | |||
MASPKILIYLLRRDLRVHDNPIFHKLTSMSSQANAPFTHLLPLYVFPAHQIEVSGFLSSSDEKSPYPEARSKVGKFWRCGQLRAKFLVESVWDLKQNLETIGSGLEIRVGMLHDVVKQL | |||
VEGFKSKGVQVKGLWMTSEEGYEEKAEERQVRKIIVNAGGDFHLWKDEKYFIDDDDIPFDDPQKYPDVFTKYRNTVEPLREAPRKVLPTPKKLPPLPQNIPPQAHPFKIPGNLKDLIAAL | |||
QKPLDAGLGLKNPPQMPSAGASSAVPFAGGATSGQKRLKHLIESGAMTRYKDTRNGMVGTDYSSKLSLWLALGSLTAREVHSALIDFEEGKTDVGKGAEGYGKGENKGTTHMRFELLWRD | |||
YMRLCTRKYGSRLFLVGGFRNARNIQWKHDNSIMQRWLEGTTGIGLVDAAQRELFLTGFTSNRARQNVASFLTKHLEQDWRLGAEWYECNLVDYDVSSNWGNWQYTAGVGNDPREDRKFN | |||
PVKQASDYDPKAEFVKAWIPEVRELQPEEAWQCWKASYAAKQKPGLRGNIMAERPLAKISFTPRSGGNDGHRGGGRSRGRAQWF* 0 | |||
>DASH_acrMil Acropora millepora (coral) cnidarian JT000937 transcript single exon based on A. digitifera BACK01017766 | |||
MDAQTSNSIAIYLIRNDLRVHDNECLSWAQQNADFVIPLFCFDKEIFGHGAETWHFKFPKTAIHRARFILDSVVDLKDSLKKGGSDLLLRSEQTMKEAVL | |||
GVIQLCRQQSIQNLSLVYQREIAKEEKDVEAELLELCKKENVAVKSFWGLTLYHVEDLPFASVRHLPDTYTEFRKSVEARCRVRPMIPAPNRLKAIPDFI | |||
TSDNMGGIPTLTELVKDQGTTPDARSAFPFCGGETAAIERMNNYLWTTDNVSKYKETRNGLIGAEYSTKLSPWLAVGALSPRKIYECVKQYEKERTANQS | |||
TYWVLFELLWRDYFKFVCFKFGDSVFYLSGIMKKVGLSWKQDMSSFDKWRFGQTGVPFVDANMRELLYTGWMSNRGRQNVASFLVKDLGLDWRLGAEWFE | |||
SLLVDHDVCSNYGNWNYSAGIGNDPRENRKFNMIKQAFDYDADGDFVRLWVPELAGLKGAKVHIPWTLTSSELKVAGITLGESYPRPMVNPPEWKRHTSK | |||
VKGPGNSTRRHNRGIDFYFKSPKDTNPSKKRH* | |||
>DASH_nemVec Nematostella vectensis (sea_anemone) XP_001623243 ABAV01026885 | >DASH_nemVec Nematostella vectensis (sea_anemone) XP_001623243 ABAV01026885 | ||
| Line 2,811: | Line 3,050: | ||
>DASH_monBre Monosiga brevicollis (choanoflagellate) XP_001745157 ABFJ01000402 | >DASH_monBre Monosiga brevicollis (choanoflagellate) XP_001745157 ABFJ01000402 | ||
0 MAKAGNPRPVVVWFRNDLRVHDNEVLLQAAK 0 | 0 MAKAGNPRPVVVWFRNDLRVHDNEVLLQAAK 0 | ||
0 ASHNHVVPVYCFDIRQ | 0 ASHNHVVPVYCFDIRQ 0 | ||
0 YSLVITHRSRRCGQFPKCGRPRARFLIESVDDLRTRLQELGSGLVVRTGLPEEEVARVAAQVGATQVFAHQEVCSEEVAAEHRLKRQLEVPLSLHWGAVTLCHLDDLDFGPRCKHLPSVFTQFRKRVEADMHVRPVVAAPARLAPLPSDLELGSIPTVEDLCPGQH 0 | |||
0 EPDERAVLPFKGGETAARARLQYYLWESNLLAS 2 | 0 EPDERAVLPFKGGETAARARLQYYLWESNLLAS 2 | ||
1 YKDTRNGLVGGDYSSKFSPWLAHGNLTARWIYHE 0 | 1 YKDTRNGLVGGDYSSKFSPWLAHGNLTARWIYHE 0 | ||
| Line 2,822: | Line 3,061: | ||
0 LSSEELAQANIQLGSTYPRPVVDRLDGGRLPLKVRDETAYGPHVQLPQQGLIKMILFDTLFTGLSW* 0 | 0 LSSEELAQANIQLGSTYPRPVVDRLDGGRLPLKVRDETAYGPHVQLPQQGLIKMILFDTLFTGLSW* 0 | ||
> | >DASH_araTha Arabidopsis thaliana (cress) NM_122394 AFMZ01019177 aka:CRY3 PDB:2VTB | ||
0 MNDHIHRVPALTEEEIDSVAIKTFERYALPSSSSVKRKGKGVTILWFRNDLRVLDNDALYKAWSSSDTILPVYCLDPRLFHTTHFFNFPKTG 1 | 0 MNDHIHRVPALTEEEIDSVAIKTFERYALPSSSSVKRKGKGVTILWFRNDLRVLDNDALYKAWSSSDTILPVYCLDPRLFHTTHFFNFPKTG 1 | ||
2 ALRGGFLMECLVDLRKNLMKRGLNLLIRSGKPEEILPSLAKDFGART 0 | 2 ALRGGFLMECLVDLRKNLMKRGLNLLIRSGKPEEILPSLAKDFGART 0 | ||
| Line 3,004: | Line 3,234: | ||
0 LQMVHEGKMHGFLRMYWAKKILEWTSSPEEALHFSLYLNDRYELDGRDPNGYV 1 | 0 LQMVHEGKMHGFLRMYWAKKILEWTSSPEEALHFSLYLNDRYELDGRDPNGYV 1 | ||
2 GCMWSICGIHDQGWAERAVFGKIRYMNYQGCKRKFDVAQFERRYHPKKFSQ* 0 | 2 GCMWSICGIHDQGWAERAVFGKIRYMNYQGCKRKFDVAQFERRYHPKKFSQ* 0 | ||
>CPD_ambMex Ambystoma mexicanum (axolotl) JK977035 fragment 81% allMis | |||
EECKRLNIPFHLLIGFAKDVLPGFIKEHAIGGVVTDFSPLRVPMQWVQDVKELLPEDVPF | |||
VQVDAHNIVPCWVASVKQEYGARTIRNKIHDKVSEFLTEFPPVLVHPHESKFQGEPIDWD | |||
ACIANLQVDRTVGEVDWAKPGTKAGMGVLKSFIAERLKFFGTDRNNPNKNALSNLSPWFH | |||
FGQVSVQRAILEVRKYRSRFKESVEGFIEEAMVRRELADNFCFYNKKYDTVEGAYDWAKNT | |||
>CPD_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01034265 | >CPD_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01034265 | ||
| Line 3,181: | Line 3,417: | ||
2 GYVGCMWSIAGIHDQGWAERAVFGKIR 21 YMNYKGCKRKFDVARFVSRFPQAVANAA* 0 | 2 GYVGCMWSIAGIHDQGWAERAVFGKIR 21 YMNYKGCKRKFDVARFVSRFPQAVANAA* 0 | ||
>CPD_araTha Arabidopsis thaliana (cress) | >CPD_araTha Arabidopsis thaliana (cress) NM_179320 AFMZ01000529 PHR1 GC-AG splice exon 6-7 | ||
0 MASTVSVQPGRIRILKKGSWQPLDQTVGPVVYWMFRDQRLKDNWALIHAVDLANRTNAPVAVVFNLFDQFLDAKARQLGFMLKGLRQLHHQIDSLQIPFFLLQ 0 | 0 MASTVSVQPGRIRILKKGSWQPLDQTVGPVVYWMFRDQRLKDNWALIHAVDLANRTNAPVAVVFNLFDQFLDAKARQLGFMLKGLRQLHHQIDSLQIPFFLLQ 0 | ||
0 GDAKETIPNFLTECGASHLVTDFSPLREIRRCKDEVVKRTSDSLAIHEVDAHNVVPMWAASSKLEYSARTIRGKINKLLPDYLIEFPKLEPPKKKWTGMMDKKLVDWDSLIDKVVR 2 | 0 GDAKETIPNFLTECGASHLVTDFSPLREIRRCKDEVVKRTSDSLAIHEVDAHNVVPMWAASSKLEYSARTIRGKINKLLPDYLIEFPKLEPPKKKWTGMMDKKLVDWDSLIDKVVR 2 | ||
| Line 3,204: | Line 3,440: | ||
0 GWKERPVFGKIRYMNYAGCKRKFDVDAYISYVKRLAGQSKKRNAEESPNPVVKLSKSQH* 0 | 0 GWKERPVFGKIRYMNYAGCKRKFDVDAYISYVKRLAGQSKKRNAEESPNPVVKLSKSQH* 0 | ||
=== Cryptochromes from cnidarians, trichoplax, | === Cryptochromes from cnidarians, ctenophore, trichoplax, sponge, choanoflagellate and diatom === | ||
This collection of cryptochromes and photolyases serves as deeper outgroups to bilateran sequences. Some do not classify clearly | This collection of cryptochromes and photolyases serves as deeper outgroups to bilateran sequences. Some do not classify clearly; others represent clade-specific gene family expansions of litle general interest. Those with minimal or novel introns may represent retroprocessed genes that were subsequently re-intronated at new locations. Those clearly orthologous to DASH and CPD have been moved to those sections. As the remaining sequences are classified, they will be moved to their corresponding orthologous class above to the extent that is useful. | ||
The | The Arabidopsis cryptochromes/photolyases have been re-named according to contemporary classification but with common synonyms provided in their fasta header. To collect all the available sequences for this or any other species, simply use web browser search (eg '_araTha') or sort the [[Cryptochrome_evolution#247_curated_refSeqs_for_metazoan_cryptochromes_and_photolyases|summary page of fasta headers]] by species. | ||
The phylogenetic position of ctenophores is uncertain (sister to cnidarians vs earlier branching). The genome assembly of Mnemiopsis contains only CRY64 and an ancient duplicated pair of CPD cryptochromes. While these bear no special relationship to cnidarian counterparts (clustering instead with invertebrate representatives), that might be attributable to rapid evolution of cnidarians rather than a non-sistered topology. Use of maximal likelihood cannot resolve this because it requires modeling evolutionary processes over a vast span (600 myrs of geologic time but tens of billions of branch length time) and so the result would be no more reliable than its untestable assumptions. | |||
>CRY64_mneLei Mnemiopsis leidyi (sea_walnut) AGCP01007162 ctenophore no introns 49% identity to closest refSeq CPD | |||
MGKIALHWFRHGLRLHDNTPLQKAIAGCTSLIPLYILDTEYFKPGKVGINRMGFLLDSLKALDTDLRAIGSRLYVAKGDPEEVIGKFIKEHKIGAVSFER | |||
DTEPYNKVMDGKIINLTKELNVEAFPLWGHTMFDPEYLLALNNGDAPLTMTSFLRLMSEAGDPPKPIDPPKSLPPPPADSLVCKDVFIFQGVPTLSDLTE | |||
YEFNPKDYTTWFVAGEKEGIRVMNEFLAQKRRVSTFEKPKTDPTALQPDTTALSPYITRGSLSSRTFYHGLKDTLKGMKSSKPPVSLKGQLYWREMAYLI | |||
GFSVPNFNQMEGNPICKQIPWLTGDDAKALLDKWEMGQTGFPAVDAVMNQLRTEGWMHHLARHLVACFLTRGDLWVTWELGRDVFEKHLVDADWSINNFS | |||
WHWLSCSAFFHQYFRCYSPIAFFKKTDPNGNYIKKHVPILAKFPDKYIYEPWTAPKGIQIACGCIIGKDYPKPMVEHQFVCQENKSRMKKAYDTSKKRDS | |||
GEPPAKKLKQNNLSKYRTK* | |||
>CPD1_mneLei Mnemiopsis leidyi (sea_walnut) AGCP01013608 ctenophore no introns 54% identity to closest refSeq CPD | |||
MASDKEPAPKRQKTEEIKFNMSRLKQLHGGQGAPSNVTAVAYYMHRDQRVQDNWALIYAQSLATRHSAPLHVVTLITTSHPEQRGATFRHLQFCFDGLKE | |||
VSEELSELNIAFHLLIDQGGKVGGGKVAKWMKECEVDCLVTDFSPLRQHRSLLAQLTKSTDLPDSATVYQVDAHNIVPVEVTSDKQEYAARTIRNKIMSK | |||
LGKYLTEFPPVTKHPFGDAAMATRHFAEETGTSAKLKEDWDSVLQNLKLDFSVKPSELYKGGTKAGMSCLEDFVERRIKRYSDKRNDPTEDAISDLSPWL | |||
HMGNLSAQRAVLYVKKHASSYSSVFIEEAVVRRELSDNFCFYNQNYDSIKGAANWAQETLKAHRDDKREYVYTRQQFLEAKTHDNLWNAAQKQLVREGKM | |||
HGFMRMYWAKKILEWTASPEEALAEALYFNDHFSLDGNDANGFVGCMWSICGVHDQGWRERSVFGKIRYMNYNGCKRKFDIEKYIAKYS* | |||
>CPD2_mneLei Mnemiopsis leidyi (sea_walnut) AGCP01006228 ctenophore no introns 49% identity to closest refSeq CPD | |||
MNYLSFNVKRCRLLGGNDNIISKSSGIAYYMHRDQRVQDNWAFVYSQDLAVKHNLPLYVLAGFNVKHPENPEGTRRAIDFTIGGLKEVEKECTELGIQFH | |||
MFKDHHVPMFERILKFIKETSVRCIVADFSPLHPHRNQMNELAKRLDSSSCLLQVDAHNIVPVWEASKKDEPRATEMRNQIMPQFDEYFTEFPRIKNHPT | |||
KVLDPLPAVNWEEIRNSVTVDETVQSCEWAKPGTSHGMQHLKRFLTEGLHIYNEKRNIPTVDAISNLSPWFHSGQVSAQRAVIEVKKFEEQYPKSVYKFV | |||
DEAVVWSEMSDNFCFYNDNYDKLSGAPQWAQDTLNEHRSDKREYVYTKEQFEKAQTHDNLWNAAQIQLRDEGKMHGFMRMYWAKKILEWSENPDEAIAIA | |||
LYLNDHYSLDGADSNGFAGVMWSIAGVHDPPKWGERPVYGKIRYMSYNGCKGKFDIFEYIKKYQSGDAASAPQGGLHKYYKGADPKQKGQGSGRVQGSWG | |||
GGKQ* | |||
>CRY1A_acrMil Acropora millepora (coral) EF202589 | >CRY1A_acrMil Acropora millepora (coral) EF202589 | ||
MSLNLKSVEDNNSAVSAEKSQGKLKAKHAIHWVRKDLRLHDNPSLLEAVKGSDTVRIIYVLDTKVDHATGIGLNLWRFLLQSLEDVDDSLRKLNSRLFVV | MSLNLKSVEDNNSAVSAEKSQGKLKAKHAIHWVRKDLRLHDNPSLLEAVKGSDTVRIIYVLDTKVDHATGIGLNLWRFLLQSLEDVDDSLRKLNSRLFVV | ||
| Line 3,353: | Line 3,613: | ||
=== The 4Fe-4S photolyases and primases === | === The 4Fe-4S photolyases and primases === | ||
A small | A small subset of available sequences are shown as these suffice for GenBank probes. PFES stands for photolyase with a 4Fe-4S redox cluster. These are entirely prokaryotic. | ||
>PFES_agrTum Agrobacterium tumefaciens (bacteria) NP_355900 aka: PhrB | >PFES_agrTum Agrobacterium tumefaciens (bacteria) NP_355900 aka: PhrB | ||
| Line 3,387: | Line 3,647: | ||
IQQEMDLLRFRFSILPKDKIQDFLKDSQLQFEAISDEEKTLREQEIVASSPSLSGLKLGFESIYKIPFADALDLFRGRKVYLEDGFAYVPLKDIVAIILNEFRAKLSKALALTARSLPAV | IQQEMDLLRFRFSILPKDKIQDFLKDSQLQFEAISDEEKTLREQEIVASSPSLSGLKLGFESIYKIPFADALDLFRGRKVYLEDGFAYVPLKDIVAIILNEFRAKLSKALALTARSLPAV | ||
QSDERLQPLLNHLSHSYTGQDYSTQGNVGKI<font color=blue>SLDQIDLLSTKSFPPCMRQLHKALRENHHLRHGGRMQYGLFLKGIGLTLEQALQFWKQEFIKGKMDPDKFDKGYSYNIRHSFGKEGKRT | QSDERLQPLLNHLSHSYTGQDYSTQGNVGKI<font color=blue>SLDQIDLLSTKSFPPCMRQLHKALRENHHLRHGGRMQYGLFLKGIGLTLEQALQFWKQEFIKGKMDPDKFDKGYSYNIRHSFGKEGKRT | ||
DYTPFSCLKIILSNPPSQGDYHGCPFRHSDPELLKQKLQSYKISPGGISQILDLVKGTHYQVACQKYFEMIHNVDDCGFSLNHPNQFFCESQRILNGGKDIKKE</font>PIQPETPQPKPSVQKT* | DYTPFSCLKIILSNPPSQGDYHGCPFRHSDPELLKQKLQSYKISPGGISQILDLVKGTHYQVACQKYFEMIHNVDDCGFSLNHPNQFFCESQRILNGGKDIKKE</font>PIQPETPQPKPSVQKT* | ||
KDASSALASLNSSLEMDMEGLEDYFSEDS* 0 | KDASSALASLNSSLEMDMEGLEDYFSEDS* 0 | ||
>PRIM2_sacCer Saccharomyces cerevisiae (yeast) P20457 aka: PRI2_YEAST primase large subunit <font color=blue>PDB|3LGB</font> | >PRIM2_sacCer Saccharomyces cerevisiae (yeast) P20457 aka: PRI2_YEAST primase large subunit <font color=blue>PDB|3LGB</font> | ||
MFRQSKRRIASRKNFSSYDDIVKSELDVGNTNAANQIILSSSSSEEEKKLYARLYESKLSFYDLPPQGEITLEQFEIWAIDRLKILLEIESCLSRNKSIK | MFRQSKRRIASRKNFSSYDDIVKSELDVGNTNAANQIILSSSSSEEEKKLYARLYESKLSFYDLPPQGEITLEQFEIWAIDRLKILLEIESCLSRNKSIK | ||
EIETIIKPQFQKLLPFNTESLEDRKKDYYSHFILRLCFCRSKELREKFVRAETFLFKIRFNMLTSTDQTKFVQSLDLPLLQFISNEEKAELSHQLYQTVS | EIETIIKPQFQKLLPFNTESLEDRKKDYYSHFILRLCFCRSKELREKFVRAETFLFKIRFNMLTSTDQTKFVQSLDLPLLQFISNEEKAELSHQLYQTVS | ||
ASLQFQLNLNEEHQRKQYFQQEKFIKLPFENVIELVGNRLVFLKDGYAYLPQFQQLNLLSNEFASKLNQELIKTYQYLPRLNEDDRLLPILNHLSSGYTI | ASLQFQLNLNEEHQRKQYFQQEKFIKLPFENVIELVGNRLVFLKDGYAYLPQFQQLNLLSNEFASKLNQELIKTYQYLPRLNEDDRLLPILNHLSSGYTI | ||
ADFNQQKANQFSENVD<font color=blue>DEINAQSVWSEEISSNYPLCIKNLMEGLKKNHHLRYYGRQQLSLFLKGIGLSADEALKFWSEAFTRNGNMTMEKFNKEYRYSFR | ADFNQQKANQFSENVD<font color=blue>DEINAQSVWSEEISSNYPLCIKNLMEGLKKNHHLRYYGRQQLSLFLKGIGLSADEALKFWSEAFTRNGNMTMEKFNKEYRYSFR | ||
HNYGLEGNRINYKPWDCHTILSKPRPGRGDYHGCPFRDWSHERLSAELRSMKLTQAQIISVLDSCQKGEYTIACTKVFEMTHNSASADLEIGEQTHIAHP | HNYGLEGNRINYKPWDCHTILSKPRPGRGDYHGCPFRDWSHERLSAELRSMKLTQAQIISVLDSCQKGEYTIACTKVFEMTHNSASADLEIGEQTHIAHP | ||
NLYFERSRQLQK</font>KQQKLEKEKLFNNGNH* 0 | NLYFERSRQLQK</font>KQQKLEKEKLFNNGNH* 0 | ||
== Alignment-ready sequences == | == Alignment-ready sequences == | ||
The sequence sets above can be aligned as-is but for most | The sequence sets above can be aligned as-is but for most purposes -- especially comparing one orthology class to another -- it is preferable to trim off non-conserved variable length regions at the N- and C-termini and also to groom sequences by removing insertions limited to a single species especially in outlier taxa as these create gap columns that distract from the central evolutionary story. | ||
Some assemblies have missing exons, so when the rate of divergence is low, the exon can be infilled ( | Some assemblies have missing exons, so when the rate of divergence is low, the exon can be infilled (ie predicted) from the closest related species. For example, zebra finch is missing the first exon of CPD but that is available from other songbirds (which are more suitable than chicken or duck). While this loses true variation, it improves products such as logos and overlays on 3D structures. | ||
However it is counterproductive to infill sequences with multiple missing exons. These are included in the sequence compilation primarily to establish the presence/absence of a gene in the given phylogenetic position. For narrowly targeted purposes, their exons can be informative. However the very fact of poor coverage speaks to the quality of sequencing project, meaning observed variation is not necessarily reliable. Thus the primary use is in confirming change (or not) implied by | However it is counterproductive to infill sequences with multiple missing exons. These are included in the sequence compilation primarily to establish the presence/absence of a gene in the given phylogenetic position. For narrowly targeted purposes, their exons can be informative. However the very fact of poor coverage speaks to the quality of sequencing project, meaning observed variation is not necessarily reliable. Thus the primary use is in confirming change (or not) implied by the complete sequences. | ||
Finally, pseudogenes are not appropriate for groomed alignment inclusion because they introduce spurious amino acid variation not informative to structure or function. The issue here is not older pseudogenes riddled with frameshifts and internal stop codons, but rather newer pseudogenes that have not yet accrued grossly inappropriate features. | Finally, pseudogenes are not appropriate for groomed alignment inclusion because they introduce spurious amino acid variation not informative to structure or function. The issue here is not older pseudogenes riddled with frameshifts and internal stop codons, but rather newer pseudogenes that have not yet accrued grossly inappropriate features. | ||
These newer pseudogenes can be recognized because their pattern of amino acid substitution does not respect the pattern of conservation within the protein -- a residue strictly invariant for a billion years | These newer pseudogenes can be recognized because their pattern of amino acid substitution does not respect the pattern of conservation within the protein -- only in a pseudogene is a residue strictly invariant for a billion years as likely to be substituted as an inconsequential loop residue. Operationally, the 'dot product' of observed substitutions of a subtle pseudogene with the conservation logo will score too high (ie outside the cluster defined by the non-pseudogene sequences). However very recent pseudogenes cannot be detected by any method. These contribute an effect similar to residual sequencing error and individual polymorphisms. | ||
=== CRY7 === | |||
The 14 CRY7 sequences become quite conserved after a ragged amino terminal start. The [http://genomewiki.ucsc.edu/index.php/Image:TrimmedGroomedCRY7.doc groomed and gapped sequence set] from initial to final regions of conservation extends for 774 amino acids, even after removing an internal spacer region. The sequences are in phylogenetic order relative to Xenopus, the only tetrapod to retain the gene. | |||
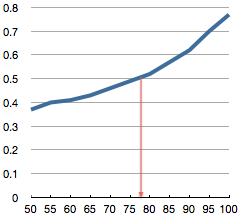
Following the curated sequences, the first alignment is conventional (full sequences for all species), allowing motifs of interest to be located by web browser text search. The second alignment, made with [http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_multalin.html Multalin2] is a difference alignment set to display an amino acid position (alignment column) only when 91% or more residues in an alignment column agree. The figure 91% is chosen so 50% of the total amino acids are displayed. | |||
CRY7 is highly conserved despite its restricted phylogenetic distribution. The sequences here omit two fragmentary genes from mollusk. The erratically evolving connector domain following the UIM motif has been trimmed out (3 spaces show its location) to allow for easier comparisons to other members of the gene family. Short non-conserved regions have also been removed from the N- and C- termini. | |||
[[Image:CRY7weblogo.gif|200px|thumb|left|CRY7 weblogo: click for full size]] | |||
The best options for modeling the spacial locations of the downstream ultra-conserved residues are Drosophila CRY64 or an Arabidopsis CRY1A. Since the percent identities are fairly low, the two fold domains will be recapitulated but the precise orientations of sidechains will remain problematic. For example it will not prove feasible to dock potential antenna molecules to that domain. Nonetheless, comparisons can be made to conserved residues in other members of the gene family, notably tryptophans. The novel upstream domain cannot be modeled at all for lack of template, though the UIM ubiquitin domain is known to be a plain alpha helix. | |||
<br clear=all> | |||
CRY7_xenTr ultra-conserved amino acid positions in frog: | |||
CRY7_xenTro ....G......F.....S.LG...T...F......L..........L......YF...............L.. ..........RR.R.K...----..........PVL.W.RRDLRL.DNPAL...L..G.PVIP.F.W...EE.G...T.A.GGA.KYWLH.AL......L...GS..... | |||
CRY7_xenTro ...-------.....L..L...TGA.T....A.YEPWL..RD......L...GV.....HSYCL..P..V.T.GVGLRGIGSVSHF..CC..N.....G..L..P..LP.P..WP....L..L.L..MP.RKDGT..DWA..IR..WDFSE.GA...L..FL.DGV..YEKES.RAD.P.TS..SP | |||
CRY7_xenTro YLHFGQ.S.R.....A........KF.RKLAWRDLAYW...LFP..P.E..RP.YK..RWS.D..HL.AWQ.G.TGYPLVDAAMR.LW.TGWM.NY.RHVVASFL.AYLH..W..GYRWFQDTL.DADVAI.AMMWQNGGM.GLDHWNFVMHPVD.A.TCDP.G.YVRKWCPEL..LPD..IHKPW | |||
CRY7_xenTro .C..S.LRRAGV..G..YP.RI..DLEERR..SL.DV..VR.......D..SGCD....P..L....LG..............FLLPVITR.EFK.....P...--.NPY..VLKGYVSR.RDE..A......FTAS...E...R.ER.....R..EGLP..........RT..-.D..S..P... | |||
>CRY7_xenTr ultra-conserved amino acids in frog for display on CRY64_droMel 1U3C 34% identity | |||
<font color =gray>QLES<font color =red>G</font>SVQADE<font color =red>F</font>LCLVL<font color =red>S</font>I<font color =red>LG</font>SSR<font color =red>T</font>YSQ<font color =red>F</font>PAILQS<font color =red>L</font>SRKEPAMYRE<font color =red>L</font>MDLHAE<font color =red>YF</font>RKEPADLETLGYETD<font color =red>L</font>EL RPRESKAKHS<font color =red>RR</font>S<font color =red>R</font>K<font color =red>K</font>KKS APSRGLVAMK<font color =red>PVL</font>V<font color =red>W</font>F<font color =red>RRDLRL</font>H<font color =red>DNPAL</font>ISA<font color =red>L</font>EH<font color =red>G</font>V<font color =red>PVIP</font>V<font color =red>F</font>L<font color =red>W</font>CIN<font color =red>EE</font>T<font color =red>G</font>QNF<font color =red>T</font>L<font color =red>A</font>T<font color =red>GGA</font>T<font color =red>KYWLH</font>H<font color =red>AL</font>LKLNQS<font color =red>L</font>QRF<font color =red>GS</font>HIIFR | |||
VAR SCEEE<font color =red>L</font>VS<font color =red>L</font>VHE<font color =red>TGA</font>D<font color =red>T</font>IIIN<font color =red>A</font>V<font color =red>YEPWL</font>KE<font color =red>RD</font>DLISET<font color =red>L</font>RRH<font color =red>GV</font>ELKKH<font color =red>HSYCL</font>YE<font color =red>P</font>DS<font color =red>V</font>S<font color =red>T</font>E<font color =red>GVGLRGIGSVSHF</font>MS<font color =red>CC</font>KR<font color =red>N</font>NSAPI<font color =red>G</font>MP<font color =red>L</font>DA<font color =red>P</font>RC<font color =red>LP</font>A<font color =red>P</font>CN<font color =red>WP</font>ESDH<font color =red>L</font>DT<font color =red>L</font>E<font color =red>L</font>GK<font color =red>MP</font>H<font color =red>RKDGT</font>LI<font color =red>DWA</font>VT<font color =red>IR</font>ES<font color =red>WDFSE</font>D<font color =red>GA</font>YTC<font color =red>L</font>AN<font color =red>FL</font>Q<font color =red>DGV</font>KH<font color =red>YEKES</font>G<font color =red>RAD</font>K<font color =red>P</font>Y<font color =red>TS</font>HI<font color =red>SP</font> | |||
<font color =red>YLHFGQ</font>I<font color =red>S</font>P<font color =red>R</font>TVLHE<font color =red>A</font>YFTKKNVP<font color =red>KF</font>L<font color =red>RKLAWRDLAYW</font>LLI<font color =red>LFP</font>DM<font color =red>P</font>S<font color =red>E</font>PV<font color =red>RP</font>A<font color =red>YK</font>SQ<font color =red>RWS</font>S<font color =red>D</font>LN<font color =red>HL</font>R<font color =red>AWQ</font>K<font color =red>G</font>L<font color =red>TGYPLVDAAMR</font>E<font color =red>LW</font>L<font color =red>TGWM</font>C<font color =red>NY</font>S<font color =red>RHVVASFL</font>V<font color =red>AYLH</font>IH<font color =red>W</font>VH<font color =red>GY</font>R<font color =red>WFQDTL</font>L<font color =red>DADVAI</font>N<font color =red>AMMWQNGGM</font>S<font color =red>GLDHWNFVMHPVD</font>S<font color =red>A</font>L<font color =red>TCDP</font>Y<font color =red>G</font>S<font color =red>YVRKWCPEL</font>AG<font color =red>LPD</font>EY<font color =red>IHKPW</font> | |||
K<font color =red>C</font>AP<font color =red>S</font>Q<font color =red>LRRAGV</font>IL<font color =red>G</font>RN<font color =red>YP</font>H<font color =red>RI</font>VL<font color =red>DLEERR</font>EQ<font color =red>SL</font>K<font color =red>DV</font>VEV<font color =red>R</font>KKHLEYL<font color =red>D</font>EV<font color =red>SGCD</font>MVQI<font color =red>P</font>DQ<font color =red>L</font>LALT<font color =red>LG</font>TSGEDEVVRNRTGS<font color =red>FLLPVITR</font>K<font color =red>EFK</font>YKTLQ<font color =red>P</font>DTK D<font color =red>NPY</font>NT<font color =red>VLKGYVSR</font>K<font color =red>RDE</font>TI<font color =red>A</font>YMNERH<font color =red>FTAS</font>TIN<font color =red>E</font>GAQ<font color =red>R</font>H<font color =red>ER</font>IERTN<font color =red>R</font>LM<font color =red>EGLP</font>APSDAKNKSR<font color =red>RT</font>PK K<font color =red>D</font>PF<font color =red>S</font>II<font color =red>P</font>PSY</font> | |||
=== CRY64 === | |||
The 27 available CRY64 sequences do not need significant trimming at either the N- or C-terminus. The latter is somewhat peculiar in that the penultimate exon is quite ragged at its end (though preserving its phase 12 splice donor) but the last exon is quite conserved in character (many basic residues) at least in vertebrates. Since the alignment from lizard to sponge is very gappy here, the alignment is splice-driven (that is, forced to agree at the start of the final exon. This exon corresponds to the third from the end in CRY1; in other words, CRY64 lacks any specialized extension as befits its straightforward DNA repair role. | |||
CRY64 has a moderate level of conservation overall but certain interior regions are quite invariant, even at the 90% level over the entirety of metazoan evolution (sponge to lizard). That localization of conservation can be seen thematically in the first image below -- it is predominantly concentrated in the catalytic FAD domain. Vertebrate CRY64 are easily modeled using one of eight Drosophila structural determinations; these have 57% identity over 91% of the length (7 IHWFR...KAAYA 488). This template does not cover the terminal basic exon of vertebrate protein. The [http://genomewiki.ucsc.edu/index.php/Image:TrimmedgroomedCRY64.doc groomed and gapped sequence set] file also contains an alignment colored by conservation and a difference alignment relative to the CRY64 consensus sequence. | |||
[[Image:CRY64p90.png|left]] | |||
[[Image:CRY64weblogo.gif|300px|thumb|left|Weblogo of CRY64]] | |||
<br clear=all> | |||
Ultra-conserved residues of CRY64 displayed within lizard sequence for purposes of alignment with paralogs: | |||
CRY64_anoCar M IHWFR LRLHDNPAL A PIFILD VG NR RFL L DLD L N RLFVVRG E! L W V LT E D EPY RD V V V HTLYD D I N G PLTY P P | |||
CRY64_anoCar Y P L EL F GGE EAL RL WV FEKP PN PSTTVLSPY FGCLS R F EV S QL WREFFYT PNF M GN Q WD N E L W E TGYPFIDA M QL EGWIH | |||
CRY64_anoCar HLARH VACFLTRGDLWISWEEG VFEE LLD D LNA NW WLS S FF Q FRVYSP FGKKTD G YI KY P L KFP YIYEPW AP Q AGCI G DYP IV H N M AY | |||
>CRY64_conSer Ultra-conserved residues of CRY64 for display on CRY64_droMel 3CVV 59% identity | |||
<FONT COLOR = GRAY><FONT COLOR = RED>M</FONT>MAHVS<FONT COLOR = RED>IHWFR</FONT>KG<FONT COLOR = RED>LRLHDNPAL</FONT>LA<FONT COLOR = RED>A</FONT>MKNSAEIY<FONT COLOR = RED>PI</FONT>F<FONT COLOR = RED>ILD</FONT>PWFPKNMQVSI<FONT COLOR = RED>NR</FONT>W<FONT COLOR = RED>RFL</FONT>IES<FONT COLOR = RED>L</FONT>K<FONT COLOR = RED>DLD</FONT>ES<FONT COLOR = RED>L</FONT>KKL<FONT COLOR = RED>N</FONT>S<FONT COLOR = RED>RL</FONT>F<FONT COLOR = RED>VVRG</FONT>RPA<FONT COLOR = RED>EV</FONT>FPE<FONT COLOR = RED>L</FONT>FTK<FONT COLOR = RED>W</FONT>K<FONT COLOR = RED>V</FONT>TR<FONT COLOR = RED>L</FONT>AF<FONT COLOR = RED>E</FONT>V<FONT COLOR = RED>D</FONT>T<FONT COLOR = RED>EP</FONT>YAR-<FONT COLOR = RED>RD</FONT>AE<FONT COLOR = RED>V</FONT>VRLAAEHG<FONT COLOR = RED>V</FONT>Q<FONT COLOR = RED>V</FONT>IQKVS<FONT COLOR = RED>HTLYD</FONT>T<FONT COLOR = RED>E</FONT>R<FONT COLOR = RED>I</FONT>IVE<FONT COLOR = RED>N</FONT>S<FONT COLOR = RED>G</FONT>KA<FONT COLOR = RED>P</FONT>L<FONT COLOR = RED>TY</FONT>TRLQTLVASLGP<FONT COLOR = RED>P</FONT>KQPVPA<FONT COLOR = RED>P</FONT>KLEDMK | |||
DCCTPVKEDHDLEYGT<FONT COLOR = RED>P</FONT>SYE<FONT COLOR = RED>EL</FONT>GQDPKTAGPHLYP<FONT COLOR = RED>GGE</FONT>T<FONT COLOR = RED>EAL</FONT>A<FONT COLOR = RED>RLD</FONT>LHMKRTS<FONT COLOR = RED>WV</FONT>CN<FONT COLOR = RED>F</FONT>K<FONT COLOR = RED>KP</FONT>ETH<FONT COLOR = RED>PN</FONT>SLT<FONT COLOR = RED>PSTTVLSPY</FONT>VK<FONT COLOR = RED>FGCLS</FONT>V<FONT COLOR = RED>R</FONT>M<FONT COLOR = RED>F</FONT>WWKLA<FONT COLOR = RED>EV</FONT>YQGRKH<FONT COLOR = RED>S</FONT>DPPVSLHG<FONT COLOR = RED>QL</FONT>L<FONT COLOR = RED>WREF</FONT>F<FONT COLOR = RED>YT</FONT>AGAGI<FONT COLOR = RED>PNFD</FONT>R<FONT COLOR = RED>M</FONT>EN<FONT COLOR = RED>N</FONT>PVCV<FONT COLOR = RED>QV</FONT>D<FONT COLOR = RED>WD</FONT>N<FONT COLOR = RED>N</FONT>Q<FONT COLOR = RED>E</FONT>Y<FONT COLOR = RED>L</FONT>RA<FONT COLOR = RED>WREG</FONT>Q<FONT COLOR = RED>TGYPFIDA</FONT>I<FONT COLOR = RED>M</FONT>T<FONT COLOR = RED>QL</FONT>RT<FONT COLOR = RED>EGWIH</FONT> | |||
<FONT COLOR = RED>HLARH</FONT>A<FONT COLOR = RED>VACFLTRGDLWISWEEG</FONT>QK<FONT COLOR = RED>VFEE</FONT>L<FONT COLOR = RED>LLD</FONT>A<FONT COLOR = RED>D</FONT>WS<FONT COLOR = RED>LNA</FONT>A<FONT COLOR = RED>NW</FONT>Q<FONT COLOR = RED>WLS</FONT>A<FONT COLOR = RED>S</FONT>A<FONT COLOR = RED>FF</FONT>H<FONT COLOR = RED>Q</FONT>F<FONT COLOR = RED>FRVYSPV</FONT>T<FONT COLOR = RED>FGKKTD</FONT>K<FONT COLOR = RED>NG</FONT>E<FONT COLOR = RED>YI</FONT>K<FONT COLOR = RED>KY</FONT>L<FONT COLOR = RED>P</FONT>F<FONT COLOR = RED>L</FONT>R<FONT COLOR = RED>K</FONT>FSN<FONT COLOR = RED>DYI</FONT>Y<FONT COLOR = RED>EPW</FONT>K<FONT COLOR = RED>AP</FONT>RS<FONT COLOR = RED>LQ</FONT>ER<FONT COLOR = RED>AGCIIG</FONT>Q<FONT COLOR = RED>DYP</FONT>KP<FONT COLOR = RED>IVEH</FONT>EKVYKR<FONT COLOR = RED>N</FONT>LER<FONT COLOR = RED>M</FONT>KA<FONT COLOR = RED>AY</FONT>ARRSPNLVIQAKDKVSQKK-GVNRKRPEAPTKAKVQAKKV<font color=red>R</font><font color=red>K</font></font> | |||
=== DASH === | |||
DASH is a relatively isolated sequence class with an uneven phylogenetic distribution that implies multiple gene loss events, both old and new. The 35 sequences provided here range from sponge to amniotes; conservation is moderate relative to other members of its gene family. The [http://genomewiki.ucsc.edu/index.php/Image:TrimmedgroomedDASH.doc groomed and gapped] sequence set shows rather stricter conservation of the catalytic FAD domain. The final region of the protein requires manual alignment utilizing the last intron to insure homological register in a very gappy region. The final RGIDFYF motif can then be aligned but some arbitrary gap placement is still necessary even in closely related species, demonstrating the lack of selective pressure outside of the splice junction and motif. | |||
[[Image:DASH.png|300px|thumb|left|DASH conservation: click for enlargement]] | |||
If a residue is displayed only when 21 or more of its 24 column mates agree (87.5% conservation), the figure at left -- which excludes the terminal exon -- shows the uneven distribution of the 276 best conserved residues. | |||
<br clear=all> | |||
>DASH_conSeq ultra-conserved consensus sequence residues for display on DASH_araTha 2VTB 53% identity | |||
MSMSSSRTV<FONT COLOR = RED>I</FONT>C<FONT COLOR = RED>L</FONT>L<FONT COLOR = RED>R</FONT>N<FONT COLOR = RED>DLR</FONT>LH<FONT COLOR = RED>DN</FONT>EVLHW<FONT COLOR = RED>A</FONT>QRNADHIV<FONT COLOR = RED>PLYCFDP</FONT>R<FONT COLOR = RED>H</FONT>YVG<FONT COLOR = RED>T</FONT>HHYNF<FONT COLOR = RED>PKTG</FONT>PH<FONT COLOR = RED>R</FONT>LR<FONT COLOR = RED>FLL</FONT>ESVK<FONT COLOR = RED>DLR</FONT>NT<FONT COLOR = RED>L</FONT>KKK<FONT COLOR = RED>GS</FONT>N<FONT COLOR = RED>L</FONT>VVRRGK<FONT COLOR = RED>PE</FONT>EVVAD<FONT COLOR = RED>L</FONT>IKQLGSVSAVAFHE<FONT COLOR = RED>E</FONT>VTK<FONT COLOR = RED>EE</FONT>LD<FONT COLOR = RED>VE</FONT>KALKQV<FONT COLOR = RED>C</FONT>AQHGVKVHTF<FONT COLOR = RED>WG</FONT>S<FONT COLOR = RED>TL</FONT>Y<FONT COLOR = RED>H</FONT>RD<FONT COLOR = RED>D</FONT>L<FONT COLOR = RED>PF</FONT>RHISRL<FONT COLOR = RED>P</FONT>DV<FONT COLOR = RED>YT</FONT>Q<FONT COLOR = RED>FRK</FONT>A<FONT COLOR = RED>VE</FONT>SQSR<FONT COLOR = RED>VR</FONT>PT | |||
FPM<FONT COLOR = RED>P</FONT>DQLKPLPPGLEE<FONT COLOR = RED>G</FONT>SI<FONT COLOR = RED>P</FONT>TAEDLGQKDPAT<FONT COLOR = RED>D</FONT>P<FONT COLOR = RED>R</FONT>SA<FONT COLOR = RED>FP</FONT>CS<FONT COLOR = RED>GGE</FONT>TQ<FONT COLOR = RED>A</FONT>LA<FONT COLOR = RED>R</FONT>LKH<FONT COLOR = RED>Y</FONT>F<FONT COLOR = RED>W</FONT>DTNLVAS<FONT COLOR = RED>YKETRNG</FONT>LI<FONT COLOR = RED>G</FONT>M<FONT COLOR = RED>DYSTKF</FONT>SP<FONT COLOR = RED>WLA</FONT>L<FONT COLOR = RED>G</FONT>CI<FONT COLOR = RED>SPR</FONT>Y<FONT COLOR = RED>IY</FONT>EQIKK<FONT COLOR = RED>YE</FONT>K<FONT COLOR = RED>ER</FONT>TA<FONT COLOR = RED>N</FONT>Q<FONT COLOR = RED>STYWV</FONT>I<FONT COLOR = RED>FEL</FONT>L<FONT COLOR = RED>WRDYF</FONT>R<FONT COLOR = RED>FV</FONT>AL<FONT COLOR = RED>K</FONT>Y<FONT COLOR = RED>G</FONT>NRI<FONT COLOR = RED>F</FONT>YLR<FONT COLOR = RED>G</FONT>LQDKSVP<FONT COLOR = RED>W</FONT>KKDMKL<FONT COLOR = RED>F</FONT>DA<FONT COLOR = RED>W</FONT>KE<FONT COLOR = RED>G</FONT>R<FONT COLOR = RED>TGVPFVDANMREL</FONT>AA<FONT COLOR = RED>TG</FONT>F<FONT COLOR = RED>MS</FONT> | |||
<FONT COLOR = RED>NRGRQNVASFL</FONT>T<FONT COLOR = RED>KDL</FONT>GL<FONT COLOR = RED>DWR</FONT>M<FONT COLOR = RED>GAEWFE</FONT>YL<FONT COLOR = RED>L</FONT>V<FONT COLOR = RED>D</FONT>H<FONT COLOR = RED>DVCSNYGNW</FONT>L<FONT COLOR = RED>Y</FONT>S<FONT COLOR = RED>AG</FONT>I<FONT COLOR = RED>GNDPR</FONT>EN<FONT COLOR = RED>RKFN</FONT>MI<FONT COLOR = RED>KQ</FONT>GL<FONT COLOR = RED>DYD</FONT>GN<FONT COLOR = RED>G</FONT>D<FONT COLOR = RED>Y</FONT>V<FONT COLOR = RED>R</FONT>L<FONT COLOR = RED>W</FONT>V<FONT COLOR = RED>PEL</FONT>QGIKGGDV<FONT COLOR = RED>H</FONT>T<FONT COLOR = RED>PW</FONT>T<FONT COLOR = RED>L</FONT>SSAA<FONT COLOR = RED>L</FONT>SQAGVS<FONT COLOR = RED>LG</FONT>ET<FONT COLOR = RED>YP</FONT>N<FONT COLOR = RED>P</FONT>IVT<FONT COLOR = RED>A</FONT>P<FONT COLOR = RED>EW</FONT>S<FONT COLOR = RED>RH</FONT>INKKPSGSGPSPRGRKGPSHTPKQHKD<FONT COLOR = RED>RGIDFYFS</FONT>RS<FONT COLOR = RED>K</FONT>LS | |||
<br clear=all> | |||
=== CPD === | === CPD === | ||
The 33 CPD sequences | The 33 CPD sequences are unalignably N-terminally (iMet wander) but are otherwise quite orderly in their conservation. The [http://genomewiki.ucsc.edu/index.php/Image:TrimmedGroomedCPD.doc groomed and gapped sequence set] from initial to final regions of conservation runs to 473 amino acids. The sequences are in phylogenetic order relative to marsupial. | ||
The first alignment is conventional (full sequences for all species), allowing regions of interest to be located by web browser text search. For example, Oryza sativa (rice) has a determined CPD structure (3UMV) marked up in its GenBank entry for alpha helices and beta sheets. Methanosarcina mazei (euryarchaeota) also has a relevant structure 2XRY that is still 49% identical to frog that | The first alignment is conventional (full sequences for all species), allowing regions of interest to be located by web browser text search. For example, Oryza sativa (rice) has a determined CPD structure (3UMV) marked up in its GenBank entry for alpha helices and beta sheets. Methanosarcina mazei (euryarchaeota) also has a relevant structure 2XRY that is still 49% identical to frog that is relevant for comparative purposes. | ||
[[File:CPDlogo.png|300px|thumb|left|Weblogo for CPD: click for full size]] | [[File:CPDlogo.png|300px|thumb|left|Weblogo for CPD: click for full size]] | ||
| Line 3,434: | Line 3,751: | ||
FC.Y...YD...G...WA...L..H..D.R...Y....LE...T.D.LWN..Q......GK.HGF.RMYWAKKILEWT..P...L.....LN.....DG.DP.GYVGC.WS..G.HD.GW.ER..FGK.R.MNY.GC.RKF | FC.Y...YD...G...WA...L..H..D.R...Y....LE...T.D.LWN..Q......GK.HGF.RMYWAKKILEWT..P...L.....LN.....DG.DP.GYVGC.WS..G.HD.GW.ER..FGK.R.MNY.GC.RKF | ||
>CPD_conSer ultra-conserved residues for display on CPD_orySat 3UMV 100% identity | |||
<font color=gray><font color=red>YWM</font>L<font color=red>RD</font>Q<font color=red>R</font>LA<font color=red>DNWA</font>LLHAAGL<font color=red>A</font>AASAS<font color=red>PL</font>AVA<font color=red>F</font>ALFPRL<font color=red>L</font>SARR<font color=red>R</font>QLG<font color=red>F</font>LLR<font color=red>GL</font>RRLAADAAARHLP<font color=red>F</font>FLFT<font color=red>G</font>GPAE-IPALVQRLGASTLVA<font color=red>D</font>FS<font color=red>PL</font>RPVREALDAVVGRREAPGVAVH<font color=red>QVDAHN</font>V<font color=red>VP</font>V<font color=red>W</font>TA<font color=red>S</font>A<font color=red>K</font>M<font color=red>E</font>YS | <font color=gray><font color=red>YWM</font>L<font color=red>RD</font>Q<font color=red>R</font>LA<font color=red>DNWA</font>LLHAAGL<font color=red>A</font>AASAS<font color=red>PL</font>AVA<font color=red>F</font>ALFPRL<font color=red>L</font>SARR<font color=red>R</font>QLG<font color=red>F</font>LLR<font color=red>GL</font>RRLAADAAARHLP<font color=red>F</font>FLFT<font color=red>G</font>GPAE-IPALVQRLGASTLVA<font color=red>D</font>FS<font color=red>PL</font>RPVREALDAVVGRREAPGVAVH<font color=red>QVDAHN</font>V<font color=red>VP</font>V<font color=red>W</font>TA<font color=red>S</font>A<font color=red>K</font>M<font color=red>E</font>YS | ||
<font color=red>A</font>K<font color=red>T</font>F<font color=red>R</font>G<font color=red>K</font>VSKVMDEY<font color=red>L</font>VE<font color=red>FP</font>ELPAVVPWDREQPEGLVD<font color=red>W</font>DALIARVCSEENVPEID<font color=red>W</font>CEP<font color=red>G</font>EEAAIEA<font color=red>L</font>DG<font color=red>F</font>LTK<font color=red>R</font>IKSYETD<font color=red>RN</font>D<font color=red>P</font>TKRAL<font color=red>S</font>G<font color=red>LSP</font>YL<font color=red>H</font>F<font color=red>G</font>HISA<font color=red>QR</font>CA<font color=red>L</font>EAKKCRHLSPKS<font color=red>V</font>DAFL<font color=red>EE</font>LVVR<font color=red>REL</font>AD<font color=red>N</font> | <font color=red>A</font>K<font color=red>T</font>F<font color=red>R</font>G<font color=red>K</font>VSKVMDEY<font color=red>L</font>VE<font color=red>FP</font>ELPAVVPWDREQPEGLVD<font color=red>W</font>DALIARVCSEENVPEID<font color=red>W</font>CEP<font color=red>G</font>EEAAIEA<font color=red>L</font>DG<font color=red>F</font>LTK<font color=red>R</font>IKSYETD<font color=red>RN</font>D<font color=red>P</font>TKRAL<font color=red>S</font>G<font color=red>LSP</font>YL<font color=red>H</font>F<font color=red>G</font>HISA<font color=red>QR</font>CA<font color=red>L</font>EAKKCRHLSPKS<font color=red>V</font>DAFL<font color=red>EE</font>LVVR<font color=red>REL</font>AD<font color=red>N</font> | ||
| Line 3,442: | Line 3,758: | ||
[[Image:CPDultras.gif|200px|thumb|left|Ultra conserved residues in CPD]] | [[Image:CPDultras.gif|200px|thumb|left|Ultra conserved residues in CPD]] | ||
When the ultra conserved residues defined by alignment of 33 eukaryotic sequences are | When the ultra conserved residues defined by alignment of 33 eukaryotic sequences are displayed (yellow, at left) on the crystallographic structure for rice CPD, a rather striking result emerges: the conserved residues predominantly cluster at the interface between the two domains and also about the catalytic FAD site and DNA binding groove. The structure and function of CPD cannot be said to be understood until an explanation is at hand for all observed conserved residues. Some of this conservation is mundane internal packing but other conserved surface residues are likely involved in protein-protein regulatory interactions while a conserved antenna pocket impies an antenna molecule, and yet other conserved aromatic residues are critical to the light sensing reaction or key to DNA binding and conformational change. | ||
Many different displays of conservation on structure are feasible, for example ultra-conserved tryptophans, a small subset of which presumably participate in electron flow. Those of CPD could be compared (homologically aligned) with those of other orthology classes. While snapshots have some value, an interactive 3D resource is ultimately much better. Whether a viewer can be embedded in this wiki for the diverse computer platforms of visitors needs further exploration. If so, then the focus can shift from static images into providing comparative genomics resources that properly initiate the interactive viewer. | |||
<br clear=all> | <br clear=all> | ||
== Article authorship and data usage policy == | == Article authorship and data usage policy == | ||
[[Image:Author.jpg|left]] | [[Image:Author.jpg|left]] | ||
I researched this article in its entirety in the winter of 2012, not paying attention | I researched this article in its entirety in the winter of 2012, not paying much attention to previous studies which are excellent on reaction mechanisms and regulatory cycles but completely clueless on comparative genomics ten years into that era. Cryptochromes are a moderately difficult topic as vertebrate genes go because the timing of gene duplications largely falls between the cracks of phylogenetic coverage and because 'model' organisms did not retain a representative set of orthologs. | ||
My interests are primarily in the long range evolutionary acquisition and divergence of function-enabling structure, starting from primases and 4Fe-4S cluster photolyases and ending with circadian, | I plan to greatly expand the treatment of 3D structural aspects of comparative genomics during the summer of 2012. My interests are primarily in the long range evolutionary acquisition and divergence of function-enabling structure, starting from primases and 4Fe-4S cluster photolyases and ending with circadian, magneto-sensing and couplings with opsins. However comparative genomics has major applications to rapid hypothesis testing in all aspects of cryptochrome and photolyase research, the main point being the strong coupling between sequence conservation (ie selective pressure) and functional importance. This means conservation never persists without a reason, and conversely that non-conserved features are not important. | ||
Although copyrighted, all the information here is in the public domain and can be used by anyone without additional permissions if properly sourced; however if data, figures or original observations are taken wholesale for a peer-reviewed scientific publication, it might be appropriate (after consultation early on) to include me among secondary co-authors. | Although copyrighted, all the information here is in the public domain and can be used by anyone without additional permissions if properly sourced; however if data, figures or original observations are taken wholesale for a peer-reviewed scientific publication, it might be appropriate (after consultation early on) to include me among secondary co-authors. | ||
| Line 3,486: | Line 3,774: | ||
Rather than make article edits yourself, please contact me by email with clarifications, corrections or additions to the content so I can make edits while maintaining a consistent approach. For broader disagreements or different interests, a better option is to simply register at the UCSC genomeWiki site and create your own page within the comparative genomics category. | Rather than make article edits yourself, please contact me by email with clarifications, corrections or additions to the content so I can make edits while maintaining a consistent approach. For broader disagreements or different interests, a better option is to simply register at the UCSC genomeWiki site and create your own page within the comparative genomics category. | ||
This is just a scientific research article on a vertebrate gene family, not a counseling resource for personal genomics, | This is just a scientific research article on a vertebrate gene family, not a counseling resource for personal genomics, recommendation for or against melatonin supplementation, or medical advice on insomnia and metabolic disorder -- thanks in advance for not sending inappropriate email. Technical terms from genetics and molecular biology are not explained in the article when keywords have a satisfactory treatment at wikipedia or in undergraduate genetics texts; because of good keywords, the scientific literature is easily searched at [http://www.ncbi.nlm.nih.gov/pubmed/advanced PubMed] so not duplicated here. | ||
My last dozen published research papers in PNAS, Nature, Science etc can be found [http://www.ncbi.nlm.nih.gov/pubmed/21709235,20164927,19020620,18464734,18266766,17984227,18085818,17975064,17322288,15608236,12045153,22012981 here]. Watch for 4 additional comparative genomics paper to appear in 2012. I've also written over a [[:Category:Comparative_Genomics|thousand pages of comparative genomics]] for other human genes, authored the original user manual to the UCSC human genome browser and in 1999 an advanced tutorial on metazoan genome annotation still [http://www.mad-cow.org/00/annotation_tutorial.html widely available online]. I thank the UCSC Genomics Group (Hiram Clawson, Brian Raney, Maximilian Haeussler) for software, manuscript and literature resources, Evim Foundation for logistical support, and the Sperling Foundation for financial support under project grant 2012.GNTCS.006. | My last dozen published research papers in PNAS, Nature, Science etc can be found [http://www.ncbi.nlm.nih.gov/pubmed/21709235,20164927,19020620,18464734,18266766,17984227,18085818,17975064,17322288,15608236,12045153,22012981 here]. Watch for 4 additional comparative genomics paper to appear in 2012. I've also written over a [[:Category:Comparative_Genomics|thousand pages of comparative genomics]] for other human genes, authored the original user manual to the UCSC human genome browser and in 1999 an advanced tutorial on metazoan genome annotation still [http://www.mad-cow.org/00/annotation_tutorial.html widely available online]. I thank the UCSC Genomics Group (Hiram Clawson, Brian Raney, Maximilian Haeussler) for software, manuscript and literature resources, Evim Foundation for logistical support, and the Sperling Foundation for financial support under project grant 2012.GNTCS.006. | ||
Latest revision as of 00:01, 30 August 2018
See also: Comparative genomics analysis of cryptochromes and photolyases
Updates: fixes and additions become difficult to locate in a long article; these are provided below in reverse chronological order linked to their approximate spot in the article. 20 May 12: added 9 new DASH sequences, fixed various sequence discrepancies and provided a groomed sequence alignment. 18 May 12: scoured ctenophore genome for photolyases and cryptochromes, finding only CRY64 and CPD. 10 Oct 13: intensive search of new amniote genomes for complete gene family sets
Introduction: curated metazoan cryptochrome and photolyase sequences
The 307 metazoan cryptochrome and photolyase reference sequences provided here are hand-curated for greater accuracy. This is necessary in part because many full length genes are implicit in genome projects but not provided as such at GenBank. When predicted by unsupervised bioinformatic algorithms, the resulting gene models too often are unacceptably inaccurate -- erroneous start codons, missing short and diverged exons, bogus splice junctions in gappy regions, uncorrected homopolymer run frameshift errors, read-through of penultimate exons to the next random stop codon, and making no use of massive and synergistic homological data.
Comparative genomic analysis of a gene family cannot be built upon a rotten foundation of faulty sequences -- this can only lead to downstream rubbish. The method of curation used here is complex but necessary. It begins with a seed set of thoroughly studied experimental sequences in model species where there is no doubt about completeness of the sequence nor its accuracy. Using the four main divisions of GenBank (nr, ESTs, transcriptome projects, whole genome assemblies), the seed set of each orthology class is slowly expanded with additional representatives in closely related species that share homology, syntenic location and exon pattern.
The build-out of the reference sequence collection grows recursively in accuracy because of four independent tools: an ever-growing blast classifier, a phylogenetically aware protein multi-aligner, a pre-computed best-blast phylogenetic overview of neighboring genes, and a BROKEN 46-species whole genome alignment based gene predictions.
The blast classifier allows homologs extracted from raw dna contigs and 46-way preliminary models to be assigned to the closest available reference sequence class. The multi-aligner highlights anomalies in the sequence collection that required additional curational focus (such as incorrect exon boundaries and regions temporarily out of reading phase). The synteny browser is often correct even when its underlying gene models are somewhat flawed and is imperative to determining orthology. However synteny dissipates fairly rapidly with phylogenetic distance, so the much deeper conservation of intron position and phase becomes critical to refining gene models and orthologous classification.
Despite these improvements, a set of sequences is never perfect because of errors in GenBank data (sequencing lab contamination, systemic errors in read technology, mis-assembled contigs, gaps in coverage, premature truncation of contigs, high levels of polymorphism, unsuspected hybrids, parasites and commensals, erroneous taxonomy, variations of the single animal sequenced unrepresentative of its species, lineage sorting during a messy speciation, horizontal gene transfer, and inevitable data handling errors.
Contaminated DNA is a significant problem in GenBank entries. This can arise somewhat excusably from embedded viruses, endosymbionts, commensals or parasites not being cleanly separated from target organism, but less excusably from DNA contamination by species previously sequenced in the same lab (eg, DNA from the Lottia genome project contaminated sponge, the subsequent project at JGI). It is not unusual to find tuna sandwich, chicken salad and lab staff DNA in older mammalian genome projects. Horizontal gene transfer must also be considered when a sequence does not fit into its phylogenetic niche, though that is a non-issue in vertebrates. Some clades, such as tunicates and nematodes, just evolve so rapidly that their sequences seem implausible; even when confirmable, these are not informative to the comparative genomics endeavor.
The focus here is entirely on metazoan sequences -- sponge to human -- supplemented slightly with homologs for intensively studied model species such as Arabidopsis, single-celled eukaryotes when an outgroup is helpful, and prokaryotic species where the fold has been determined. The instructive class of 4Fe-4S photolyases occurs only in prokaryotes so an exception was made there; a few primases are also included because of deep fold homology to photolyases.
Because of multiple events of gene duplication, followed by accelerated asymmetric divergence and sometimes much later loss in various lineages, orthologous correspondences are not so straightforward in this gene family. Because it is not feasible to experimentally study every protein in every species, it is imperative to correctly establish orthologous families to optimize the accuracy of annotation transfer. That gives far more accurate outcomes than across paralogs -- and when it doesn't, leads to discovery of unsuspected developments in protein evolution.
By definition, two genes in two species are orthologous only when they are vertically descended from a single gene in their last common ancestor. Orthology is strictly genetic -- its definition makes no reference to function but instead describes evolutionary descent. It cannot be reliably determined by reciprocal best blast. Classification by non-heritable and possibly revertible attributes such as enzyme vs signaling, presence or absence of antenna molecule, ssDNA or dsDNA substrate, peak wavelength adsorption and so forth are not productive.
Orthologs can and do drift in exon number and position, cofactors, post-translational modifications, tissue and intra-cellular location of expression, regulation of that transcription, protein binding partners, enzymatic substrates and products, and relative catalytic efficiency. However, mostly they don't -- genes can persist for trillions of years of cumulative branch length time with very few changes in any attribute. Annotation transfer to orthologs provides a default hypothesis that can only be refined by actual experiment.
The five main tools for determining orthology are primary and tertiary sequence relatedness, location and size of indels (insertions and deletions), positions and phases of introns, and retention of syntenic chromosomal position. These tools are independent of each other and so synergistic; any model of orthology relationships must be compatible with and fully utilize the information of all five. Lazy methods such as uncurated downloads, best reciprocal blast and evolution-modeling software can only provide crude provisional outcomes.
Previous efforts in the comparative genomics of cryptochromes are not reviewed here. These have floundered on the twin shoals of overly sparse phylogenetic sampling relative to gene duplication event rate and over-reliance on modeling tools based on simplifying but unwarranted assumptions about clade-specific variations in the tempo and mode of nucleotide evolution over a vast evolutionary time span involved. Unpretentious blast clustering in the classifier accomplishes the same thing with far greater speed and transparency.
Since today the phylogenetic tree is largely geologically dated and topologically fixed, individual gene trees -- which are commonly discrepant -- can be clamped (subordinated) to the species tree even if that is less parsimonious. After a gene duplication, if one copy retains parental location and main role, the other copy may re-functionalize and evolve rapidly as it adapts better to its newly specialized role. This scenario can account for anomalies in gene trees and is likely applicable here to the shift from repair enzyme to light sensor.
In the view of Piatigorsky, parental genes are usually multi-functional to begin with, so the duplicated copy already has a pre-existing niche available to take over -- it does not need to scramble desperately for a function before pseudogenization from lack of selective pressure support. All fixed gene duplications of cryptochromes within Bilateria have been segmental, meaning upstream regulatory and enhancer regions were likely carried along, which is not the case with retropositioned gene copies which duplicate only a matured mRNA transcript and not its promoter regions.
The sequence collection has been split from a companion analysis page to separate data from analysis. Not every conceivable cryptochrome gene fragment at GenBank is represented below; the intensity of curational effort increases in proportion to that needed to resolve specific evolutionary issues. For example, vertebrate CRY1 genes evolve very slowly in their main region but acquired additional terminal exons; the issue there is where those originated, what parts of them are conserved, and what they are doing. Here a terminal gene fragment from a phylogenetically critical position is more informative than infilling static regions of the tree with intermediate species.
The sequences are provided in advanced fasta format -- parsed into exons, preceded by splice acceptor overhang, followed by splice donor coding phase overhang, with lower case amino acids indicating uncertainty, spaces indicating gaps in coverage, and stop codons shown when explicitly in the data. Frameshifts in homopolymer runs -- a common form of sequencing error -- have been edited to restore reading frame.
The sequences are grouped as orthologs, re-named as necessary to avoid confusion with historical use, and presented in phylogenetic order relative to human. This does not determine order in subtrees (eg rodents) but there the order can be set by assembly quality (eg mouse is the best murid assembly.
The fasta header lines are simple space-delimited databases showing first gene name, then genus, species, common name, accession number if not a simple genomic blat or whole genome alignment output, PubMed accession if specifically studied in a journal article, followed by an unstructured comment field. The headers and exons are easily reformatted into a spreadsheet single lines by replacing spaces and paragraph returns with tabs.
Here gene names are designed for consistency across all Eukaryota, with special attention to phylogenetically localized gene duplications. Following international agreement, no greek letters, roman numerals, hyphenation, lower case letters, subscripts, or superscripts can be used in gene names. It is completely unworkable with 10,000 vertebrate genomes to use narrowly conceived lab conventions such as mCRY1 for mouse -- even 6 letters for genusSpecies can provide no more than a mnemonic. Since annotation today is done by gene family, the gene name comes first to preserve orthologous clustering when listed alphabetically, which also provides clarity in alignment tool output.
The availability of some orthology classes is quite limited due to recent origin and restricted phylogenetic persistence. Sequencing effort is unfortunately spread very unevenly across the phylogenetic tree of metazoans. For species with good assemblies, the entire repertoire of cryptochromes and photolyases can be extracted. For large genes with numerous exons, absence from the assembly usually means genuine absence from the genome. When only an exon or two gene fragment is available, the classifier can still usually assign the correct orthology class. However it is risky to assemble an entire gene from small unlinked contigs and that was not done here with the exception of critical clades such as cartilaginous fish that lack coherent assemblies and comprehensive transcript collections. It sometimes suffices just to establish the presence/absence of a given gene at a given time of divergence.
A remarkable amount of the data below surfaced at GenBank during the last six months, implying much better phylogenetic coverage will surface during 2012-14. This promises to resolve the timing of gene duplications to the extent that extant species can determine it. The mysterious role of 4Fe-4S clusters in photolyases, primases, helicases and other DNA repair enzymes may also be clarified experimentally, providing a better foundation to understanding the deeper evolutionary origin of photolyases and cryptochromes.
In terms of massive gene losses in this family supporting the recently revisited 1942 Wall hypothesis concerning a deep nocternal epoch for mammals, the record is still somewhat spotty in the critical region of early tetrapods. Amphibians other than frogs remain very poorly represented at GenBank. Only four photolyases/cryptochromes can be confirmed as of Oct 13, all transcripts from a single species of salamader. Only only one of these -- DASH -- is full length.
Thus it remains difficult to determine the full crytochrome/photolyase repertoire at the various amniote ancestral nodes. The Wall hypothesis has narrower versions, such as predicting losses in say, coelocanths, which live year-round in an extremely dark deep-sea environment and so might also be expected to have lost most genes in this family. Recently the genome of a cave fish has become available -- it too will likely have lost its crytochrome/photolyases as well as most opsins (to the extent these have no light-independent functions), depending on how many millions of years it has been out of genetic contact with surface fish.
An exonic pseudogene must be fairly new (several million years) to still be recognizable so indicates not only contemporary gene loss but that the species possessed an operational gene in the recent past. Actually, the same is true for a processed (intronless) pseudogene because it implies the existence (possibly continuing but just not in the current assembly) of a parent gene that gave rise to it. The difference is that processed pseudp\ogenes are far more persistent in terms of Blast dectectability because the longer length of their contiguous (formerly) coding region.
For example, DASH consists of 524 amino acids located in 14 widely separated exons with lengths varying from 20 to 59 residues (average of 37). As frameshifts from indels and substitutions from nucleotide changes accrue over time, these exons become increasingly difficult for Blast to confidently identify, a situation exacerbated by the best probe itself (necessarily from another species, potentially only ~85% identical) and potential mis-attribution to other members of the gene family and their pseudogenes.
Vertebrate CRY1 reference sequences
>CRY1_homSap Homo sapiens (human) 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSAENIPGCSSSG 1 2 SCSQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSMGTGLSGGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_panTro Pan troglodytes (chimpanzee) XM_509339 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSAENIPGCSSSG 1 2 SCSQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSVGTGLSGGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_ponAbe Pongo abelii (orangutan) XM_002823690 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDASLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSAENVPGCSSSG 1 2 SCSQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSMGTGLSGGKRASQEEDTQSIGPKVQRQSTN* 0 >CRY1_nomLeu Nomascus leucogenys (gibbon) XM_003269977 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSAENIPGCSSSG 1 2 SCSQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSMGTGLSGGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_macMul Macaca mulatta (rhesus) NM_001194159 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGINYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSTENIPGCSSSGS 1 2 CSQGSGILHYTHGDSQQTHLLKQ 1 2 GRSSMGTGLSGGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_calJac Callithrix jacchus (marmoset) XM_002752946 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSAENIPGCTSSG 1 2 SCSQGSGILHCAHGDSQQTHLLKQ 1 2 GRSSMSTGISGGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_saiBol Saimiri boliviensis (squirrel_monkey) nearly identical to marmoset 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSAENIPGCTSSG 1 2 SCSQGSGILHCAHGDSQQTHLLKQ 1 2 GRSSMSTGLGGGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_tarSyr Tarsius syrichta (tarsier) ABRT010205577 unsure if exon 2 is CRY1 or CRY2 0 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 0 0 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMGYSPAENTPGCSSSG 1 2 SCSQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSVGTGLSGGKRPSQEEDPQSIGPKVQRQSTN* 0 >CRY1_micMur Microcebus murinus (mouse_lemur) 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 IIELNGGQPPLTYKRFQTLISK SEVIEKCTTPLSDD 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 0 0 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGFMEYSPAENIPGCSSSG 1 2 NCSQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSMGTGFSGGKRPSQEEDTQSIGSKVQRQSTN* 0 >CRY1_otoGar Otolemur garnettii (bushbaby) AAQR03016495 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITRLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 0 0 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEIIEKCTTPLFDDHDEKYGVPSLEEL 1 2 0 2 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPASFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGINYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGSFMEYSPPENIPGCSSSG 1 2 NCSQGSGILHYAPGDGQQPHLLKQ 1 2 GRSSMGTGLSGGKRPSQEEDMQSVGPKVQRQSTN* 0 >CRY1_tupBel Tupaia belangeri (treeshrew) 0 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 0 0 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKFGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRVNSNSLLASPTGL 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLSSSGSCSQGSGILHYAHGDSQQTHLLKQGRSSMGP 1 2 1 2 GCSQGSGILHYAHGDSQQTHLLKQ 1 2 GLSSGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_musMus Mus musculus (mouse) NM_007771 all transcripts support longer exon 10 lost splice donor 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLVSKMEPLEMPADTITSDVIGKCMTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLER 0 0 KAWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNSNGNGGLMGYAPGENVPSCSSSGNGGLMGYAPGENVPSCSGG 1 2 NCSQGSGILHYAHGDSQQTHSLKQ 1 2 GRSSAGTGLSSGKRPSQEEDAQSVGPKVQRQSSN* 0 >CRY1_ratNor Rattus norvegicus (rat) NM_198750 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLVSKMEPLEMPADTITSDVIGKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYAPGENVPSGGSGGG 1 2 NCSQGSGILHYAHGDSQQTNPLKQ 1 2 GRSSMGTGLSSGKRPSQEEDAQSVGPKVQRQSSN* 0 >CRY1_criGri Cricetulus griseus (hamster) XM_003505292 0 MGVNAVHWFRKglrlhDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDSNLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLVSKMEPLEMPAETITSDVIGKCMTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLER 0 0 KAWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYTTGENLPSCSGGG 1 2 SCSQGSGILHYAHGDSQQAHLLKQ 1 2 GRSSMGTSLSSGKRPSQEEETRSVDPKVQRQSSN* 0 >CRY1_spaJud Spalax judaei (blind_molerat) AJ606298 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRW 2 1 RFLLQCLEDLDANLRKLNSRLLVIRGQPADVFPRLFKEWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERR 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTGSDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYTPGENIPNCSSSG 1 2 SCSQGSGILHYAHGDSQQAHLLKQ 1 2 GSSSMGHGLSNGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_dipOrd Dipodomys ordii (kangaroo_rat) ABRO01202522 ABRO01202521 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRW 2 1 RFLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCSVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYAAGDNLPGSSSSG 1 2 SCSQGSGILHYAHGDSQQMHLLKQ 1 2 GRSSMGTGLSSGKRPSQEEDSQSIGPKVQRQSTN* 0 >CRY1_hetGla Heterocephalus glaber (naked_molerat) AHKG01030319 revised hystricognath 0 MAVNAVHWFRKGLRLHDNPALKECTRGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCVTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYAPGESIPGGSSSG 1 2 SCAHGSGILPCAHTDGQQAHLLKP 1 2 GRNCVGPVLSSGKRPSQEEDAQSIGPKLQRQSTD* 0 >CRY1_cavPor Cavia porcellus (guinea_pig) AAKN02054126 last two exons diverged, only 69 bp separation 0 MGVNAVHWFRKGLRLHDNPALKECIRGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVVEKCVTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVHHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLLGYAPGESTPGSGGG 1 2 SCVPGSSSAGVSHCAQGEAPQAPP 1 2 GRDPAGPGLGGGKRPSQEEDAQSTGHKIQRQSPD* 0 >CRY1_octDeg Octodon degus (degu) AJSA01012260 AJSA01012259 hystricognath 0 MGVNAVHWFRKGLRLHDNPALKECIRGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCMTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQLYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLLGYAAGETLPGGGGGG 1 2 SCTPGSSILHCAHQDGQQAPLLKP 1 2 GRSSGGPAPGSGKRPSQEEDTQSIGHKVQRQSTT* 0 >CRY1_speTri Spermophilus tridecemlineatus (squirrel) AGTP01024540 AGTP01024543 Ictidomys 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHEASLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMAyAPGENIPGCSSSG 1 2 SCTQGSSILHNAHGDSQQTHLLKQ 1 2 GRSSMGTGLSSGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_oryCun Oryctolagus cuniculus (rabbit) 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRW 2 1 RFLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITRLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSDVIEKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGQTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGINYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYSPGENIPGCSSSG 1 2 SCSQGSGILHYAQGDTQQTQLLKQ 1 2 GRSSMGTGLSSGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_oviAri Ovis aries (sheep) NM_001129735 19341811 19150926 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIIRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVMEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYSPGENIPGCSSSA 1 2 SCTQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSTAAGLGSGKRPSQEEDTQSVGPKVQRQSTN* 0 >CRY1_bosTau Bos taurus (cow) NM_001105415 XM_616063 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYSPGENIPGCSSNA 1 2 SCTQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSTGAGLGSGKRPSQEEDTQSIGPKVQRQSTN* 0 >CRY1_susScr Sus scrofa (pig) XM_003126079 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYSPGENIPGCSSSG 1 2 SCPQGSGILHYAHGESQQNHLLKQ 1 2 GRSSTGSGLSSAKRPSQEEDTQSIIGPKVQRQSTN* 0 >CRY1_ailMel Ailuropoda melanoleuca (panda) XM_002927658 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPANPNGNGGLMGYSPGENIPGCSSSG 1 2 SCSQGSGILHYAHGDSQQTHLLKQ 1 2 GRSSMGSGLSSGKRPSEEEDTQSIGPKVQRQSTN* 0 >CRY1_loxAfr Loxodonta africana (elephant) XM_003405313 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWDITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVIGKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYSPGENTPGCNSSG 1 2 SCSQGSGILHYVHGDSLLKQ 1 2 GRSPTGTGVSSGKRPSQDEETQTLGPKVQRQSTN* 0 >CRY1_triMan Trichechus manatus (manatee) AHIN01036366 AHIN01036362 very similar to elephant 0 MGVNAVHWFRKGLRLHDNPALKECIQGADTIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWDITKLSIEYDSEPFGKERDAAIKKLATEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLEIPVETITSEVMEKCTTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCSIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYSPGENIPGCSSNG 1 2 SCPQGNGILHYAHRDSQQAHLLKQ 1 2 GRSPTGTGVSSGKRPSQEEETQSIGPKVQRQSAN* 0 >CRY1_monDom Monodelphis domestica (opossum) XM_003341966 0 MGVNAGHWFRKRLRLPHNPALKGCIQGADTVCCVYIRDPWFGGSSNFGANEWK 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMEPLATPVETITPEVMHKCVTPLSDEHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGSLMAYTPGENIPGCSSGG 1 2 GAPVGASDGQILQACVLPEPPTGTSGVQQP 1 2 GYSQGSGISHYSHEDSQQAYMLKQ 1 2 GRSSLGVGGGKRPRQEEETQSINPKVQRQSTN* 0 >CRY1_macEug Macropus eugenii (wallaby) assembly frameshift 0 MGVNAVHWFRKGLRLHDNPALKECIEGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPLGRNAMQLLKNWASEAGVEVIVRISHTLYDLDK 2 1 1 2 gFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGVQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGSLMGYTTGENIPTCSSSGG 1 2 GAPAGASDGQILQACVLPEPPTGTSGVQQP 1 2 GYSQGSGISHYSHEDSQQAYVLKQ 1 2 GRNSLGGGKRHRQEEETQSIGSKMQRQSVN* 0 >CRY1_sarHar Sarcophilus harrisii (tasmanian_devil) nearly identical to oppossum 0 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSINAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKVAKCLIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYTSGENGPACNSGG 1 2 GAPVGASDGQILQSCALPEPPAGASCIQQS 1 2 GYSQGSGISHYSHEDSQQAYILKQ 1 2 GRSSLSGGKRPRQEEETQSVGPKVQRQSVN* 0 >CRY1_triVul Trichosurus vulpecula (possum) EC362500 terminal transcript 1 IERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGGLMGYAPGENIPACSSSGG 1 2 GAPAGVGDGQILQACALPEPPTGASGVQQP 1 2 GYSQGSGISHYAHEDSQQAYMLKQ 1 2 GRSSLSGGGKRHRQEEEAQSIGPKMQRQSVN* 0 >CRY1_ornAna Ornithorhynchus anatinus (platypus) XM_001508563 = rubbish, genomic frameshift, continuing exon 12 0 MGVNAVHWFRKGLRLHDNPALRDCVRGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYELDK 2 1 IIELNGGQPPLTYKRFQTLISRMDPLAMPVETITAEVMGKCMTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATSNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNANGSGGLMAYSPGENIPGCSSGGG 1 2 GVQMGASESH LLQTCVLGESHLGPSGIQQQ 1 2 GYCQGSGVLYYANGESHLTQ 1 2 GRSSLTPGLSGGKRPCQEEESQSIGPKVQRQSTD* 0 >CRY1_tacAcu Tachyglossus aculeatus (echidna) SRR000649.130490 short read transcripts corrected for frameshifts, skipped penultimate exon 0 2 1 0 0 2 1 ETITAEVMGKCVTPLSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGEAEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLTPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATSNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAImtqlrqeGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGIQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNANGSGGLMAYSPGENIPGCSSGG 1 2 GAQIGASESHLLQTCVLGESHLGPSGIQQQ 1 2 1 2 GRSSLTPGLSGGKRHCQEEESQSIGPKVQRQSTD* 0 >CRY1_galGal Gallus gallus (chicken) PMID: 11684328,17324421,15459395 altSplExon11: GIVGVPICRGSADLCN* BU143111 0 MGVNAVHWFRKGLRLHDNPALRECIRGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWSIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITPEVMQKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERH 0 0 LERKAWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESVQKAAKCVIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGNGGLMSFSPGESISGCSSAG 1 2 GAQLGTGDGQTVGVQTCALADSHTGGSGVQQQ 1 2 GYCQASSILRYAHGDNQQSHLMQP 1 2 GRASLGTGISAGKRPNPEEETQSVGPKVQRQSTN* 0 >CRY1_melGal Meleagris gallopavo (turkey) XM_003202363 altSplExon11: GTVGVPICRGSANWYK* 0 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWSIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITPEVMQKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERH 0 0 LERKAWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESVQKAAKCVIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGNGGLMSFSPGESISGCSSAG 1 2 GAQLGTGDGQTVGVQSCALGDSHTGGNGVQQQ 1 2 GYCQASSILRYAHGDNQQPHLMQP 1 2 GRASLGTGISAGKRPNPEEETQSVGPKVQRQSTN* 0 >CRY1_anaPla Anas platyrhynchos (duck) scaffold157 altSplExon11: GMTGVLVCRGSPGSHNYGKKDKT* 0 MGVNAVHWFRKGLRLHDNPALRECIRGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITPEVMEKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESIQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGSGGLMGFSPGESVSGCGSTG 1 2 GAQLGTGDSQTVGVQTCALGDSHAGTSGIQQQ 1 2 GYCQASSILHYAHGDNQQSHLLQA 1 2 GRTSLGTGISAGKRPNPEEETQSVGPKVQRQSTN* 0 >CRY1_eriRub Erithacus rubecula (robin) AY585716 aka: CRY1A altSplExon11: GIMAVPVCRGSPNACNYGKPDKTSK* CRY1B 0 MGVNAVHWFRKGLRLHDNPALRECIRGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITPEVMKKCTTPVFDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 ASVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESIQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGNGGLMGYSPGESISGCGSTG 1 2 GAQLGTGDGHTVVQSCTLGDSHSGTSGIQQQ 1 2 GYCQASSILHYAHGDNQQSHLLQA 1 2 GRTALGTGISAGKRPNPEEETQSVGPKVQRQSTN* 0 >CRY1_sylBor Sylvia borin (warbler) AJ632120 aka: CRY1A PMID:15381765 altSplExon11: GIVAVAVCRGSPNPCNYGKPDKTSE* sylBor DQ838738 CRY1B 0 MGVNAVHWFRKGLRLHDNPALRECIQGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITPEVMKKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESIQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGNGGLMGYSPGESISGCGSTG 1 2 GAQLGAGDGHSVVQSCALGDSHTGTSGVQQQ 1 3 GYCQASSILHYAHGDNQQSHLLQA 1 2 GRTALGTGISAGKRPNPEEETQSVGPKVQRQSTN* 0 >CRY1_taeGut Taeniopygia guttata (finch) XM_002196518 altSplExon11: GIMAVPVCRGSPNPCNYRKPDKTSK* 0 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 0 0 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITPEVMKKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESIQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGNGGLMGYSPGESISGCGSTG 1 2 GAQLGTGDGHSVVQSCALGDSHTGTSGIQQQ 1 2 GYCQASSILHYAHGDNQQSHLLQA 1 2 GRTALGTGISAGKRPNPEEETQSVGPKVQRQSTN* 0 >CRY1_melUnd Melopsittacus undulatus (parakeet) AGAI01062111 altSplExon11: GIMAVPVCRGSSNPCNCGKTDKTSK* 0 MGVNAVHWFRKGLRLHDNPALRECIQGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITPEVMEKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESIQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGNGGLMGYSPGESISGCSSAGG 1 2 GAQLGTGDGHTVVQPCALGDSHTAGASGIQQQ 1 2 GYCQASSILHYAHGDNQQSHLLQAG1 2 VGTGISAGKRPNPEEETQSVGPKVQRQSTN* 0 >CRY1_parWeb Paradoxornis webbianus (parrotbill) JR867166 TSA transcript 0 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITPEVMKKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESIQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGNGGLMGYSPGESISGCGSTG 1 2 GAQLGTGDGHSVVQSCALGDSHTGTSGIQQQ 1 2 GYCQASSILHYAHGDNQQSHLLQA 1 2 GRTALGTGISAGKRPNPEEETQSVGPKVQRQSTN* 0 >CRY1_allMis Alligator mississippiensis (alligator) genome/blat 0 MGVNAVHWFRKGLRLHDNPALRECIQGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMKKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGVQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNPNGNGNGGLMGYSPGENVSGCGSTG 1 2 GAQMGSSDGHTVSVQPCALGESHGGSNGIQQQ 1 2 GYFQASSILHFPHGDDQQSHLLQQ 1 2 GRTSLSSGISAGKRPNPEEETQSIGPKVQRQSTN* 0 >CRY1_croAda Crotalus adamanteus (rattlesnake) JU174244 TSA 0 MGVNSVHWFRKGLRLHDNPALREVIEGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNISKLSIEYDSEPFGKERDAAIKKLATEAGLEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMSTCITPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGVK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPILRGFPAKYIYDPWNAPEGVQKAAKCTIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNGNGNGNGGLMGYSPSENLPGCVGTN 1 2 GTQMGTSEGHTGNVQACTLGETHTGTSGIHQQ 1 2 GYCQGNSGILHYAHGDSKKTHLMQQ 1 2 GRTSLSIGGCTGKRPNPEEGIQSIGPKVQRQSSN* 0 >CRY1_pytMol Python molurus (python) AEQU010547455 0 MGVNSVHWFRKGLRLHDNPALREVIQGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFk 0 0 EWNISKLSIEYDSEPFGKERDAAIKKLATEAGLEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMSKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLER 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPILRGFPAKYIYDPWNAPEGVQKAAKCIIGINYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 2 1 2 GAQMGTSEGHTGNVQACTLGETHTGTSGIQQQ 1 2 GYSQGNSGILHYAHGDSQKTLLMQ 1 2 GRTSLSVGVCTGKRPNPEEGIQSIGPKVQRQSSN* 0 >CRY1_chrPic Chrysemys picta (turtle) AHGY01469963 AHGY01469969 0 MGVNAVHWFRKGLRLHDNPALRECIQGADSVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNIAKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMEKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNRSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVSFGRRTDPNGDYIR 2 1 RYLPILRGFPAKYIYDPWNAPESIQKAAKCVIGVHYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLATVPSNP NGNGGLMGYSPGENISGCSSAS 1 2 GAQMGSNDGHTVGVQTCSLEDSHAGSSGIQQH 1 2 GYSQGNSIVHYAQGDHQQSHLLQQG 1 2 GRTVSTGISTGKRPNPEKETQSIGPKVQRQSTN* 0 >CRY1_anoCar Anolis carolinensis (lizard) XM_003220923 AAWZ02014443 0 MGVNAVHWFRKGLRLHDNPALREVIQGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYELDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEIPVETITAEVMSKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTDGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPDGVQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQMYQQLSRYRGL 1 2 GLLASVPSNGNGNGNGGLMGYSTGENIPGCTNTN 1 2 GSQMGMNEGHIGNVQACTMGESHTGTSGIQQQ 1 2 GYSQGSGILLYSHGDNQKTHSAQKGRISLGT 1 2 GVCTGKRPSPEVETQSVGPKVQRQSSN* 0 >CRY1_podSic Podarcis siculus (wall_lizard) DQ376040 16809482 0 MGVNAVHWVRQGLRLPDNRALREVIQGADTARCVYILDPSFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMEPLEMPVETITAEVMSKCTTPVSDDHDEKCGVPSLEEL 1 2 GFDTGGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARRAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGVQKVAKCVIGVNYPKPMVNHTEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPLMEWNVIVLMGYHQGTLLVYHAS 1 2 GSQMGTNEAHTGSVQTCTLGESHTGTSGIQQQ 1 2 GYPQGSDILHYAHGEGQKTHLIQQGRASLVA 1 2 GVCTGKRPNPEEETQSIGPKVQRQSSK* 0 >CRY1_xenTro Xenopus tropicalis (frog) NM_001087660 11533577 final four exons confirmed by many ESTs 0 MGVNAVHWFRKGLRLHDNPALRECIQGADTVRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWKITKLSIEYDSEPFGKERDAAIKKLASEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISKMDPLEIPVETITAEVMEKCTTPVSDDHDEKYGVPSLEEL 1 2 GFDTEGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASTTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMDGNPICVQIPWDRNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGKRTDPNGDYIR 2 1 RYLPILKGFPPKYIYDPWNAPETVQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNPNGNGNGGLMSYSPGESMSGCSNNG 1 2 GGQMGVNEGSSASNPNANKGEVHPGTSGLQ 1 2 GYWQGSSILHYSHSDSQQSYLMQ 1 2 ARNPLHSVVSSGKRPNPEEETQSIGPKVQRQSSH* 0 >CRY1_ambMex Ambystoma mexicanum (axolotl) JV201332 fragment 154-312 QTLISRMEPLEMPVETIGPEVMEKCTTPVLDDHDDKYGVPSLEELGFDTEGLPSAVWPGGE TEALTRLERHLERKAWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK VKKNSLPPLSLYGQLLWREFFYTAATNNPRFDKMEGN >CRY1A_latCha Latimeria chalumnae (coelocanth) AFYH01018055 AFYH01018053 AFYH01018050 0 MGVNAIHWFRKGLRLHDNPALREAIQGGDCIRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNISRLSFEYDSEPFGKERDAAIKKLASEAGVEVIVRITHTLYDLDR 2 1 IIELNGGQPPLTYKRFQTLISRMEPLDLPVETINSEVMGNCSTPVSDDHDEKYGVPSLEEL 1 2 GFDTEGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPEGVQKAAKCIIGVNYPKPMVNHAEASRLNIERMKQIYQQLSRYRGM 1 2 GLLASVPSNPNGNGGLGCSLAENIPVCNSAA 1 2 GAQMGGDDGHKVSVLAYTQGDSRAGEIEMQQQ* 1 2 GYGPGSTILRYAHGDSQHTRAVLLQQ 1 2 GRSPIGSSVTSGKRPSPDEETQSIGPKVQRQNSN* 0 >CRY1B_latCha Latimeria chalumnae (coelocanth) last exons uncertain 0 MVVNSVHWFRKGLRLHDNPALQEALKGADTVRCVYMLDPRFAGSSNLGINRWR 2 1 FLLQCLEDLDASLRKLNSRLFVIRGQPANVFPRLFK 0 0 EWKITRLTIEYDSEPFGKERDAAIKKLASEAGVEVIVKISHTLYDLDK 2 1 IIELNGGHPPLTYKRFQTLVSRMEHPEMPVEAITASFMGKCITPVSEDHHEKYGVPSLEEL 1 2 GFDTEGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFEKPRMNANSLLASPTGLSPYLRFGCLSCHLFYFKLTDLYKK 0 0 VKKNGSPPLSLYGQLLWREFFYTAATNNPQFDKMEGNPICVRIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEDLLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWSAPESVQTAAKCIIGVHYPKPVVNHAEASRLNIERMKQIYQQLSRYRGM 1 2 GLLASVPSNHNGNGNGGMMVYSPVEPTLGTNAAH 1 2 GFSIASGHSNTSILLSYSNDEQPGPSGIQ 1 2 GWSNYGNANGNSSGKRDREPEQDGSTDDETHMALVK 1 2 * 0 >CRY1A_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01025403 0 MVVNTVHWFRKGLRLHDSPALRESLKGAASLRCVYILDPWFAGSSNVGINRWR 2 1 FLLQCLEDLDASLRKLNSRLFVIRGQPADVFPRLFK 0 0 EWNINRLSFEYDSEPFGKERDAAIKKLASEADVEVIVRVSHTLYDLDK 2 1 IIELNGGQPPLTYKRFQTLISRMDPLETPAEPITAEVMGKCTTPVSDDHDEKFGVPSLEEL 1 2 GFDTEGLPSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRHEGWIHHLARHAVACFLTRGDLWVSWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPDSVQKAAKCVIGVHYPKPMVHHAEASRLNIERMKQIYQQLSCYRGL 1 2 GLLASVPSNPNGSGGAGMTGFSSGEGLHSSNAA 1 2 GYQIPSGRSSLPSYSSGDSQAGSNLLPQ 1 2 GYTRGSTVVHYTQGDSQQTS 1 2 GRGHLPSVLTTVKRPSPEESTQGIVPKVQRQSSN* 0 >CRY1B_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01016727 AHAT01016728 0 MGPNSIHWFRKGLRLHDNPALKEAIRGASTVRCVYFLDPWFAGSSNLGVNRWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPANVFPRLFK 0 0 EWKISRLTFEYDSEPFGKERDAAIKKLAKEAGVEVIVKISHTLYDLDR 2 1 IIELNGGQPPLTYKRFQTLISRMDPPELPVDPLSEVFMGKCVTPVSDDHGDKYGVPSLEEL 1 2 GFDTEGLPSAVWPGGETEALTRIERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYRK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATTNPRFDKMEGNPICVRIPWDRNSEALAKWAEAKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPDSVQNAAKCVIGVHYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSNHNGNGNGGMMAYSPGEQQPAHGTTY 1 2 GPGSSVASGNGSSSILLNFDGEEQPGPSGVHH 1 2 1 2 GRWAHGGAGGGSSAPGKRERETERDAAGDEEARSSA* 0 >CRY1A_danRer Danio rerio (zebrafish) NM_001077297 whole genome duplicate of retained CRY1 duplicate 0 MVVNTVHWFRKGLRLHDNPSLRDSILGAHSVRCVYILDPWFAGSSNVGISRWR 2 1 FLLQCLEDLDASLRKLNSRLFVIRGQPTDVFPRLFK 0 0 EWNINRLSYEYDSEPFGKERDAAIKKLANEAGVEVIVRISHTLYDLDK 2 1 IIELNGGQSPLTYKRFQTLISRMEAVETPAETITAEVMGPCTTPLSDDHDEKFGVPSLEEL 1 2 GFDTEGLSSAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYRK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPRFDKMEGNPICVQIPWDKNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVSFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESVQKAAKCIIGVHYPMPMVHHAEASRLNIERMKQIYQQLSCYRGL 1 2 GLLAMVPSNPNGNGENSTSLMGFQTGDMTKEVTTPS 1 2 GYQMPPTSQGEWHGRTMVYSQGDQQTSSIMTSQ 1 2 GFGNNGSTMCYRQDAQQIT 1 2 GRGLHSSIIQTSGKRHSEESGPTTVSKVQRQCSS* 0 >CRY1A2_danRer Danio rerio (zebrafish) BC044558 AW184635 olfactory old teleost CRY1 duplicate syntenically retained as tetrapod CRY1 0 MVVNTVHWFRKGLRLHDNPSLRDSIKGADNLRCVYILDPWFAGSSNVGISRWR 2 1 FLLQCLEDLDASLRKLNSRLFVIRGQPTDVFPRLFK 0 0 EWKISRLSYEYDSEPFGKDRDAAIRKLATEAGVEVFVRISHTLYDLDK 2 1 IIEFNGGQSPLTYKRFQTLISRMDPVEMPAETITAEIMGKCSTPVSDDHDDKFGVPSLEEL 1 2 GFETEGLSTAVWPGGETEALTRLERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYRK 0 0 VKKNSTPSLSLYGQLLWREFFYTAATNNPHFDKMEFNPICVQIPWDRNPEALAKWAEGQTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPILRGFPAKFIYDPWNAPESVQKVAKCIIGVHYPKPMVNHAEASRINIERMKQIYQQLSCYRGL 1 2 GLLATIPSNTANNEDSNGAGPSSGDTQESTHIT 1 2 VHTLPASSQSEFPAFAVSCGQQLVNSVQPQP 1 2 GYSGAGMVGQAHKQTQR 1 2 VRGMTAFKRHGEDRTTDFPSKIQRNDCN* 0 >CRY1B_danRer Danio rerio (zebrafish) BC095305 EB921055 aka CRY2A whole genome duplicate of lost CRY1 duplicate 0 MAPNSIHWFRKGLRLHDNPALQEAVRGADTVRCVYFLDPWFAGSSNLGVNRWR 2 1 FLLQCLDDLDSNLRKLNSRLFVVRGQPANVFPRLFK 0 0 EWKISRLTFEYDSEPFGKERDAAIKKLAMEAGVEVIVKTSHTLYNLDK 2 1 IIELNGGQPPLTYKRFQTLISRMDPPEMPVETLSNSIMGCCVTPVSEDHGDKYGVPSLEEL 1 2 GFDIEGLPSAVWPGGETEALTRIERHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYRK 0 0 VKKTSTPPLSLYGQLLWREFFYTAATTNPRFDKMEGNPICVRIPWDKNPEALAKWAEAKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDFIR 2 1 RYLPILRGFPAKYIYDPWNAPDSVQAAAKCIIGVHYPKPMVNHAEASRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSTHNGNGNGMAYSPGEQQSGTNTP 1 2 APAVSSGSVASGNRSGSILLNFDSEEHQGPSGIQQQHQQQQQQQQQQQL 1 2 GYHHMPDSGHNSRFYKSNVSHDMA 1 2 GHLLHKGGSVTGKRERESERDLDGEDESLSTSHKLQRQIAEVTSVYASSGNQSSMVNITLLNL* 0 >CRY1C_danRer Danio rerio (zebrafish) BC164795 EE210836 aka CRY2B old CRY1 duplicate lost in tetrapods CRY1 C12ORF23 CRY4 0 MSVNSVHWFRKGLRLHDNPALLEALNGADTLRCVYFLDPWFAGASNLGVNRWR 2 1 FLLQSLEDLDASLRKLNSCLFVIRGQPADIFPRLFK 0 0 EWKVSRLTFEFDSEPFGKERDAAIKKLACEAGVEVIVKISHTLYDLDR 2 1 IIELNGGQSPLTYKRFQTLVSSMEPPDPPLSSPDRGMMGKCVTPISENHRDKYGVPLLEEL 1 2 GFDTEGLAPvVWPGGESEALKRMERHLdrt 0 0 AWQENFERPKMNASPLMASPLGLSPYLRFGCLSCRLFYCKLTQLYKK 0 0 VKKNMNPSISLYDKILWREFFYTAATNNPRFDRMEGNPICIRIPWDRNAEALAKWAEAKTGFPWIDAIMMQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWLCHSCSSFFQQFFHCYCPVGFGRRIDPNGDFIR 2 1 RYLPVLRDFPAKYIYDPWNAPHDVQLAAKCVIGVDYPKPMVNHAEASRLNIERMRQIYQQLSRYRGL 1 2 GLLATVPSNHIEDAAANVNRGAMAPPTRRKTRNDASSY 1 2 RVRREQPGPSGAKHRHRHSLS 1 2 HRPSPYATQSELTDHNHEFAVPQQP 1 2 GRVSWPNVGGTGKKEGEAEHNCCRGDNGPSSSALKAQQQDNEVCRK* 0 >CRY1A_leuEri Leucoraja erinacea (skate) AESE010236716 AESE011153531 AESE010038968 AESE010673288 AESE012524396 0 MGCNSIHWFRKGLRLHDNPALLESVQGVDTIRCVYIVDPWFAGTSSLAINRWR 2 1 FLIESLEDLDISLRKLKSRLFVVRGQPTDVFPRLFK 0 0 2 1 IERNGGQPPLTYKHFQKLISQMGPVEMPVKMITSETMGRCTMPVLDDHGDRFGVPSLEEL 1 2 GFETEGLTSAVWPGGETEALARLERHLERK 0 0 AWVANFERPGMNSKSLLASPTGLSPYLRFGCLSSRLFYFKLTELYKK 0 0 VKKNSSPPLSLYGQLFWREFFYAAGTNNPKFDKMEGNPICVQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGNWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGYPAKHIYDPWNAPEDVQRAANCVIGVDYPRPMVHHAEASRLNIERMKQIYQQLSCYRGL 1 2 1 2 * 0 >CRY1B_leuEri Leucoraja erinacea (skate) AESE011669465 AESE012563587 AESE010604630 AESE011547252 0 MVRNAVHWFRKGLRLHDNPALQRALSGADTVRCVYILDPCFASSSNIRINWWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPANVFPRLFK 0 0 KWNVTQLTFEYDSEPFGKERDAAIMKLAKIAGVEVIVQISHTLYDVDR 2 1 IIELNGGQPPLTYKRFQTLISRMDPPELPVDPLSEVFMGKCVTPVSDDHGDKYGVPSLEEL 1 2 GFDTSELGCAVWQGGETKALAWLHKHFEGK 0 0 ACTASYESSNLNPDSLMSSSTGLCPYLCLGCLSCRKFYYTLLEFYRM 0 0 IKDHNKSPLPLYGELLRREFFYTAATNNPRFDRMEGNPVCVQIPWDKNQQALAKWADGQTGFPWIDAIMKQLRQEGWIHHLARRAVGCFLTRSDLWISWEEGMK 0 0 VFNELLLDADFSVSAGWWMWLSCSAFFQRFLPCYCPVAFGRRTDPNGDYVR 2 1 1 2 1 2 * 0 >CRY1A_calMil Callorhinchus milii (shark) AAVX01551101 AAVX01266331 AAVX01354908 AAVX01055947 0 MVVNSVHWFRKGLRLHDNPALQAAVSGADTVRSVYILDPWFAGSSNVRINKWR 2 1 FLLQCLEDLDISLRKLKSRLFVVRGQPTDVFPRLFK 0 0 EWNVTRLTFEYDSEPFGKERDVAIVNLARKAGVETIVHTSHTLYDVNR 2 1 1 2 GFDTEGLSSAVWPGGESEALTRLERHLERK 0 0 AWVANFERPRMNSNSLLASPTGLSPYLRFGCLSCRLFYFKLTDLYKK 0 0 SLYGQLLWREFFYAAATNNPRFDKMEGNPICVQIPWDRNAEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYLPVLRGFPAKYIYDPWNAPESVQKAAKCILGVHYPKPMVNHAEASCLNIERMKQIYQQLSCYRGL 1 2 GLLDSVPSNPNGNGG IMGYSAGENTHGCNNLT 1 2 VPSNPNGNGGIMGYSAGENTHGCNNLT 1 2 1 2 * 0 >CRY1B_calMil Callorhinchus milii (shark) AAVX01090452 AAVX01101328 AAVX01636526 AAVX01201905 0 MVVNSVHWFRKGLRRHDNPVLQAAVSGADTIRSVYILDTWFTGSSNVHINKWR 2 1 FLLQCLEDLDANLRKLNSRLFVIRGQPADVFPRLFK 0 0 2 1 1 2 0 0 0 0 WADGLTGFPWIDAIMKQLRKEGWIHHLARHAVSCFLTRGDLWISWEEGMK 0 0 VFEELLLDADFSINAGSWMWLSCSAFFQQFFHCYCPVGFGKRTDPSGDYIR 2 1 RYLPLLKEYPTKYIYEPWNAPLAVQKAAKCIIGIDYPKPIVNHAEASRLNIERMKQIYQQLSHYKGL 1 2 1 2 1 2 1 2 * 0 >CRY1_petMar Petromyzon marinus (lamprey) Contig24766 0 MVLNSIHWFRKGLRLHDNPALREALQGADTVRCVYILDPWFAGASNVGINRWR 2 1 FLLQSLEDLDASLHKLNSRLFVIRGQPADVFPRLFK 0 0 EWNISRLTFEYDSEPFGKERDAAIKKLASEAGVEVLVRISHTLYNLDR 2 1 IIELNEGQAPLTYKRFQALVSRMEPPERPVGTITSEVMGPCRTPLWEDHDERYGVPSLEEL 1 2 GFDTDGLTTAVWQGGESEALTRLDRHLERK 0 0 AWVANFERPRMNANSLLASPTGLSPYLRFGCLSCRLFHYKLTELYKK 0 0 VKKNSSPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICVQIPWDRNPEALAKWAEGRTGYPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVGFGRRTDPNGDYIR 2 1 RYIPLLRAFPAKYIYDPWNAPESVQKTARCIIGVDYPKPMANHAEASRLNIERMKQIYQHLSRYRGL 1 2 1 2 1 2 1 2 * 0 >CRY1_braFlo Branchiostoma floridae (amphioxus) XM_002609455 end uncertain 0 MKSTIHWFRKGLRLHDNPSLRDALQMLQPGDVWRCVYVLDPWFAGSSNVGVNRWR 2 1 FLLQCLEDLDASLRKLNSRLYVIRGQPTDVFPRLFK 0 0 EWKVSCLSFEEDSEPFGRERDMAVMKLAKEAGVKVILRTSHTLYKLQD 2 1 IIDVNGGQPPLTYKRFQAILSKMDPPDQPVDSITASTVDNLTTPIRDDHDEKFGVPSLEEL 1 2 GFDTDNLNPVVWPGGETEALTRLERHLERK 0 0 AWVASFERPKMTASSLLASPTGLSPYLRFGCLSPRLFYKKLTELYKK 0 0 VKRSNHPPLSLYGQLLWREFFYTAATNNPNFDKMVNNTICVQIPWDKNPEALAKWAE 00 GQTGFPWIDAIMAQLQQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSSFFQQFFHVYCPVNFGRRTDPNGDFIR 2 1 KYLPKLRGFPAKYIYDPWNSPESVQKAAKCII 12 GKDYPLPMVNHAETSRINVERMKQVYQQLSRYRGA 1 2 GLLATIPYPDGGNGNPTGHKKHAIVQGMERQHAERREHGQPAHVSRQHI* 0 >CRY1A_strPur Strongylocentrotus purpuratus (urchin) XM_001194752 same split exons as braFlo, end of gene uncertain, partial duplicate elsewhere 0 MPKRKHRSSRRDREEKPGCLVHWFRKGLRLHDNPALKEGLKSASGFRCIYILDPWFAGSCSKGVNRWR 2 1 FLLECLEDLDSSLRKLNSRLFLIRGQPADVLPRLFK 0 0 EWKVTQLSFEEDSEPFGRTRDKAISTLAQEAGVKVISKVSHTLYDPQE 2 1 ILALNNNEPPLTYKRFQDIISLMGIPIYPAEALEAEDVEGLDTFIDPNHEDKYGIPTLEEL 1 2 GFDPEDVPPPMWIGGETEAKQRLDRHLERK 0 0 AWVANFERPRMSPASLMASPAGLSPYLRFGCLSPRTFYWKLTELYQK 0 0 VRKTTNTPLSLHGQLLWREFFFTVACNNRQFDHMVDNPICIQIPWDKNTALLNKWAN 00 GETGYPWIDAIMTQLRLEGWIHPLARHAVACFLTRGDLWISWEEGMK 0 0 VFDEYLLDADWSVNAGNWIWLSCSSFYQQFFHCYCPVKFGRRTDPNGDYVR 2 1 KYLPFLKNFPSKYIFEPWTAPIEVQEKAKCII 12 GKDYPLPIVDHAEASHRNIERMRKVYNNLSRRGGP 1 2 GILGSVPSSQAASAPPQLGKFNMSSKHGQLLCITVYYCVLLCITV* 0
Vertebrate CRY2 reference sequences
>CRY2_homSap Homo sapiens (human) 11 exons 0 MAATVATAAAVAPAPAPGTDSASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGLVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKAFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSMSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPTPELPSKDA* 0 >CRY2_panTro Pan troglodytes (chimp) 0 MAATVATAAAVAPAPAPGTDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKAFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSMSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPTPELPSKDA* 0 >CRY2_gorGor Gorilla gorilla (gorilla) 0 MAATVATAAAVAPAPAPGTDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKAFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSVSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPTPELPSKDA* 0 >CRY2_ponAbe Pongo pygmaeus (orangutan) 0 MAATVATAAAVAPAPAPGTDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKAFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSVSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPTPELPSKDA* 0 >CRY2_rheMac Macaca mulatta (rhesus) CJ488220 testis 0 MAATVATAAAVAPAPAPGTDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLER 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSVNSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRAKVAELPTPELPSKDA* 0 >CRY2_papHam Papio hamadryas (baboon) 0 MAATVATAAAVAPAPAPGTDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLER 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSVNSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRAKVAELPTPELPSKDA* 0 >CRY2_calJac Callithrix jacchus (marmoset) 0 MAATVATAAAAVPAPAPGTDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSMSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPTPELPSKDV* 0 >CRY2_micMur Microcebus murinus (mouse_lemur) 0 MATAVATAAAAPTPASSTDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWr 2 1 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNSGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSMSST 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVTELPTPELPSKDA* 0 >CRY2_musMus Mus musculus (mouse) CF898022 0 MAAAAVVAATVPAQSMGADGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQALISRMELPKKPAVAVSSQQMESCRAEIQENHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPGSSQAGSISNT 1 2 GPRALSSGPASPKRKLEAAEEPPGEELTKRARVTEMPTQEPASKDS* 0 >CRY2_ratNor Rattus norvegicus (rat) DN948283 prostate 0 MAAAAVVAATVPAQSMGADGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQALISRMELPKKPVGAVSSQHMENCRAEIQENHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYRK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPGSSQAGSISNT 1 2 GPRPLSSGPASPKRKLEAAEEPPGEELSKRARVTEMPAQEPPSKDS* 0 >CRY2_criGri Cricetulus griseus (hamster) XR_135830 0 MAAAAVVAGAPRGARVPALTMGADGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGAVTSQQMENCRAEIQENHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSISNT 1 2 GSRPLSSGPASPKRKLEAAEEPPGEELSKRARVTEMPAQEPQTKDA* 0 >CRY2_spaJud Spalax judaei (blind_molerat) AJ606300 0 MAAASVVVATSAAPAMAVDGGSSVHWFRKGLRLHDNPSLLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIDLNGQKPPLTYKRFQAIISRMELPKKPVVAVTSQQMESCRAEIQDNHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFPTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAARCIIGVDYPRPMVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQPGSITNT 1 2 GPRPLSTGPASPKRKLEAAEEPPGEELSKRARVTELPAPEPASKDA* 0 >CRY2_dipOrd Dipodomys ordii (kangaroo_rat) 0 MAAAMVTAAVAVPAPPSGADGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLQEGWIHHLARHAVTCFLTRGDLWVSW 0 0 2 1 RYLPRLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSISST 1 2 GPRPLSSGPASPKRKLEAAEEPPGEELSKRARVTEMPAQEPPSKDS* 0 >CRY2_hetGla Heterocephalus glaber (naked_molerat) EHA99865 hystricognath 0 MAAAVGTGTGAAPTPATGAEGACSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGAVSSQQMKSCRAEIQENHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPTLSQAGSSSST 1 2 GPRSLPDGPASPKRKLEAAEEPPGEELSKRARVAELPAPEPTSKDA* 0 >CRY2_cavPor Cavia porcellus (guinea_pig) hystricognath 0 MAAAVGTGTAAAPTPVTGAEGACSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGAVSSQQMESCRAEIQENHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSLSQAGSSTST 1 2 GPRPLPGGPASPKRKLEAAEEPPGEELSKRARVAELPTPEPPSKDA* 0 >CRY2_octDeg Octodon degus (degu) AJSA01043118 hystricognath 0 MAAAVGTGTSAAPTPAAGTEGACSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPAEVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGVVSSQQMEGCRAEIPENHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNSGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIVGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSSTST 1 2 GPRPLPSGSASPKRKLEAAEEPPGDELSKRARVAELPTPEPSSKDA* 0 >CRY2_speTri Spermophilus tridecemlineatus (squirrel) 0 MSASVVTTSATLLTPTSADVSSVHWFRKGLRLDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHT 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGAVSSQQMESCKAEIQEDHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKKNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPGSSQAGSISST 1 2 GPRPLSSGQASPKRKLEAAEEPPGEELSKRARVAELPTPELPSKDA* 0 >CRY2_oryCun Oryctolagus cuniculus (rabbit) 0 MAAAAAAAAAAVPAPAASANGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPAGSVTSQQLEGCRAEIQDNHDETYGVPSLEEL 1 2 GFPTEGLSPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVR 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSHPVAEPSSSQAGSVSGA 1 2 APRPLPSGPASPKRKLEAAEEPPGEELSKRARVAEVPTPELPSKAV* 0 >CRY2_turTru Tursiops truncatus (dolphin) 0 MAAAVATSAVAAPAPAARAEGASSVHWFRKGLRLHDNPALQAAVRGAHCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDK 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMEGCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRSSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCMEDLSNPVAEPSSSQAGGMSSA 1 2 GPRPLPSGPASPKRKLGAAEEPPGEELSKRARVAGLPPSELLSKDV* 0 >CRY2_bosTau Bos taurus (cow) EG706191 lens 0 MAAAAAAATQAPAARGDGASSVHWFRKGLRLHDNPALLAAVRGAHCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDRSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDTAIMKMAKEAGVEVVTENSHTLYDLDK 2 1 IIELNGQKPPLTYKRFQAIISRMELPRKPVGSVTSQQMEGCRAEIQESHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCVIGVDYPRPIVNHAEASRLNIERMKQVYQQLSRYRGL 1 2 CLLASVPSCVEDLSNPVAEPSSIQAGSSSSV 1 2 GPRPLPGGPASPKRKLEAAEEPPGGELSKRARVAESLPSELPSRGV* 0 >CRY2_oviAri Ovis aries (sheep) NM_001129736 PubMed:19341811 0 MAAAAAATASAAAAAQAPAPRGDGASSVHWFRKGLRLHDNPALLAAVRGAHCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDRSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDK 2 1 IIELNGQKPPLTYKRFQAIISRMELPRKPVGSVTSQQMEGCQAEIQESHDETYGVPSLEEL 1 2 GFPTEGLGPAVWRGGETEALARLDKHLERK 0 0 AWVASYERPRMNASSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYRK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMAQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAEASRLNIERMKQVYQQLSRYRGL 1 2 CLLASVPSCVEDLSTPVAEPSSSQAGSSSSA 1 2 GPRPLPGGPASPKRKLEAAEEPPGGELSKRARVAESLPSELPSRGV* 0 >CRY2_susScr Sus scrofa (pig) XM_003122835 0 MAAAVATAAASSPAPAAGAEGASSVHWFRKGLRLHDNPALLAAVRGAHCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPALFQ 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDK 2 1 IIELNGQKPPLTYKRFQALISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSNPVAEPSSSQAGSVSAA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPPTELPGRDV* 0 >CRY2_equCab Equus caballus (horse) 0 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPRKPVGSVTSQQMESCRADIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNAASLLASPTGLSPYLRFGCLSCRLFYYRLWDLYRK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEAKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKAFPSRYIYEPWNAPEAVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSNPVAEPSSSQAGSVSSA 1 2 GPRPPPSGPASPKRKLEAAEEPPGEELSKRARVAEPPS* 0 >CRY2_canFam Canis familiaris (dog) XM_540761 0 MAAAVVAAAAAAPVPTAGVDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDK 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRADIQDNHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPILKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSNPVAEPSSSQTGSMSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPTPELPCRDV* 0 >CRY2_ailMel Ailuropoda melanoleuca (panda) XM_002922310 iMet lost to assembly gap 0 PAPAAGVDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDK 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRADIQESHDDTYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPILKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSNPVAEPSSSQTGSMSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPAPEPPSRDV* 0 >CRY2_myoLuc Myotis lucifugus (microbat) 0 MAANAVTAAAAAPAPAAGTDGASSVYWFRKGLRLHDNPALLAAVRGARCVLCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVASVTRHQMESCPAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSSPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGFR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCMEDLSNPVAEPSLSQTGSMSSA 1 2 GPKPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPATELPSRDV* 0 >CRY2_pteVam Pteropus vampyrus (macrobat) 0 MAATVGTAAAAASAPAAGTDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQHMESCTAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLNNPVAEPSSSQTGSMNNA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAELPAPELPSRDV* 0 >CRY2_loxAfr Loxodonta africana (elephant) 0 MAAAVVTAGAAALVPIPSMDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSSPVAEPSSSQAGSSNSA 1 2 GPRPLPSGPTSPKRKLEAAEEPPGEELSKRARVAKLPGPELPSKDV* 0 >CRY2_triMan Trichechus manatus (manatee) AHIN01126950 AHIN01126951 0 MAATVVTAAAAALAPAPSIDGASSVHWFRKGLRLHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDRSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPSLEEL 1 2 GFPTEGLGPAVWQGGETEAMARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCAEDLSNPVAEPSLSQAGSMSSA 1 2 GPRPLPSGPASPKRKLEAAEEPPGEELSKRARVAKLPSPELPSKDV* 0 >CRY2_choHof Choloepus hoffmanni (sloth) 0 MAATAVMAGSAAPAPASGTEGASSVHWFRKGL*LHDNPALLAAVRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIMKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVGSVTSQQMESCRAEIQENHDETYGVPsleel 1 2 gFPTEGLGPAVWQGGEAEALARLDKHLERK 0 0 AWVANFERPQMNANSLLASPTGLSPYLQFGCLSCQLFYFKLTDLYKK 0 0 VKRNSTPPLSLFGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNSGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPKLKGFPSRYIYEPWNAPESIQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSNPVAEPSSSQAGSMSSA 1 2 GPRPVPSGPASPKRKLEAAEEPPGEELSKRARVTELPTPELPSKDV* 0 >CRY2_macEug Macropus eugenii (wallaby) FY652314 testis 0 2 1 FLLQSLEDLDISLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIVKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVSCVTSQQMEQCQAEIQENHDDTYGVPSLEEL 1 2 GFPTDGLGPAVWQGGEMEALARLDKHLERK 0 0 AWVANYERPRMNATSLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 0 VKRNNTPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVR 2 1 RYLPQLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPKPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCMEDLSSPMAETSMGQAGSVSST 1 2 GPKPLPCSPASPKRK LEATEEASREEHSKRARMAALPVPELAS* 0 >CRY2_monDom Monodelphis domestica (opossum) 0 MAAAVVTMTAAAPAPAPSPEGASSVHWFRKGLRLHDNPALQAALRGARCVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDISLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIVKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPRKPVNSVTSQQMERCQAEIQENHDDAYGVPSLEEL 1 2 GFPTDGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNAASLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 0 VKRNNTPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNSGSWMWLSCSAFFQQFFHCYCPGFGRRTDPSGDYVR 2 1 RYLPQLKGFPARYIYEPWNAPEPVQKAAKCIIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCLEDLSSPMVEASLGQAGAVSGP 1 2 GLKPLPCSPASPKRKLEATEEAPGEEHSKRARVARLPGSELAGKDV* 0 >CRY2_ornAna Ornithorhynchus anatinus (platypus) 0 LHDNPALQAALRGARSVRCVYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIVKMAKEAGVEVVTENSHTLYDLDR 2 1 IIELNGQKPPLTYKRFQAIISRMELPKKPVSCVTSQQMEGCKAEIQDNHDDTYGVPSLEEL 1 2 GFPTDGLGPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWDLYKK 0 0 VKRNSTPPLSLYGQLLWREFFYTAATNNPRFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDFIR 2 1 RYLPKLKAFPSRYIYEPWNAPESVQKAAKCVIGVDYPRPIVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSSAAAESGLGQASGSNIST 1 2 APRPPPPPYGPASPKRKLEAAEGLPGEELCKRPKVAGRPGPEPPGEDA* 0 >CRY2_galGal Gallus gallus (chicken) AJ396745 bursa 19456395 15459395 0 MAAAASPPRGFCRSVHWFRRGLRLHDNPALQAALRGAASLRCIYILDPWFAASSAVGINRWR 2 1 FLLQSLEDLDNSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIIKLAKEAGVEVVIENSHTLYDLDR 2 1 IIELNGNKPPLTYKRFQAIISRMELPKKPVSSIVSQQMETCKVDIQENHDDVYGVPSLEEL 1 2 GFPTDGLAPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 0 VKRNSTPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICIQIPWDKNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVK 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPKPMVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDLSGPVTDSAPGQGSSTST 1 2 AVRLPQSDQASPKRKHEGAEELCTEELYKRAKVTGLPAPEIPGKSS* 0 >CRY2_taeGut Taeniopygia guttata (finch) FE716439 brain 0 IYILDPWFAATSAVGINRWR 2 1 FLLQSLEDLDNSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIVKLAKEAGVEVVIENSHTLYDLDR 2 1 IIELNGHKPPLTYKRFQAIISRMELPKKPVSTVISQQMETCKVDIQENHDDVYGVPSLEEL 1 2 GFPTDGLAPAVWQGGETEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 0 VKRNSTPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVK 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPKPMVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCVEDISGPVPDSASGQGCSTST 1 2 AVRLSQAEQASPKRKHEGAEEPCSEELYKRAKVTGLPTSEISGKSL* 0 >CRY2_allMis Alligator mississippiensis (alligator) genome/blat 0 MAASRSFPSSVPARAGPCRAVHWFRRGLRMHDNPALQAALRDAASVRCIYILDPWFAASSAVGINRWR 2 1 FLLQSLEDLDNSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTRLTFEYDSEPFGKERDAAIIKLAKEAGVEVVIENSHTLYDLDR 2 1 IIELNGNKPPLTYKRFQAIISRMELPKKPVSSITSQQMEKCKAEIQENHDDVYGVPSLEEL 1 2 GFPMDGLAAAVWQGGEMEALARLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 0 VKRNSTPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWECGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVK 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPKPMVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 GLLASVPSCAEDLSGPVTDPASVQGCSTST 1 2 ALKPSQSGQASPKRKHEGIEEMCTEDLYKRAKVTGLHGPEIPSKSL* 0 >CRY2_croAda Crotalus adamanteus (rattlesnake) JU174243 TSA 0 MAALPGPVPRPRSGGGCCSVHWFRRGLRLHDNPALQAAIREATSVRCIYILDPWFAASSAVGINRWR 2 1 FLLQSLEDLDNSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWGVTHLTFEYDSEPFGEERDAAIVKLAKEAGVKVTTENSHTLYDLDR 2 1 IIELNGHKPPLTYKRFQAIISRMDLPKKPVSTITSQQMEMCQTKIQENHDETYGVPSLEEL 1 2 GFFTEGLAPAVWQGGETEALTRLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 0 VKRNSTPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAAACFLTRGDLWISWESGVR 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYLPQLKGFPSRYIYEPWNASESVQKAAKCIIGVDYPKPMVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCIEDFSNITTDLSSGSCSNTTL 1 2 RQSHCSQTSPKRKCDAEEDNNEELHKRVRMMVHQVSKKPQQLSQPEFGN* 0 >CRY2_anoCar Anolis carolinensis (lizard) XM_003214641 0 MAALPGPLGRCSVHWFRRGLRLHDNPALQAAIRDGGPVRCIYILDPWFAASSSVGINRWR 2 1 FLLQSLEDLDNSLRKLGSRLFVVRGQPTDVFPRLFK 0 0 EWRVTRLTFEYDSEPFGKERDAAIVKLAKEAGVEVITENSHTLYDLDR 2 1 IIELNGHKPPLTYKRFQTIISRMDLPKKPVASITHQQMEMCKTEIQDNHDETYGVPSLEEL 1 2 GFPTESLAPAVWLGGETEALTRLDKHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLWELYKK 0 0 VKRNSTPPLSLYGQLLWREFFYTAATNNPKFDRMEGNPICIQIPWDRNPEALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWESGVR 0 0 VFDELLLDADFSVNSGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVR 2 1 RYLPKLKGFPSRYIYEPWNAPESVQKAAKCIIGVDYPKPMVNHAETSRLNIERMKQIYQQLSRYRGL 1 2 CLLASVPSCMEDLSNPVADTSSSHGNCIGT 1 2 ASRQTHCGQTSPKRKHDVVQEYKEELYKRAKVVASQFAENPRQELVHHEMGN* 0 >CRY2_xenTro Xenopus tropicalis (frog) NM_001088670 AY049035 CX389867 CX841030 11533577 discrepancies 0 MEGKPSVSSVHWFRKGLRLHDNPALLSALRGANSVRCVYILDPWFAASSSGGVNRWR 2 1 FLLQSLEDLDTSLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWGVSRLTFEYDSEPFGKERDAVIMKLAKEaGVEVVVENSHTLYDLDR 2 1 VIELNGHSPPLTYKRFQAIISRMELPRRPAPSVTRQQMEACRAEIKRNHDETYGVPSLEEL 1 2 GFHsEiKGpsIWPGGETEALARLDRHLERK 0 0 AWVANYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLKELYKK 0 0 VKKNSPPPLSLYGQLLWREFFYTAATNNPKFDQMEGNPICVQIPWDKNPKALAKWAEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWNSWECGVK 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVR 2 1 RYLPVLKAFPSRYIYEPWSAPESVQKEAKCIIGIDYPKPIVNHAEASRMNIERMKQTYQQLSRYRGL 1 2 CILASVPSCMEDLGGPMADSSQNISEA 1 2 APKQSLCNVESPKRKRKVSEEEPHMKVRVLCAPETERPAKDF* 0 >CRY2_ranCat Rana catesbeiana (bullfrog) GO458565 AY256684 extra SS removed 0 MEGPAVSSVHWFRKGLRLHDNPALLAALRGARCVRCVYILDPWFAASSSGGVNRWR 2 1 FLLQSLEDLDSSLRKLGSRLFVGRGQPADVFPRLFK 0 0 EWGVTRLTFQYYSEPFGKERDAAIMKLAKEAGVEVIVESSHTLYDLDK 2 1 IIELNGNSPPLTYKRFQAIVSRMELPRRPVPSITRQQMEKCRAEIKSTHDDTYGVPSLEEL 1 2 GFPRDNPGAAVWPGGETEALARLDRHLERK 0 0 AWVAHYERPRMNANSLLASPTGLSPYLRFGCLSCRLFYYRLRELYQK 0 0 VKKNSPPPLSLFGQLLWREFFYTAATNNPKFDQMEGNPICVQIPWDKNPEALAKWTEGKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWNSWECGVK 0 0 VFDELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYVR 2 1 RYLPVLKGYPSRYIYEPWNAPESVQKEAKCIVGVDYPKPVVNHAEASRLNIERMKQTYQQLSRYRGL 1 2 CILASVPSSVEDLSGPMADPASSQQGSS 1 2 DPAPRLCAVDSPKRKHEDLDGELCKKARLQCVQEMERAAKDFL* 0 >CRY2_latCha Latimeria chalumnae (coelocanth) AFYH01005158 AFYH01005161 AFYH01005164 0 MVVNSVHWFRKGLRLHDNPALQEALKGADTVRCVYMLDPRFAGSSNLGINRWR 2 1 0 0 EWNVTRLTFEYDSEPFGKERDAVIIKVARESGVEVIMRDSHTLYNLNR 2 1 IIELNGHKPPLTYKRFQAIISRMELPRRPVAAVTQQQMETCKTDIGNDHDEQYGVPSLAEL 1 2 GFQVENDPAIWQGGETEALARLEKHLDRK 0 0 AQVANFERPWINANSLLANPTSLSPYLRFGCLSCRLFYYRLLELYKK 0 0 IKGNSSPPLSLYGQLLWREFFYTAATNNPRFDRMEGNPICVQIPWDSNPKALAKWAEGQTGFPWIDAIMTQLRQEGWIHHLARHSVACFLTRGDLWISWECGMK 0 0 VFEELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDFVR 2 1 1 2 1 2 * 0 >CRY2_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01038797 0 MVVNSVHWFRKGLRLHDNPALQEALNISDTVRCVYILDPWFAASANVGINRWR 2 1 FLLESLEDLDASLRKLNSRLFVVRGQPADVFPRLFK 0 0 EWNVTHLTFEYDSEPFGKERDAAIMKLARESGVETIVQESHTLYSLDR 2 1 IIELNNNSPPLTFKRFQAIVSRLGLPKKPLGTVTPQQMERCHTPISENHDECYRVPTLEEL 1 2 GFKTHGLSPSIWKGGETEAVRRLNKHLERK 0 0 AWVASFERPKINVYSLIASPTGLSPYLRFGCVSCRLFYYSLLELYKK 0 0 VRKSSSPPLSLFGQLLWREFFYTAATNNPKFDQMEGNPICVQIPWDRNPEALAKWAEGRTGYPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWVSWESGMR 0 0 VFEELLLDADFSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYIPKLKEYPNRYIYEPWNAPEIVQKAANCVIGVDYPKPMINHAQASRLNIERMKQIYQQLSHYKGL 1 2 GLLASVPSVPDDARSPVTDDPHGHSS 1 2* 0 >CRY2_danRer Danio rerio (zebrafish) aka CRY3 NM_131786 0 MVVNSVHWFRKGLRLHDNPALQEALNGADTVRCVYILDPWFAGSANVGVNRWR 2 1 FLLESLEDLDTSLRKLNSRLFVVRGQPTDVFPRLFK 0 0 EWNVTRLTFEYDSEPYGKERDAAIIKMAQEYGVETVVRNTHTLYNPDR 2 1 IIEMNNHSPPLTFKRFQAIVNRLELPRKPLPTITQEQMARCRTQISDNHDEHYGVPSLEEL 1 2 GFRTQGDSLHVWKGGETEALERLNKHLDRK 0 0 AWVANFERPRISGQSLFPSPTGLSPYLRFGCLSCRVFYYNLRDLFMK 0 0 LRRRSSPPLSLFGQLLWREFFYTAGTNNPNFDHMEGNPICVQIPWDHNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWESGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYIPKLKDYPNRYIYEPWNAPESVQKAANCIVGVDYPKPMINHAESSRLNIERMKQVYQQLSHYRGL 1 2 SLLASVPTIQEEAEPPMSDDSQASSSST 1 2 GQASSPPHLSITAPSTPPLSESSSPNSSPTASTSVPHTQRKRVRPSETPSKQKTKVKHTSQARGLELKMDDKQ* 0 >CRY2_oreNil Oreochromis niloticus (tilapia) XM_003449249 split exon 7 also in gasAcu, oryLat, tetNig not danRef or lepOcu 0 MVVNSVHWFRKGLRLHDNPALQEALNGADAVRCVYILDPFFAGAANVGINRWR 2 1 FLLEALEDLDSSLKKLNSRLFVVRGQPTDVFPRLFK 0 0 EWNVTRLTFEYDPEPYGKERDGAIIKMAQEFGVETIVRNSHTLYNLDR 2 1 IIEMNNNSPPLTFKRFQTIVSRLELPRRPLPPITQQQMDKCHTKIGDNHDQLYSIPSLEEL 1 2 GFRTAGLPPAVWRGGESQALDRLSKHLDKK 0 0 VWVTSLEHPRVNTCSLYASPTGLSPYLRFGCLSCRVLYYNLRELYMK 0 0 VRKRCSPPLSLFGQLLWREFFYTAATNNPNFDRMDGNPICVQ 0 0 IPWDQNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWESGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPTGDYIR 2 1 HYIPILKDYPNRYIYEPWNAPESLQKAANCVVGVDYPKPMINHAESSRLNIERMKQVYQQLSHYRGL 1 2 SLLASVPTIQEEAEPPMTDESQTSSGP 1 2 DSPQRVPANTEGAGCYEAPDSSTVCPSSTSAPHPDQEHQAPCISSGCHTDPS* 0 >CRY2_sigGut Siganus guttatus (spinefoot) AB643456 full length? imputed introns Percomorpha PUBMED 22163321 lunar phase-recognition 0 MVVNSVHWFRKGLRLHDNPALQEALNGADTVRCVYILDPWFAGSANVGINRWR 2 1 FLLEALEDLDNSLKKLNSRLFVVKGQPTDVFPRLLK 0 0 EWKVTRLTFEYDPEPYGKERDGAIIKMAQEFGVETIVRNSHTLYNLDR 2 1 IIEMNSNNPPLTFKRFQTIVRRLELPRRPLAPITQQQMDKCRTKIADNHDQLYSIPSLEEL 1 2 GFRTEGLPAAVWRGGESEALDRLNKHLDKK 0 0 VWVANLEHPRVNTCSLYASPTGLSPYLRFGCLSCRVLYYNLRELYMK 0 0 LCKHSSPPLSLFGQLLWREFFYTAATNNPNFDRMEGNPICVQ 0 0 IPWDQNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHVARHAVACFLTRGDLWISWESGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSAFFQQFFHCYCPVGFGRRTDPSGDYIR 2 1 RYIPILKDYPNRYIYEPWNAPESVQKAANCVVGVDYPKPMINHAEGSRLNIERMKQVYQQLSHYRGL 1 2 SKSTHQFLPV 1 2 * 0 >CRY2_tetNig Tetraodon nigroviridis (fugu) CAAE01010345 0 MVVNSVHWFRKGLRLHDNPALQEALSGADSLRCVYVLDPWFAGAANVGINRWR 2 1 FLLEALEDLDCSLKKLNSRLFVVRGQPTDVFPRLLK 0 0 EWKVTRLTFEYDPEPYGKERDGAIIKMAQQFGVETIVRNSHTLYNLDR 2 1 IVEMNNNSPPLTFKRFQTIVSRLELPRRPLPSVTQQQMDKCGTPVADNHDQLYSIPSLEEL 1 2 GFRTGGLAPAVWRGGESEALERLRKHLEKK 0 0 VWVSHLEHSRANTCSLYASPAGLSPYLRFGCLSCRVFYYNLREHYIR 0 0 LCKGCSPPLSLFGQLLWREFFYTAATNNPNFDRMAGNPICVQ 0 0 IPWDQNPEALAKWAEGRTGFPWIDAIMTQLRQEGWIHHQARRASACFLTRGDLWISWECGMK 0 0 VFEELLLDADWSVNAGSWMWLSCSAFFQQFFKCYCPVGFGRRTDPSGDYIR 2 1 RYIPILKDYPNRYIYEPWNAPEAVQKAANCVVGVDYPRPMINHAEGSRLNIERMKQVYQQLSHYRGL 1 2 SLLASVPTIQEEADTP 1 2 * 0 >CRY2_takRub Takifugu rubripes (fugu) HE592015 0 MVVNSVHWFRKGLRLHDNPALQEALSGADSLRCVYVLDPWFAGAANVGINRWR 2 1 FLLEALEDLDCSLRKLSSRLLVVRGQPTDVFPRLLK 0 0 DWKVTRLTFEFDPEPYGKERDGAIIKLAQQFGVETIVRNSHTLYNLDR 2 1 IIEVNNNSPPLTFKRFQTIVSRLELPRRPLPTVTQHQIHKCGAKMADSQEQLYSIPSLEEL 1 2 GFRTEGLPPAVWRGGESEALERLHKHLDKK 0 0 VWVANLEHSRVSTCSLYASPAGLSPYLRFGCLSCRVLYYNLRELYVK 0 0 LRKGCSPPPSLFGQLLWREFFYTAATNNPNFDRMEGNPICVQ 0 0 IPWDQNPEALAKWAEGHTGFPWIDAIMTQLRQEGWIHHQARRAVACFLTRGDLWISWE 0 0 VFEELLLDADWSVNAGSWMWLSCSAFFQQFFKCYCPVGFGRRTDPSGDYIR 2 1 RYIPILKDYPNRYIYEPWNAPEAVQKAANCVVGVDYPRPMINHAEGSRLNIERMKQVYQQLSHYRGL 1 2 SLLASVPTIQEEAEPPMTDEFQNSSGP 1 2 * 0
Invertebrate cryptochromes
>CRY1B_strPur Strongylocentrotus purpuratus (sea_urchin) XM_001183029 echinoderm lacks final 2 exons 0 MPGGACIHWFRHGLRLHDNPALLEGMTLGKEFYPVFIFDNEVA 1 2 GTKTSGYNRWRFLHDCLVDLDEQLKAAGGRLFVFHGDPCLIFKEMFL 0 0 EWGVRYLTFESDPEPIWTERDRRVKALCKEMKVECIERVSHTLWNPDI 2 1 IIEKNGGTPPITYSMFMECVTEIGHPPRPMPDPILTKVNMKIPSDFEERCALPSLEVM 1 2 GVNMECTEQEKKVWKGGETRALELFRVRILHEEE 0 0 AFKGGYCLPNQYMPDLLGTPKSLSAYLRFGCLSVRRFYWKIHDTYSEVRS 0 0 EVSPSHLTAQVIWREYFYTMSVGNIHFNKMKENPICLNIEWKEDDEKLKAWTD 0 0 GRTGYPWIDACMKQLKYEGWIHQVGRHATACFLTRGDLWISWEDGLQ 0 0 VFDKYLLDADWSICAGNWMWISSSAFEKFLQCPNCFCPVRYGRRMDPTGEYVR 2 1 RYLPVLKDMPIRYLFEPWKAPRAVQERAKCIVGKDYPMPVVEHKSASAANHEQMEKVVNKLRDT 1 2 GIVHCAPSTQREVREFVWLPEKMAGGGSCRADQNCEGILGL* 0 >CRY1B_lytVar Lytechinus variegatus (sea_urchin) AGCV01081039 echinoderm many small contigs 0 MPGGACIHWFRHGLRLHDNPALLEGMTLGTEFYPIFIFDNEVA 1 2 GTSTSGFNRWRFLHDCLADIDEQLKAGGGRLFVFQGDPCHVFKEMFL 0 0 EWGVRYLTFESDPEPIWTERDRKVKALCKEMNVECIERVSHTLWNPDI 2 1 IIEKNGGTPPITYSMFMECVTEIGHPPRPMPDPIFTKVKMNVPADFEERFSLPSLEAL 1 2 GVRLECPEQEKKVWKGGETRALELFRVRILHEEE 0 0 AFKGGYCLPNQYMPDLLGTPKSLSAYLRFGCLSVRKFYWKIHDTYSEVRS 0 0 EVSPSHLTAQVIWREYFYTMSVGNIHFNKMKENPICLNIEWKKDDEKLEAWTE 0 0 GRTGYPWIDACMKQLKYEGWIHQVGRHATACFLTRGDLWISWEDGLR 0 0 VFDKYLLDADWSICAGNWMWISSSAFEKFLQCPNCFCPVRYGRRMDPTGEYVR 2 1 RYLPVLKDMPIRYLFEPWKAPRAVQERAKCIVGKDYPMPIVEHKSASAANTEQMEKVINRLRDS 1 2 GIVHCAPSTQKEVREFVWLPEKMAGGGSCRADQNCEGILGL* 0 >CRY1B_parLiv Paracentrotus lividus (sea_urchin) AM599080 echinoderm many transcripts 0 MPSGACIHWYRHGLRLHDNPALLEGMTLGTEFYPIFIFDNEVA 1 2 GTRTSGYNRWRFLHDCLQDLDDQLKEAGGRLYVFHGDPCLVFKEMFI 0 0 EWGVRYLTFESDPEPIWTERDLKVKALCKEMNVECIERVSHTLWNPEI 2 1 IIEKNGGTPPITYSMFMECVTEIGLPPRPMPDPVLNKVTFKIPPDFEERFALPSLEKM 1 2 GVNMECKEQERKIWKGGGTRALELFKVRILHEEE 0 0 AFKGGYCLPNQYMPDLMGTPKSLSAYLRFGCLSVRKFYWKIHDTYSELKN 0 0 EVSPSHLTAQVFWREYFYTMSVGNIHFNKMKENPICLNIEWKKDDEKLEAWTE 0 0 GPTGYPWIDACMIQLKYEGWIHQVGRHATACFLTRGDLWISWEDGLK 0 0 VFDKFLLDADWSICAGNWMWISSSAFEKFLQCPNCFCPVRYGRRMDPTGEYVR 2 1 RYLPVLKDMPIRYLFEPWKAPRAVQERAKCIVGKDYPMPIVEHKSASAANHEMMEKVINRLRDS 1 2 GIVHCAPSTQKEVREFVWLPEKMAGGGSCRASQNCEGRTGSDGVGRGRTGDR* 0 >CRY1B_aplCal Aplysia californica (sea_hare) FF067636 AASC02010117 scaffold_151 mollusc 0 MEENDTKSHVSVHWFRHGLRLHDNPALLDSLENCREFYPVFILDGNVA 1 2 GISSAAFPRMQFLFETLQDLDDNLRSHGSRLFVLRGKPVDVFAKMFE 0 0 EWGVTRLTFEQDPEPVWQERDNSVKALCDKQNVEWIERVSHMLWEPRL 2 1 ILEENGGEPPLTYAMFNQVAQVVGPPPKPVGDPDFSGISLPCRKDDEESYKLPSLEEI 1 2 GIRPESEEQASPFGRYLG GETKALKLLSSRLEKEHE 0 0 AFSIGQCLPTQLNPNLIGMPMSLSPHLRFGSLSVRKFYWSLRDTFSQ 0 0 lfpdrpvPGSITSQLVWREYFYCMSVNNPMYNRMKENPICLDIDWYDDEQKFEKWKK 0 0 GQTGFPWIDACMRQLLQEGWIHQVCRHAVACFLTRGDLWIDWQKGLK 0 0 VFDRYLLDADWSVCAGNWMWVSSSAFEKVLQCPRCICPVRYGKRNDPTGAYVR 2 1 RYVPELKRMPLQYLFEPWKAPLKLQEEVDCIIGKDYPEPMVDHRTASKECKTKMEAIKKVSK 1 2 DVPHIAPANEEEVLTLMWSGKQTRSELMDACTHNLDKIWCVWSLSFVYTLSVNLKAYKKFFNPYSC* 0 >CRY1B_octVul Octopus vulgaris (octopus) JR450373 transcript assembly mollusc 0 MKLEKKQKIAVHWFRHGQRLHDNPALLDALKDCDEFYPVFIFDGEVA 1 2 GTKLCGFNRWRFLLENLKDLDESFSEYGGRLYTFQGKPVEVFANLQN 0 0 EWGITHITAEIDPEPIWQERDDAVKEFCQKSGIKCDFFNSHTLWDPKR 2 1 LLKKNGGTPPLTFELFQLVTSSLGPPPRPIDAPTFEGIKMPLPENHDKFSVPTLKSL 1 2 GIYPEFEEQKNPINVFIG GEKRALVLLKARLEKEAQ 0 0 SFRHGQCLPNHQEQPELLARAVSLSPYLRFGCVSIRKTYWDICDTYKR 0 0 IKKVEAPKEIVCQLYWREYFYIMSIDNINFDKIENNPYCLKINWQYNEEFLKKWEM 0 0 GQTGYPWIDAIMNQLRFEGWNHHVGRHAVSCFLTRGDLWVSWEDGLK 0 0 LFLKYQLDADWSVCAGNWMWVSSSAFEKALQCPTCYSPVMYGMRMDKNGDFVKTYVPVLKDMPL 2 1 KYLFCPWKAPLEIQEKANCIIGKDYPEPIVMHRDASKQNMAKMYKVKEHLLHQ 1 2 DVPHCGPTNETEVWKFAWLPPIEHHDLAHNI* 0 >CRY1B_craGig Crassostrea gigas (oyster) GQ415324 HS189569 mollusc 0 GLRLHDNPSLIDGLSECDRFYPVFIFDGEVA 1 2 GTKTAGYNRFRFLLECLQDLDKNLKAAGTRLYCFQGQPTDILERLIE 0 0 EWGVTKVTFEADPEPIWQERDRLVRELLDKKNVQCVEKVSHTLWDPYE 2 1 IIENNGGSPPLTFSLFNLVTSTIGPPPRPVEDPDFTDISLPVSQNHDKQFGIPSLEDL 1 2 NVRPECEEQNKRLVEWLGGESKALELLAIRMKHEE 0 0 KAYENGYVMPNQYHPDLLSPPLSLSAHLRFGCLSVRKFYWSIHDKFEE 0 0 VKPSMGAPVSLSAQLMWREYFYTMAINNINYDKMETNPICLNIPWYDNPEHEEKWTQ 0 0 GETGYPWIDAIMKQLRYEGWVHHVARHAVSCFLTRGDLWLNWEVGLK 0 0 VFYKYLLDADWSVCAGNWMWVSSSAFEKVLQCPNCFCPVRYGKRMDPSGEYVR 2 1 RYLPVLKDMPLRYLFEPWKAPLPVQQKAKCIVDYPKPMVDHQKASKECIQNMKAVKDALIGK 1 2 EIPHCAPSEEIEARRFSWLP* 0 >CRY1B_rudPhi Ruditapes philippinarum (clam) JO113369 mollusc 1 RYLYEPWKAPKAVQEKANCVVGVDYPYPIVEHNKASKECYNLMMEVKNKLVQQ 2 1 GKDLEHCRPTNVEEVRMFVWMPGAHKGACGQEVPLDDKELCDGLSIM* 0 >CRY1B_vilLie Villosa lienosa (mussel) JR510441 transcript assembly mollusc fragment 0 1 2 GTSTAAFDRMRFLLECLHDLDESFKKFGGRLFVFHGKPKAIFTQLFE 0 0 EWGVTKLTFEQDPEPVRKARDNEIKQLCYEKGVTCIECISHTLWDPKQ 2 1 IIEANEGCPPLTYEIFRQVTSVLGPPPKPLDIPDFSKIKIPVQHNHAELYGIPCMAFL 1 2 GINPDCQEQLQRVHEWVG GETKGQELLGVrIKVERK 0 0 AFEEGIALPNQVHPDLLGTPRTLSPHLRFGCISVRKFYWCLYSQYCEV 0 0 0 0 YPGETPPHSIYGQLLWIHHVCCHAVSCFLTHRDLWISWEEEIK 0 0 2 1 1 2 * 0 >CRY1B_lymSta Lymnaea stagnalis (snail) ES576734 mollusc 0 MSDSERKSHVTVHWFRHGLRLHDNPALLDGLEDCREFYPVFIFDGSVA 2 1 GTSNVALPRMQFLLETLKDLDDNLKGLGTRLYVFKGDPVEVFDKLIE 0 0 EWGVTRITFEQDPEPVWQERDNSVKDLCAKQGVEWIERVSHTLWDPQL 2 1 IIKENGGSPPLTYAMFCQVTEIVGQPPTPRDDPDFEGITLPVCEDHDKKYHLPTLAQL 1 2 GVESESDQQTSPYCRYLGGESKALKLLATRLESERR 0 0 AFSLGQSLPNQLFPNLSGMPMSLSPHLRFG* 0 >CRY1B_plaDum Platynereis dumerilii (clam_worm) GU322429 annelid mRNA fragment 0 DGEVA 1 2 GTKLVSYPRMKFLLECLKDLDDSLKKHGGRLYVVKGPSDVVIKQLIE 0 0 EWGVTRVTCEIDPEPIWQPRDKAVKDLCATKGVKWFDYNSHLLWDPKAVCDANGGRPPHTYKLFC 0 0 QVTDLLGKPETPHPDPDFSHVQMPVSDDFDDKFGLPTLKEL 1 2 GCEPECEEQEKPFNKWQGGETGALELLETRLMIERTAYKA 2 1 GYIMPNQYIPDLVGPPRSMSPHLRFGALSIRK 2 1 FYWDLHNNYAEVCGGEWLGALT 1 2 AQLVWREYFYCMSYGNPSFDKMEGNPICLQIPWYKDEEALEKWKQGQ 2 1 TGFPWIDACMRQLRYEGWMHHVGRHAVACFLTRGDLWISWVDGLEAFYKYMLDGDW* 0 >CRY1B_dapPul Daphnia pulex (water_flea) ACJG01002273 FE370447 FE356368 crustacean 0 MNVLWFRRGLRIHDNPALLSALENSKDFIALFVFDTTFQ 1 2 DPGYKPYHMNGFLLECLQDLNESLESVGTKLHVFQGCPLEVFRHLHNIKPINKLCFIQDCEPIFHERDIAAK 1 2 DLCSELDIEVYEHVAHTLWDPMDIIASNGGTPPLTYEMFV 2 0 HVAMSVGDPPKPVADPEWKGVKFLTLEPFESNRFT 0 0 LFGGVPTLSQIGVPENPVGCRGRRIF 1 2 ETNALKHFAIRLQAEETAFRSGFYQPNQARPDLLGPPLSLSAAISVGAISVRL 2 1 FYWRIHEIFDKVNRGNPPAWLGITGQIIWRDYFYAMSRMNPKFDKEVDNPICLQIPWVENEEFFEKWKNGQTGYPFIDAGMRQLNQE 1 2 GWMHHSVRNAVAMFLTRGDLWLNWDIGAEYMANQLVDSDWSVNSGNWMWVSSSAFERLLDCSVCINSVLYGKRLEPSGDYIRRYVPELANFEFEYIHEPWKAPINIQRTANCII 1 2 GQDYPAQMVVHEEVLPRNLEWMKEFRQKFKETPAHCQPSSNSEVYKFFCLPDDSLPF* 0 >CRY1B_diaNig Dianemobius nigrofasciatus (cricket) AB291231 orthoptera 0 MDNRSNVHKKVAVHRFRHGLRLHDNPALLDAVKDCDAFLPIFIFDGESAGTKLVGYNRMKFLLESLQDIDSQLKKYGGNLYLFHGTPLCVFQYISQTIGLHK LCFEQDCEPIWQHRDDLVKKFCKENGIKCIERVSHTLWNPHDVIKTNGGIPPLTFEMFVHTVSVIGPPPRPVEDVEWSVVNFGVLPMSSIPSDIKVFKNFPTPEDFGISSEVGNMNR IIQWIGGESQALRHLQERLKVEENAFREGYCLPNQARPDLLGPPTSQSAALRFGCLSVRKFYWSIQDMYSSICGPSPNQNITSQLIWREYFYTMSVGNEYYAEMDRNPICLNI PWKNDYGSDFNKWKEGKTGYPFIDAIMRQLIQEGWIHHVARNAVACFLTRGDLWISWEEGLNFFLQYLLDADWSVCAGNWMWVSSSAFEQLLDCSHCMCPVNYGRRLDPWGQYI KRYIPELKNYPVEYLYEPWKAPLHVQETAGCIVGKDYPERIIDHQIASEKNRSYMDEIRNRLMNPPPHCRPSSEKETRQFMWFPDDCSEHSSQ* 0 >CRY1B_acyPis Acyrthosiphon pisum (aphid) NM_001171061 ABLF02032292 HP303737 hemiptera 0 MTVAVHWFRNGLRLHDNPALIEAHNNAEKLITLFIFDETTF 1 2 NPKWYGYNPMRFLLdSLIDLNNNLALVGGRLYILQGNPVNIFKMIKEKIGLHFITYEQ 0 0 DCAHLGRTRDEKVKSFCDENDIKCIETVSHTLWNPKSIIEKNGGVPPFTFKQFQ 0 0 NTAKQIGHPPSPVGNVDWLSVIFEELPASIQDEFK 0 0 NLHNPTPETFGIYPEVPENLTSTYRWYGGETRALEQLKERLEYEKEAFVNGFYLPNQVNPDLLSPPSSLSAALRYGCLSIRK 2 1 FYWELTKLFINKFEGDLLPMYSVTSQLIWRDYFYTMSIDNKNFGQIEDNPACISIPWNDIKIPENKKMLECWKTGKTGYPFIDAGMRQLMQ 0 0 EGWIHHVVRNSLASFLTRGDLWISWVEGLNHFMKYLLDADFSVCSGNWIWVSSSTFEQLLDCPLCVCPVSYGLRLDPSGEYIK 2 1 RYVPELKNMPAQFLYEPWKCPESVQKQVGCIIGKDYPNCIVDHTIASKGNRK 0 0 KMLALRVSMTNENRVPHCCPSDREEVQKFMYLPDECMQQLLPLENQDSKAYDIYKSH* 0 >CRY1B_danPle Danaus plexippus (butterfly) AY860425 AGBW01012954 lepidoptera 0 MLGGNVIWFRHGLRLHDNPSLHSALEDASSPFFPIFIFDGETA 1 2 GTKMVGYNRMRYLLEALNDLDQQFRKYGGKLLMIKGRPDLIFRRLWEEF 0 0 GIRTLCFEQDCEPIWRPRDASVRALCRDIGVSCREHVAHTLWNPDTVIKANGGIPPLTYQMFL 0 0 HTVEIIGNPPRPVDDVDLNGVNFGSLPESFYREFVVFDK 0 0 APKPEDLGVFLENEDIRMIRWVGGETAALKQMQERLAVEYETFCR 2 1 GSYLPTHGNPDLLGPPISLSPALRFGCLSVRR 2 1 FYWSLQDLFQQVHQGRLASTQFIT 1 2 GQLIWREYFYTMSVNNPNYAQMSGNPICLDIPWKEPENDELQR 2 1 WKEGRTGFPFVDAAMRQLRTEGWLHHVVRNTVASFLTRGTLWLSWEHGLQHFLKYLLDADW 2 1 SVCAGNWMWVSSSAFEALLDSGECACPVRLGRRLEPTGHYVRRYVPELARMPGEYIYEPWRAPLEVQEAAGCVIGRDYPAPVVDHTAAAARNRANMQELRRLLEKAPPHCCPSSEDEVRQFMWLGDDSQPELTTT* 0 >CRY1B_bomMor Bombyx mori (silkworm) NM_001195699 wrong BABH01015108 moth lepidoptera 0 MLGGSVLWFRHGLRLHDNPSLHSALEETSGPFFPIFIFDGETA 1 2 GTKVVGYNRMRYLLEALDDLDKQFKKYGGRLLLVKGKPSAVFRRLWEEF 0 0 GIRKLCFEQDCEPVWRPRDESVKTACREIGVTCREHVSHTLWEPDTVIKANGGIPPLTYQMFL 0 0 HTVATIGDPPRPVDNAKLRGIKFGTLPLCFYEEFTVYDK 0 0 VPNPEDLGVFLENEDIRMIRWVGGETAALKQMQHRLAVEYETFCR 2 1 GSYLPTHGSPDLLGPPISLSPALRFGCLSVRK 2 1 FYWSLQDLFQQVHQGSLCSTQYIT 1 2 GQLIWREYFYTMSVNNPHYGQMTDNPICLDIPWKSPEGDELER 2 1 WASGRTGFPFVDAAMRQLRLEGWLHHAVRNTVASFLTRGTLWLSWEHGLAHFLKYLLDADW 2 1 SVCAGNWMWVSSSAFEALLDSGECACPVRLGQRLDPSGEYVRRYVPELARVPTEYI 2 1 YEPWKAPLDVQERANCIIGKDYPAPVVNHIVAAQRNRNAME 0 0 ELRMLLEKAPPHCCPSSEDEIRQFMWLNE* 0 >CRY1B_mamBra Mamestra brassicae (moth) AY947639 Glossata lepidoptera 0 MLGGSVLWFRHGLRLHDNPSLHSALEEKGFPFFPVFIFDGETAGTKVVGYNRMRYLLEALEDLDNQLKKHGGRLIMIKGKPNVVFRRLWEEFGIRRLCFEQD CEPVWRARDDSVKSACKEIGVVCKEHVSHTLWEPDTVIKANGGIPPLTYQMFLHTVATIGDPPRPVSDIDFTGVKFGSLPESFYHEFTVYDKTPKPEDLGVFLENEDIRMIRWVGG ETTALKQMQQRLAVEYETFLRGSYLPTHGNPDLLGPPISLSPALRFGCLSVRSFYWSVQDLFRQVHQGRLATQSASHFITGQLIWREYFYTMSVNNPNYGQMAGNPICLDIPWKEP EGDELQRWVEGRTGFPFVDAAMRQLRTEGWLHHAARNTVASFLTRGTLWLSWEHGLNHFLKYLLDADWSVCAGNWMWVSSSAFEALLDSGECACPVRLGQRLDPSGEYVRRYVPEL ARMPVQYIYEPWKAPIDIQERATCIIGKDYPAPVVNHLVAAQRNKNAMKWLGRSVAGRLDKEKWFELVGELRHFLQKAPPHCCPSSEDEIRQFMWLNE* 0 >CRY1B_helArm Helicoverpa armigera (cotton_bollworm) JN997418 moth lepidoptera 0 MLGGSVLWFRHGLRLHDNPSLHSALEEKGFPFFPIFIFDGETAGTKLVGYNRMRYLLEALDDLDSQFKKFGGRLIMLKGKPNVVFRRLWEEFGIRKLCFEQD CEPVWRARDDSVKSACKEIGVVCREHVSHTLWEPETVIKANGGIPPLTYQMFLHTVATIGDPPRPVPNIDFTGVKFGSLPECFYQEFTVYDKTPKPEDLGVFLENEDIRMIRWVGG ETTALKQMQQRLSVEYETFLRGSYLPTHGNPDLLGPPISLSPALRFGCLSVRSFYWAVQDLFRQVHQGRLTTNSASHFITGQLIWREYFYTMSVNNPNYGQMAGNPICLDIPWKNP EGDELQRWVEGRTGFPFVDAAMRQLRTEGWLHHAARNTVASFLTRGTLWLSWEHGLNHFLKYLLDADWSVCAGNWMWVSSSAFEALLDSGECACPVRLGQRLDPSGEYVRRYVPEL ARMPVEYIYEPWKAPIDVQERATCVIGKDYPAPVVNHLVAAQRNKNAMKELRHMLQKAPPHCCPSSEDEIRQFMWLNE* 0 >CRY1B_droMel Drosophila melanogaster (fruit_fly) AB019389 diptera PubMed:22080955 PDB:3TVS 0 MATRGANVIWFRHGLRLHDNPALLAALADKDQGIALIPVFIFDGESA 1 2 GTKNVGYNRMRFLLDSLQDIDDQLQAATDGRGRLLVFEGEPAYIFRRLHEQVRLHRICIEQDCEPIWNERDESIRSLCRELNIDFVEKVSHTLWDPQLVIETNGGIPPLTYQMFL 0 0 HTVQIIGLPPRPTADARLEDATFVELDPEFCRSLKLFEQLPTPEHFNVYGDNMGFLAKINWRGGETQALlLLDERLKVEQHAFERGFYLPNQALPNIHDSPKSMSAHLRFGCLS VRRFYWSVHDLFKNVQLRACVRGVQMTGGAHITGQLIWREYFYTMSVNNPNYDRMEGNdICLSIPWAKPNEnLLQSWRLGQTGFPLIDGAMRQLLAEGWLHHTLRNTVATFLTRGG LWQSWEHGLQHFLKYLLDADWSVCAGNWMWVSSSAFERLLDSSLVTCPVALAKRLDPDGTYIKQYVPELMNVPKEFV 2 1 HEPWRMSAEQQEQYECLIGVHYPERIIDLSMAVKRNMLAMKSLRNSLITPPPHCRPSNEEEVRQFFWLADVVV* 0 >CRY1B_anoGam Anopheles gambiae (mosquito) DQ219482 diptera PubMed:16332522 0 MTINNILWFRHGLRLHDNPSLLEALKSDCVNQSSEAVKLFPIFIFDGESA 1 2 GTRIVGYNRMKFLLESLADLDRQFRDLGGQLLVFRGDSVTVLRRLFEELNIKKLCYEQDCEPIWKERDDAVAKLCRTMDVRCVENVSHTLWNPIEVIQTNGDIPPLTYQMFL 0 0 HTVNIIGDPPRPVGAPNFEYVEFGRVPALLASELKLCQQMPAPDDFGIHYDGNARI AFQKWIGGETRALEALGARLKQEEEAFREGYYLPTQAKPEILGPATSMSAALRFGCLSVRMFYWCVHDLFAKVQSNSQFKYPGGHHITGQLIWREYFYTMSVQNPHYGEMERNPIC LNIPWYKPEDDSLTRWKEGRTGFPMIDAAMRQLLAEGWLHHILRNITAT 2 1 FLTRGGLWLSWEEGLQHFLKYLLDADWSVCAGNWMWVSSSAFERLLDSSKCTCPIALARRLDPKGDYVKRYLPELANYPAQFV 2 1 HEPWKASREQQIEYGCVIGEKYPAPMVDLAIVSKRNAHTMASLREKLVDGGSTPPHCRPSDIEEIRQFFWLADDAATEA* 0 >CRY1B_neoBul Neobellieria bullata (fleshfly) FJ373353 diptera 0 MSLNIIWFRHGLRLHDNPALLDAISDKDEGIELLPIFIFDGESAGTQSVGYNRLKFLLDSLEDINNQLRSVSLSLGRLYIMKGNPVQIFRSLHEQRGIKKLC FEQDCEPIWNRRDNAVKNLCHDLGITCIERISHTLWDPKKVINTNGGIPPLTYQMFLHTVQIIGLPPRPVPDPDWHNVKFVKLSEKLIKDLGVFLESPTPEDFSVYPDSLSYLAKV KWIGGETQALLHLDQRLKVEESAFKCGYYLPNQAKPNILESPKSMSAHLRFGCLSVRKFYWDVHDLFKDVQQQAECLGMLMSGGAHITGQLIWREYFYTMSVNNPNYDRMEGNEIC LNIPWAETNHIQLQRWTQGKTGFPLIDAAMRQLLAEGWLHHTLRNTVATFLTRGGLWQNWEHGLRYFLKYLLDADWSVCAGNWMWVSSSAFERLLDSSLVTCPIALAKRLDPMGQY IKQYVPELANVPKEYIHEPWRMPMNIQEDSDCVIGIHYPERLIDLNVACKRNTIAMRTLRNALIAEGAPDNGPPHCRPSNEEEIRNFFWLAD* 0 >CRY1B_bacCuc Bactrocera cucurbitae (melon_fly) AB517608 diptera 0 MTKRANVLWFrHGLRLHDNPALLEAISDKSEGIALIPLFIFDGESAGTKTVGYNRMSFLLNSLADIDKQLKAIRGASDISGKLYLFQGNPATVFRRLSEYYR LNKICFEQDCEPIWNRRDDSVRSLCNDLDIEAVEKVSHTLWDPRTVISTNGGIPPLTYQMFLHTVEIIGVPPRPVEDPDWDGVEFLKLTDNMLMELNAFWRFPTPEDFNVYPDNVS YVAKVKWHGGEQQALLHLDERLKVEERAFKNGYYLPNQANPNILESPKSMSAHLRFGCLSVRRFYWSVHDLFKHVQIEALRQRVHMAGGEHITGQLIWREYFYTMSVNNPYYDRME GNAICLNIPWAAPNKEQLQSWRSGQTGFPLIDAAMRQLLAEGWLHHTLRNTVATFLTRGALWQSWEHGLRHFLKYLLDADWSVCAGNWMWVSSSAFERLLDSSLVSCPIAFSKRLD PKGEYIRQYVPELANVPQEYIHEPWRMPQELQENCECVIGVQYPERIVDLANVSKRNVxAMQTLRQSLIAGGAPDEGPPHCRPSNEEEVHQFFWLVE* 0
>CRY1A_dapPul Daphnia pulex (water_flea) FE418063 FE356487 ACJG01001137 crustacean 0 MYRQQKNIMSGYDSEPREKQVVHWFRKGLRLHDNPSLKDGLKGCSTYRCIFILDPWFAGSSNVDINKWR 2 1 FLLESLEDLDQNLRKLNSRLFVIRGQPAGVLPKLFK 0 0 EWETTCLTFEEDPEPFGRVRDQNIITMCKDFNIEVITRASHTLYHPQK 2 1 IIEKNGGKAPLTYRQFQNIIASVDAPPPPESDITFESIGRGYTPMDESMDDRFSVPTLEEL 1 2 GFDTDGLMPAVWHGGETEALTRLERHLERK 0 0 AWVASFGRPKMTPQSLLASQTGLSPYLRFGCLSVRLFHQQLTNLYKKIKKAQPPLSLHGQVLWREFFYCAATNNPNFDKMIGNPICVQIPWDSNAEALAKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGTWMWLSCSSFFHQFFHCYCPVRFGREVDPNGDFIK 2 1 KYQPVLKNFPLQYIHEPWNAPESVQRAAKCVIGKDYPLPMVNHLEVSQLNIERMKQVYQRLTQYRGT 1 2 GLMSHSPQSDNGIIINVGNKNKNENSHAKQFRTDELRQNAVQRNQSNLN* 0 >CRY1A_eupSup Euphausia superba (krill) FM200054 contig crustacean 0 MTRGGEKHVVHWFRKGLRLHDNPTLKAGLKGATTFRGIFIIDPWFAGSSNVGINKWR FLLQCLEDLDTTLRKLNGRLFVVRGQPAHVLPQLFK TWGTTCLTFEKDPEPFGKVRDANITHIAREMGIQVIIKTSHTLYKLEK 2 1 IISKLPLTYKTFQNVLSTMEPPPLPASPVTVRDVGDAFTPIDEDHDEKYGVPTLEEL 1 2 GFETENLAPSIWKGGESEALARLEHHLERKAWVASFGRPKMTPQSLYPSRTGLSPYLRFGCLSARRFFAELNDLYRK 0 0 IKKSPAPLSLHGQLLWREFYYTAATNNPKFDHMEGNPICVQIPWDKNAEALAKWAH 0 0 GRTGFPWIDAIMSQLRKEGWIHNVARHAVACFLTRGDLWVSWEEGMK 0 0 VFDELLLDADWSVNAGSWMWLSCSSFFQQFFHCYCPVRYGRKADPNGDYIR 2 1 TYLPVLKNFPTKYIHEPWTAPENVQKQSKCVIGRDYPMPMVDHVKQSQANLMRMKQVYQQLSHYRVTPPKITSLLKSCPPPLPVICDKSKNTKNNKNEITRESNYTQQMQPA* 0 >CRY1A_pedHum Pediculus humanus (louse) XM_002430500=wrong AAZO01005932 phthiraptera very similar intron pattern to vertebrate but lacks last 4 exons 0 MSDNSSQNSGKHTVHWFRKGLRLHDNPSLREGIKNSVTFRCVFVIDPWFAGSSNVGINKWR 2 1 FLLQCLEDLDRSLRKLNSRLFVIRGQPADTLPKLFK EWGTTNLTFEEDPEPFGRVRDLNIMAMCKELGISVVSKSSHTLYKLEH 2 1 IIEKNGGNPPLTYHQFQTIIANIGPPPEPEETVTANLLEGSQTPLSDDHDNKYGVPTLEEL 1 2 GFETDKLKPTVWSGGESEALARLERHLERK AWVASFGHPKMTPQSLLASQTGLSPYLRFGCLSTRLFYYQLVELYKK 0 0 IKKAMPPLSLHGQLLWREFFYCAATKNPNFDKMNENPICVQIPWDENAQALAKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFDELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVRFGRKADPNGDYIR KYLPILKNFPTKYIHEPWLAPETEQRAAKCIIGKDFPLPMVNHSIVSKINIERMKQVYRQLSKYRSQGT* 0 >CRY1A_acyPis Acyrthosiphon pisum (aphid) NM_001171102 ABLF02035823 hemiptera cry2-2 PubMed:20482645 end uncertain 0 MQNKHTVHWFRKGLRIHDNPSLREGLINAKTFRCIFILDPWFAGASNVGINKWR 2 1 FLLQCLSDLDNSLKKLNSRLFVIKGQPAEALPKLFR 0 0 QWGTTNFTFEEDPEPFGRVRDQNIKVMCSEMGISVITRCSHTLYQLDK 2 1 IINVNGGKAPLTYHLFQKLLECIDPPERAVPSIDKEFLGNAFTPIKYDHDEIFGVPTLEEL 1 2 GFKEINNITRHVWVGGETEALIRLQCHLERKAFIASFGKPKMTPQSLVASPTGLAPYLKFGCLSTRLFFSELNELYKK 0 0 IRKSQPPLSLHGQLLWRDFFYCASTNNPNFDRMVGNPICVQIPWDKNPRALSKWAN 0 0 GQTGYPWIDAIMIQLRQEGWIHCIARHAVACFLTRGDLWLSWEEGMK 0 0 VFDELLLDADWSVNAGYWMWYSCSSFYQEFIHCYCPVRFGRKVDPNGDYIR 2 1 RYIPALNNMPNQYIHEPWLAPESIQFSANCIIGIDYPLPIVNHINASKINLERMKLAYQQLSN 1 2 1 2 * 0 >CRY1A_ripPed Riptortus pedestris (bean_bug) AB379863 hemiptera PubMed:18547745 0 MSEKHTVHWFRKGLRLHDNPSLRHGLNGAKTFRCIFILDPWFANASNVGINKWR 2 1 FLLQCLEDLDRSLMKLNSRLFVIRGQPADILPKLLKEWGTTCLTFEEDPEPFGRVRDQNIMAMCRGMNITVISLVAHTLYKLEC 2 1 IIERNGGRAPLTYHQFQSVVASMESPPLPVPPVTSTVVGDAVSPISDDHDEKYGVPTLEEL 1 2 GFDTEGLLPGVWQGGESEALSRLERHLERKAWVASFGKPKMTPQSLLPSQTGLSPYLRFGCLSTRLFYYQLNDLYRK 0 0 IKKAVPPLSLHGQVLWREFFYCAATKNPNFDRMIGNPICVQVPWDKNPEALAKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVRFGRKADPNGDYIR 2 1 KYLPVLKNMPTKYIHEPWNCPESVQKAAKCTIGVDYPLPMLNHSVVSKHNIERMKQVYLQLMKYRQP 1 2 GIPPSLNSD 1 2 AALEKKKKNEDNNFDDNIFATPSPTFKGTISK* 0 >CRY1A_triCas Tribolium castaneum (flour_beetle) AAJJ01000096 coleopetera 0 MSGVVGDAGRASKGQDKHMVHWFRRGLRLHDNPSLREGLKGARTFRCVFVLDPWFAGSSNVGINKWR 2 1 FLLQCLEDLDRSLRKLSRLFVIRGQPADALPKLFK EWGTTALTFEEDPEPFGGVRDHNLTTLCQELGISVVQKVSHTLYHLQD IIDRNGGRAPLTYHQFLAIIACMGPPPQPEPPVTFNSLNGAHTPLTDDHDEKYGVPTLEEL GFDTEGRLPPVWQGGESEALARLERHLERK AWVASFGRPKMTPQSLLPSQTGLSPYLRFGCLSTRLFYYQLTDLYKK IKKAFPPLSLHGQLLWREFFYCAATKNPNFDKMIGNPICVQIPWDKNAEALAKWAN 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWLSWEEGMK 0 0 VFEELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPIKFGRKADPNGDYIR 2 1 KYLPVLKNMPSQYIHEPWTAPEQVQRAAKCIIGKDYSLPMVNHAAASRVNIQRMKQVYQQLSNYRNVDNSAKFKDGFQEQPNVVTVGNPSRK* 0 >CRY1A_bomImp Bombus impatiens (bumble_bee) EF110521 AEQM02008194 hymenoptera PubMed:17244599 0 MTGSRSSEINPDVTVRGEGGKHTVHWFRKGLRLHDNPSLREGLTGATTFRCVFVLDPWFAGSTNVGINKWR 2 1 FLLQCLEDLDCSLRKLNSRLFVIRGQPADALPKLFK EWGTTNLTFEEDPEPFGRVRDHNISALCKELGISVVQKVSHTLYKLDE 2 1 IIERNGGKPPLTYHQFQNVVASMDPPEPSVPTVTSACIGSAYTPLKEDHDDHYGVPTLEEL 1 2 GFDTEGLLPPVWVGGESEALARLERHLERK AWVASFGRPKMTPQSLLPSQTGLSPYLRFGCLSTRLFYYQLTDLYKK 0 0 IKKAVPPLSLHGQLLWREFFYCAATKNPNFDRMQGNPICVQIPWDKNVEALAKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFDELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVRFGRKADPNGDYIR 2 1 RYLPILKNFPTRYIHEPWNAPLSVQRAAKCIIGKDYSLPMVNHSKSSRINIERMKQVYQQLNKYRGN 1 2 GASLKGETV 1 2 GLLNALPPPSVKENEEEKKKTKQSPPPPENQSKMEVLAKTTQHQQQHHQHQ* 0 >CRY1A_apiMel Apis mellifera (bee) NM_001083630 AADG06001305 hymenoptera 0 MTGSRSSEINPKEGLYDEGGKHTVHWFRKGLRLHDNPSLREGLAGASTFRCVFVLDPWFAGSTNIGINKWR 2 1 FLLQCLEDLDCSLRKLNSRLFVIRGQPADALPKLFK EWGTTNLTFEEDPEPFGRVRDHNISALCKELGISVVQKVSHTLYKLDE 2 1 IIERNGDKPPLTYHQFQTVVASMDPPEPPVPTVTSACVGSAYTPLKEDHDDHYGVPTLEEL 1 2 GFDTEGLLPPVWVGGESEALARLERHLERK AWVASFGRPKMTPQSLLPSQTGLSPYLRFGCLSTRLFYYQLTDLYKK 0 0 IKKAVPPLSLHGQLLWREFFYCAATKNPNFDRMQGNPICVQIPWDKNVEALAKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFDELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVRFGRKADPNGDYIR 2 1 RYLPVLKNFPTRYIHEPWNAPLNVQRAAKCIIGKDYSLPMVNHSKSSRINIERMKQVYQQLNKYRGN 1 2 GVSLKGETV 1 2 GLLNALPPSSMKETEEEKKKTKQSLSPSENQSKMEILPKTTQRQHHH* 0 >CRY1A_attCep Atta cephalotes (ant) ADTU01021771 hymenoptera 0 MGQAVTSGVRGDGGKHTVHWFRKGLRLHDNPSLKEGLAGASTFRCVFVLDPWFAGSTNVGINKWR 2 1 FLLQCLEDLDCSLRKLNSRLFVIRGQPADALPKLFKEWGTTDLTFEEDPEPFGRVRDHNISALCKELGISVVQRVSHTLYRLDE 2 1 IIERNSGKPPLTYHQFQNVVAGMDPPEPPVPTVTAACIGSAYTPLKDDHDDHYGVPTLEEL 1 2 GFDTESLLPPVWVGGESEALARLERHLERKAWVASFGRPKMTPQSLLPSQTGLSPYLRFGCLSTRLFYYQLTDLYKK 0 0 IKKAVPPLSLHGQLLWREFFYCAATKNPNFDKMQGNPICVQIPWDKNVEALAKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFDELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVRFGRKADPNGDYIR 2 1 RYLPVLKNFPTRYIHEPWNAPLSIQHAAKCIIGKEYSLPMVNHNKSSRINIERMKQVYQQLNKYRDN 1 2 GTSFKGENI 1 2 GLLNALLASPTKDGDEEKRKQDSPNRENEQKMEAINSPTQQQ* 0 >CRY1A_exoRob Exoneura robusta (bee) HP928681 hymenoptera fragment 0 VFDELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVRFGRKADPNGDYIR 2 1 RYLPVLKNFPTRYIHEPWNAPLSVQRTAKCIVGKDYSLPMVNHSKSSRINIERMKQVYQQLNKYRGN 1 2 GTSLKGETV 1 2 GLLNSLPPPPVKKTEEEKKKTEQSPPRAENQMNMEALVKTTQHQQQHHHRQHQQRH* 0 >CRY1A_nylPub Nylanderia pubens (crazy_ant) JP792144 hymenoptera fragment 0 iKKAVPPLSLHGQLLWREFFYCAATKNPNFDRMQGNPICVQIPWDKNVEALAKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFDELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVRFGRKADPNGDYIR 2 1 RYLPVLKNFPTRYIHEPWNAPLSIQHTAKCIIGKEYSLPMVNHSKNSRINIERMKQVYQQLNKYRGN 1 2 GASFKGENI 1 2 GLLNALLAPPTDAEEEQRKQDSANRENEQKMETINNStqqq* 0 >CRY1A_nasVit Nasonia vitripennis (wasp) XM_001606355 AAZX01001169 hymenoptera N-term shortened 0 mGKKHTVHWFRKGLRLHDNPSLREGLAGASTFRCVFVLDPWFAGSANVSINKWR 2 1 FLLQCLEDLDRSLHQLNSRLFVIRGQPADALPKLFREWGTTSLTFEEDPEPYGRVRDENITTLCKELGITVVQRVSHTLYKLDE 2 1 IIEKNGGKPPLTYHQFQNVIARMDPPEYPAAAVTAACIGSAYTPLKDDHDDFFGVPTLEEL 1 2 GFDTEGLMAPVWVGGETEALARLERHLERKAWVASFGRPKMTPQSLLPSQTGLSPYLRFGCLSTRLFYYQLADLYKK 0 0 IKKTIPPLSLHGQLLWREFFYCAATNNPNFDRMHGNPICVQIPWDKNVVALSKWAN 0 0 GQTGFPWIDAIMTQLREEGWIHQLARHAVACFLTRGDLWISWEEGMK 0 0 VFDELLLDADWSINAGMWMWLSCSSFFQQFFHCYCPVRFGRKADPNGDYIR 2 1 RYLPVLKNFPTRYIHEPWNAPLSVQRAAKCIIGQEYALPMVNHSKSSRINVERMKRVYQQLSKYRAN 1 2 GVSLKGETI 1 2 GLLSIVPTTPSLPFDQQQQQQDDDKKQQQNSLNVDNSPNNDCPASHLHQQQLQQHVHQHQYH* 0 >CRY1A_antPer Antheraea pernyi (silkmoth) EF117812 lepidoptera PubMed:17244599 dropped long C-terminus 0 MSAAAETLSAPRARSHVPAASLTSAPTPRRPHTKHTVHWFRKGLRLHDNPALREGLVNATTFRCVFIIDPWFASSSNVGINKWR 2 1 FLLQCLEDLDSSLKKLNSRLFVVRGQPADALPKLFR EWGTTALTFEEDPEPYGRVRDHNITTKCREVGINVISRVSHTLYKLDK 2 1 IIERNGGKAPLTYHQFQALIASMPPPQPAEAPISIETLNGAKTPVSVDHDDRFGVPTLEEL 1 2 GFETEDLKPPMWMGGESEALARLDRHLERK AWVASFGRPKMTPQSLLASQTGLSPYLRFGCLSTRLFYYQLTELYKK 0 0 IKRVRPPLSLHGQILWREFFYCAATRNPNFDRMEGNPICVQIPWEKNQEALSKWAN 0 0 GQTGYPWIDAIMIQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFDELMLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVRFGRKTDPNGDFIR 2 1 KYIPALKNMPTRYIHEPWVAPESVQQAAQCVVGCDYPLPMLDHTKASQINLERIKQVYAQLAKYKPQ 1 2 VTLQTVQRPNVMQSSPSPTSIIASINQSNLLCSTTPESQSAATQMIYKDPS* 0 >CRY1A_anoGam Anopheles gambiae (mosquito) DQ219483 diptera dropped long C-terminus 0 MRDKHTVHWFRKGLRLHDNPALREGLRGARTFRCVFIIDPWFAGSSNVGINKWR 2 1 FLLQCLDDLDRNLRKLNSRLFVIRGQPADALPKLFKEWGTTCLTFEEDPEPFGRVRDHNISEMCKELGIEVISAASHTLYNLER 2 1 IIEKNGGRAPLTYHQFQAIIASMDAPPQPEAAITLDVIGNANTPQYDDHDDKYGVPTLEELGFETEALRPPVWIGGETEA LARLERHLERKAWVASFGRPKMTPQSLLASQTGLSPYLRFGCLSTRLFYYQLTDLYKKIKKACPPLSLHGQLLWREFFYCAATKNPTFDKMAGNPICVQIPWDRNAEALAKWASGQ TGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMKVFEELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVKFGRKADPNGDYIRRYLPVLK 2 1 NFPTRFIHEPWNASESVQRAAKCLIGKDYPLPMVNHAIASRANMERIKQVYQHLAKYRTPSGGCYEGDCTEKGGSAIAGVMTAAKVQHMNTSSMNDSPSPTTILTSVNSSGNYMCRSNPPAQSDHNGKI* 0 >CRY1A_aedAeg Aedes aegypti (mosquito) XM_001655728 diptera dropped long C-terminus 0 MTKQHNLVHQQKTTNTATSTIGSGGTGTGTNSSIGTQKQQHQQKHTVHWFRKGLRLHDNPALREGLKDAVSFRCVFVIDPWFAGSSNVGINKWR 2 1 FLLQCLEDLDRNLRKLNSRLFVIRGQPADALPKLFK EWGTTCLTFEEDPEPFGKVRDHNISEMCKELGIEVISAVSHTLYKLER 2 1 IIEKNGGRAPLTYHQFQAIIASMDAPPQPEPAITLDTIANATTPQYEDHDDKYGVPTLEELGFETEGLKPPIWVGGETEALARLERHLERKAWVASFGRPKMTPQSLLASQTGLSPYLR FGCLSTRLFYYQLTDLYKKIKKACPPLSLHGQLLWREFFYCAATKNPNFDKMAGNPICVQIPWDRNAEALAKWASGQTGFPWIDAIMTQLREEGWIHHLARHAVACFLTRGDLWISWEEGMK VFEELLLDADWSVNAGMWMWLSCSSFFQQFFHCYCPVKFGRKADPNGDYIRRYLPVLK 2 1 NFPTRYIHEPWNAPETVQRTAKCIIGKEYPLPMVNHAIASRANMERIKQVYQQLAKYRSPSSSFDSECPEKGGSAIAGVMTAAKVQHMNASSLNDSPSPTTIMTNLNSSGNYMCRTIPPAQTDPNAKIITYHQ* 0 >CRY1_vilLie Villosa lienosa (mussel) JR505030 mollusc transcript assembly mollusc 0 MDEPPKKYVVHWFRKGLRLHDNPALCEAFKGASTFRCVYILDPWFAGVSQVGINKWR 2 1 FLLQCLEDLDSSLRKVNSRLFVIRGQPADVFPRLFK 0 0 EWQITSLSFEEDPEPFGKERDAAISAMAKEAGVEVIIRMSHTLFNLQK 2 1 IITENNGTPPLTFKRFQSILKTVGPPTKPVETVTLTTIGTARTPIENDHDDRYGVPSLEEL 1 2 GFDIDGLKPSVFQGGETEALLRLDRHLERK 0 0 AWVASFEKPKMTSQSLFPSQTTISPYLKFGCLSSRLFYWKLNDLYRR 0 0 VKKKSDPPLSLHGQLLWREFFYLAATNNPKFDRMVGNPICVQVPWDRNKEALAKWAEGKTGFPWIDAIMIQLREVGWIHHLARHSVACFLTRGDLWISWEEGMK 0 0 VFDELLLDADWSVNAGMWLWLSCSSFFQQFLNCYCPVGFGKRADPAGDFIR 2 1 HYIPQLKGFHPKYIYEPWTAPYEVQVAAKCIIGKDYPQPMVDHNEVSRQNMERMKQVYQVLAMRASG 1 2 VITKTLTDDTISKHPHSKISYITSCSNHISGNKPSKAAILLGGDSMNKHGHTSDEDNTGNSTN* 0 >CRY1_tetUrt Tetranychus urticae (spider-mite) CAEY01002034 chelicerate N-terminus uncertain 0 1 2 GLQGCDTFRCIYLIDPWFSTESNCGINKWR FLLQSLEDLDSNLHKLGSRLFVIRGQPAEVFPLLFK 1 2 EWMITHLSFEEDPEPFGKIRDAKIAKMCKEMNIEVICETSHTLYRLED IINHCSGEIPLTYLQFQNVISELGKPEEPAQAITHEVIGKAYTPVPRDHAKKYYTVLFQLIFD 1 2 GFTTQPNHRSVWVGGETEALQRLSRHLEHK AWVATFGKPKLTSLSLLCASQTGLGPYLR 2 1 FGCISPRLLYHQLSNLYSKLKKGNPPLTLCGQLLWREFFYCVATNNPNFDKMQNNPLCVQIAWNSNAEALNKWANGQTGYPWIDAIQTQLRREGWIHQIAKHATVCFLTRGILWISWEEGMK 0 0 VFDELLIDADWSVNAGTWMWLSCSSFFQRICHIYCPVSFGRKMDPQGDYIR RYLPVLRNIPNEYIHEPWKAPIKVQREANCKIGKHYPQPIVDHQEAMRINQERMKQVFRILIKGLDLKREDVQD* 0 >CRY1_aplCal Aplysia californica (sea_hare) scaffold_2275 mollusc small fragment 0 2 1 0 0 2 1 1 2 0 0 0 0 GMTGFPWIDAIMVQLKKEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFDEMLLDADWSVNAGMWMWLSCSAFFQQFFHCYCPVGFGKRADPSGDFV 2 1 HYLPVLKAMPTKYIYEPWTAPESVQKVAKCIVGKDYPMPMVVHSDVSRINLERMRQVYKKRLVLQSS 1 2 * 0
Cryptochrome CRY4 sequences
>CRY4_galGal Gallus gallus (chicken) NP_001034685 CRY4 PumMed:19663499 synteny: ADIPOR1 UBE2T CRY4 LRIF1 DRAM2 CEPT1 0 MRHRTIHLFRKGLRLHDNPALLAALQSSEVVYPVYILDRAFMTSSMHIGALRWHFLLQSLEDLRSSLRQLGSCLLVIQGEYESVVRDHVQKWNITQVTLDAEMEPFYKEMEANIRGLGEELGFQVLSLMGHSLYNTQR 2 1 ILELNGGTPPLTYKRFLRILSLLGDPEVPVRNPTAEDFQ 2 1 RCSPPELGLAECYGVPLPTDLKIPPESISPWRGGESEGLQRLEQHLADQ 0 0 GWVASFTKPKTVPNSLLPSTTGLSPYFSTGCLSVRSFFYRLSNIYAQ 0 0 AKHHSLPPVSLQGQLLWREFFYTVASATPNFTKMAGNPICLQIRWYEDAERLHKWKT 0 0 AQTGFPWIDAIMTQLRQEGWIHHLARHAAACFLTRGDLWISWEEGMK 0 0 VFEELLLDADYSINAGNWMWLSASAFFHHYTRIFCPVRFGRRTDPEGQYIR 2 1 KYLPILKNFPSKYIYEPWTASEEEQKQAGCII 12 GRDYPFPMVDHKEASDHNLQLMKQAREEQHRIAQLTR 1 2 DDADDPMEMKLKRDHSEESFTKTKAARMTEQT* 0 >CRY4_melGal Meleagris gallopavo (turkey) XM_003212851 0 MRHRTIHLFRKGLRLHDNPALLAALQSSEVVYPVYILDRAFMTSSMHIGALRWHFLLQSLEDLRSSLCQLGSCLLVIQGEYDAVVRDHVQKWNITQVTLDAEMEPFYKEMEANIRALGEELGFEVLSLMGHSLYNTQR 2 1 ILELNGGTPPLTYKRFLHILSLLGDPEMPIRNLTAEDFQ 2 1 RCSPPELCLAECYRVPLPTDLKIPPESISPWRGGESEGLQRLEQHLADR 0 0 GWVASFTKPKTVPNSLLPSTTGLSPYFSMGCLSVRSFFYRLSNIYAQ 0 0 AKHHSLPPVSLQGQLLWREFFYTVASATPNFTKMAGNPICLQIRWYEDAERLHRWKT 0 0 AQTGFPWIDAIMTQLRQEGWIHHLARHAAACFLTRGDLWISWEEGMK 0 0 VFEELLLDADYSINAGNWMWLSASAFFHHYTRIFCPVRFGRRTDPEGQYIR 2 1 KYLPILKNFPSKYIYEPWTASEEEQKQAGCII 12 GRDYPFPMVDHKEASDHNLQLMKQVREEQYRTAQLTR 1 2 DDADDPMEMKLKRDHSEENLTKTKAARMTKQT* 0 >CRY4_anaPla Anas platyrhynchos (duck) scaffold1663 0 MPHRTIHLFRKGLRLHDHPALLAALESSEVVYPVYILDRKFMTSVMHIGALRWHFLLQSLEDLQKNLCRLGSHLLVTQGEYEPVLRDHVQKWNITQVTLDAEMEPFYKEMEANIRCLGEELGFEVLSVVGHSLYDTKR 2 1 ILDLNDGTPPLTYKRFLHILSLLGDPEVPVRNLTAEDFQ 2 1 RCRPPDLDLAECYRVPLPADLKTSPEDISPWRGGETEGLQRLEQHLADQ 0 0 GWVASFTKPRTIPNSLLPSTTGLSPYFSMGCVSVRTFFYRLSNIYAQ 0 0 AKHHSLPPVSLQGQLLWREFFYTVASATPNFTKMAGNPICLQISWYEDAERLHKWKT 0 0 AQTGFPWIDAIMTQLRQEGWIHHLARHAAACFLTRGDLWISWEEGMK 0 0 VFEELLLDADYSINAGNWMWLSASAFFHQYTRIFCPVRFGKRTDPEGQYIR 2 1 KYLPVLKNFPSKYIYEPWTASEEEQKQAGCII 12 GKDYPFPMVDHKEASNHNLQLMKHVREEQYRTAQLTR 1 2 DDTDDPMEMKLKRDHSEENVTKAKTARMTEQT* 0 >CRY4_taeGut Taeniopygia guttata (finch) XM_002198497 0 MLHRTIHLFRKELRLHDNPVLLAALESSEALYPVYILDRAFLTSSMHIGALRWNFLLQSLEDLHKNLGQLGSCLLVIQGEYEIVLRDHIQKWNITQVTLDAEMEPFYKEMEANIQRLGAELGFEVLSLVSHSLYNTQR 2 1 ILDLNGGSPPLTYKRFLHILSLLGDPELPVRNLTAEDFQ 2 1 RCRAPEPGLAECYRVPLPVDLKISPESLSPWRGGETEGLRRLEQHLIDQ 0 0 GWVTSFAKPRTSPNSLLPSTTGLSPYFSMGCLSVRTFFYRLSNIYAQ 0 0 AKHHSLPPVSLQGQLLWREFFYTVASATPNFTQMAGNPICLQISWYKDAERLHKWKT 0 0 AKTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADYSINAGNWMWLSASAFFHQYTRIFCPVRFGKRTDPQGNYIR 2 1 KYLPILKNFPSKYIYEPWTASEEEQKLAGCII 12 GQDYPFPMVNHKEASDHNLQLMKQVREEQHRTVQLTR 1 2 DDADDPMEIRVKRDHSEENISKGKVARTTE* 0 >CRY4_pasDom Passer domesticus (sparrow) AY494987 16687285 fragment 0 KGLRLHDNPVLLAALESSEALYPVYILDRAFLTSSMHIGALRWNFLLQSLEDLHKNLGQLGSCLLVIQGEYEIVLRDHIQKWNITQVTLDAEMEPFYKEMEANIQRLGVELGFEVFSLVSHSLYNTQR 2 1 ILDLNGGSPPLTYKRFLHILSLLGDPEVPVRNVTAEDFQ 2 1 RCRAPDPGLAECYRVPLPVDLKISPESLSPWRGGETEGLQRLERHLTDQ 0 0 GWVTSFTKPRTVPNSLLPSTTGLSPYFSMGCLSVRTFFYRLSNIYAQ 0 0 AKHHSLPPVSLQGQLLWREFFYTVASATPNFTQMAGNPICLQICWYKDAERLHKWKT 0 0 AQTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLW 0 0 2 1 12 1 2 * 0 >CRY4_anoCar Anolis carolinensis (lizard) FG650345 synteny: UBE2T CRY4 LRIF1 DRAM2 verified indel exon 3 0 MSHRTIHLFRKGLRLHDNPTLLAALKSSEIVFPVYILDRKFMTSAMQLGTIRWHFLLQSLADLQASLKKLNSCLWIIQGEYEAILREHVKKWNITQVTFDDEIEPFYKEMERTIQNIGKEMGFEVFSMVGHTLYDVKR 2 1 IVELNRGTPPLTYKRFLQILTTLGDPEMPVEDLTAENSQ 2 1 KLCPLPEEDLDECYRVPLPEDLGISKEHLSPWRGGETTGLQRLEEHMKNQ 0 0 GWIAKFTKPRTIPNSLLPSTTGLSPYFSLGCLSVRMFFYRLSKIYAQ 0 0 SKNHSLPPVSLQGQLLWREFFYTVASATPNFTKMVGNPICLQIEWYEDAEKLHKWKT 0 AQTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDSDYSINAGNWMWLSASAFFHQYSRIFCPVRFGKRTDPEGHYIR 2 1 KYLPVLKNFPSKYIYEPWTAPAEAQREAGCII 12 GKEYPFPMVNHKEVSDNNLQLMKQVREEQYKTAQLTR 1 2 DDVDDPMEMGLKHRHSEEVTPKGKHARKAEEL* 0 >CRY4_xenTro Xenopus tropicalis (frog) NP_001123706 0 MPHRTIHIFRKGLRLHDNPTLVTALETSDVVYPVYILDRNFMTSSSVIGSKRWNFFLQSIEDLHCNLQKLNSCLFVIQGDYERVLREHVEKWNITQVTFDLEIEPYYKGLDERIRAMGQELGFEVVSMVAHTLYDIKK 2 1 ILALNCGKPPLTYKNFLRVLSMLGNPDKPARQITSEDFIK 2 1 CITPTKLAAEEYYRIPKPEDLGISKDCPTNWIGGESEALSRLEQHLEKQ 0 0 GWVANFKKPQTIPNSLLPSTTGLSPYFSLGCLSVRVFFHRLSNIYAQ 0 0 SKNHSLPPVSLQGQLLWREFFYTVASSTPNFTHMVGNPICLQIDWYKNEEQLQKWKE 0 0 AKTGFPWIDAIMTQLHNEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEEFLLDADYCINAGNWMWLSASAFFHHYTRIFCPVRFGKRSDPEGNYIR 2 1 KYLPVLKNFPAKYIYEPWTAPEEIQKQAGCLI 12 GKDYPLPMVDHKTASEHNLQLMRQVREEQQKTAQLTK 1 2 DIADDPMELNLKHPFKNDKECGLENNKVKRRCLEEMTAEDGTRQSDLVQSLDPCEVKVC* 0 >CRY4_xenLae Xenopus laevis (frog) BC167313 only 89% identical to CRY4_xentro 0 MPHRTIHIFRKGLRLHDNPTLVAALETSDIIYPVYILDKNFMTSSSVIGSKRWNFLLQSIEDLHCNLQKLNSCLFVIQGDYQSVLREHVQKWHITQVTFDLEIEPYYKGMDERIRAMGQELGFDVVSKVAHTLYDVKS 2 1 ILALNYGKPPLTYKNFLRVLSVLGDPDKPARQITLEDFIK 2 1 CTTPTEFAAEEYYRIPKPEDLGICRDCAPNWKGGESEALCRLEQHLEKQ 0 0 GWVANFQKPQTVPNSLLPSTTGLSPYFSFGCLSARVFYHRLSNIYAQ 0 0 SKNHSLPPVSLQGQLLWREFFYTAASSTPNFTHMVGNPICLQIEWYKNEEQLQKWRE 0 0 GKTGFPWIDAIMAQLHEEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADYSINAGNWMWLSASAFFHHYTRIFCPVRFGRRTDPEGNYIR 2 1 KYLPVLKNFPAKYIYAPWTAPEEIQKQSGCLI 12 GKDYPLPMVDHTTASEHNLQLMRLVREAQQKTAQLTT 1 2 DIADDPMELNLKHPFKTDKESVLENKVKRRCLEETAGEDGNRQSDLVQSLDPCEVKVC* 0 >CRY4_latCha Latimeria chalumnae (coelocanth) AFYH01009222 0 MTHRTIHIFRKGLRLHDNPILLAALEFSRVVYPVYILDRKLLESGVIIGALRWRFILQSLEDLHRNLVKLNSRLFVIQGDYEQILREYVQKWTITQVTFDTEIEPFYKEMDKKVRLMGKEMGFTVLFSVAHALYDVAR 2 1 IVENNGGQPPLTYKKFLHVLSKLGDPERPVRDITVDDFQ 2 1 KCMPPNPGLKELFRVPLPEELGLQSEHNASWIGGESQGLQRLEQHLENQ 0 0 GWIAHFTKPRTIPNSLLPSTTGLSPYFTLGCLSVRNNFYRLSHIYAQ 0 0 SKSHSLPPVSLQGQLLWREFFYTVASATPNFTKMVGNPICLQIGWYEDPEGLRKWRT 0 0 AETGFPWIDAIMTQLRQEGWVHHLARHAVACFLTRGDLWISWEGGMK 0 0 VFEELLLDADYSMNAGNWMWLSASAFFHQYTRILCPVRFGKRTDPEGHYIR 2 1 KYLPVLKNFPSKYIYEPWTAPEEVQQQAGCVI 12 GKDYPFPMVNHKEVSKNSLHLMKQVREEQHKVTQLTR 1 2 DVADDPMEMRMKRKFPDDFKNKGIFSETAERSSTT* 0 >CRY4_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01016726 0 MTHRTIHLFRKGLRLHDNPTLLGALESSAVLYPVFILDRAFMEEAMARGVLRWRFILQSLEDLDTRLQAIGSRLFVLCGSTANILRELVAQWGITQISYDTEVEPYYTRMDKDIQTVAQENGLQTYTCVSHTLYDVKR 2 1 IIKANGGEAPLTYKKFLHVLALLGAPETPARRITREDFR 2 1 CRTPTEEESEEKYRVPSLEDLGIVVESEALWVGGETEGLQRLEQHMQNQ 0 0 GWIENFTKPRTIPNSLLPSTTGLSPYFSLGCLSVRMFYHRLSNIYAQ 0 0 SKNHSLPPVSLQGQVLWREFFYTVAAATPNFTRMAGNPICLQIDWYKDQEALDRWKT 0 0 ARTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEEHLLDADYSVNAGNWMWLSASAFFHQYTRIFCPVRFGRRTDPEGHYLR 2 1 KYLPVLKNFPSQYIYEPWTAPQQVQLEAGCII 12 GKDYPLPMVNHREVSESNLALMREVRREQEKTAQLTR 1 2 DTADDPMEVGKKRACQREAETDRSDAFLEKPERAKRFSHAEGEAKALACSWPSETLRLPALGREVM* 0 >CRY4_danRer Danio rerio (zebrafish) BC164413 adjacency to lost CRY1 suggests relationship 0 MSHRTIHLFRKGLRLHDNPSLLGALASSSALYPVYVLDRVFLQGAMHMGALRWRFLLQSLEDLDTRLQAIGSRLFVLCGSTANILRELVAQWGITQISYDTEVEPYYTRMDKDIQTVAQENGLQTYTCVSHTLYDVKR 2 1 IVKANGGSPPLTYKKFLHVLSVLGEPEKPARDVSIEDFQ 2 1 RCVTPVDVDRVYAVPSLADLGLQVEAEVLWPGGESHALQRLEKHFQSQ 0 0 GWVANFSKPRTIPNSLLPSTTGLSPYLSLGCLSVRTFYHRLNSIYAQ 0 0 SKNHSLPPVSLQGQVLWREFFYTVASATPNFTKMEGNSICLQIDWYHDPERLEKWRT 0 0 AQTGFPWIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWITWEEGMK 0 0 VFEEFLLDADYSVNAGNWMWLSASAFFHKYTRIFCPVRFGRRTDPQGEYLR 2 1 KYLPVLKNFPSQYIYEPWKAPEDVQLSAGCII 12 GKDYPRPIVSHIEASQRNLALMRQVRTEQQTTAELTR 1 2 DVADDPMEAGLKRELREEEGLLEEAESQCTSKRFSGSSDHKSRPCSWTPETLQLSELSGEVM* 0 >CRY4_molTec Molgula tectiformis (tunicate) CJ347377 CJ411442 CJ358785 fragment imputed introns 0 2 1 PVQTISEEHFK 2 1 VCVTPAIPEGNTEHEIPTLEKLGFQPPDNPPLWEGGETAALKRLNQRFLEQ 0 0 GWLDGKPRRVKQNMLLPTTTGLSPYFNFGCISPRYFFARLTSVKYA 0 0 NQKDILPPVSLQGQILWRDFFYTVAGHTPNFTRIKENPICLQINWYENEEQLNAWKQ 0 0 GRTGYPWIDAIMRQMNQEGWIHHLARHAVACFLTRGNLWLSWEKGQE 0 0 YFEEKLVDADYSMNAGNWMWLSASAFCHNYTRVFHPVCFGKRVDPYGEYVK 2 1 RYVPELKDFPLRYIYEPWKAPLEIQKKANCIIGQGYPEPIVEHGKVWKKNMKMMAEIRDSQKATVALTD 1 2 DKCSESMRLPTISTKKEVLVEIPFNLEEDKTIVDGKFTKKE* 0 >CRY4_braFlo Branchiostoma floridae (amphioxus) Un:610812841 XM_002609457 exon 4,7,8 wrong 0 MEVREQHNTIHWFRKGLRFHDNPSLLHALRTSRHVYPVFVMDLDFMKDFKIRSGANQWRFVIECLQDLDTRLRAYGLRLFVARGNAEAFFAEHFRKWNITQLTHDVETEHYHRFRDAAVRKIAVDAGVEVVNYVAHTLYNIDK 2 1 IIEYNGGTAPLTYRQFQKVLKDFGAPPQASETATAEHFASCSVPLEALRDERYNMVTLEELGMRCEHPSKFVGGETEGLRRVEKHLQNQ 0 0 GWVTQFEKPKTAPTSLLPSTTGLSPYFSFGCLSVRHFYHRLDKIYAK 0 0 AKNHSLPPVSLQGQLLWREFFYVAAAQTPNFVVMAGNPICLQIPWYENREHLRRWRN 1 2 AETGFPWIDAIMSQLQKEGWVHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDADYSVNAGNWMWLSGSAFYHQYTRIFCPVRFGKRTDPDGTFIR 2 1 AYLPVLKDFPAQYIYEPWKAPLNVQEAAGCIVGMDYPFPMVDHEIVSQQNLELMREIREQQRQITLLTL 1 2 * 0
Cryptochrome 6-4 photolyases
>CRY64_anoCar Anolis carolinensis (lizard) XM_003225714 6-4 photolyase synteny: DCPS TIRAP CRY64 SRPR FOXRED1 0 MAHVSIHWFRKGLRLHDNPALLAAMKNSAEIYPIFILDPWFPKNMQVSINRWRFLIESLKDLDESLKKLNSR 2 1 LFVVRGRPAEVFPELFTKWKVTRLAFEVDTEPYARRDAEVVRLAAEHGVQVIQKVSHTLYDTER 2 1 IIVENSGKAPLTYTRLQTLVASLGPPKQPVPAPKLEDMK 1 2 DCCTPVKEDHDLEYGTPSYEELGQDPKTAGPHLYPGGETEALARLDLHMKRT 0 0 SWVCNFKKPETHPNSLTPSTTVLSPYVKFGCLSVRMFWWKLAEVYQG 0 0 RKHSDPPVSLHGQLLWREFFYTAGAGIPNFDRMENNPVCVQVDWDNNQEYLRAWRE 0 0 GQTGYPFIDAIMTQLRTEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDADWSLNAANWQWLSASAFFHQFFRVYSPVTFGKKTDKNGEYIK 2 1 KYLPFLRKFSNDYIYEPWKAPRSLQERAGCIIGQDYPKPIVEHEKVYKRNLERMKAAYARRSPNLVIQAKDKVSQKK 1 2 GVNRKRPEAPTKAKVQAKKVKTKSS* 0 >CRY64_chrPic Chrysemys picta (turtle) AHGY01135270 AHGY01135271 no synteny 0 MKHVSIHWFRKGLRLHDNPALLAATTDCRKLYPIFVLDPWFPKNMRVSVNRWRFLVESLRDLDESLQKLNSR 2 1 LFVVRGRPTEIFPVLFKEWKVTRLTFEVDTEPYSRQRDSEVASLAAEHGVQVIQKVSNTLYDTDR 2 1 VIAENNGKVPLTYVRLQTLLATLGPPKRPVPAPTLENLK 1 2 DCCTPCKDNHDAEYGIPTLEELGQDPEQAGLRLYPGGESEALCRLDLHMNRT 0 0 AWVCNFQKPQTEPNSLNPSTTVLSPYIKFGCLSVRTFWWRLAEVYQG 0 0 KKHSNPPVSLHGQLLWREFFYTAGAGIPNFNKMEGNPVCVQVDWDNNPEHLRAWSE 0 0 GRTGYPFIDAIMAQLRTEGWIHHLARHAVACFLTRGDLWISWEQGQK 0 0 VFEELLLDADWSLNAGNWQWLSASAFFHQFFRVYSPIAFGKKTDKSGEYIK 2 1 KYLPFLRKFPAEYIYEPWKAPRSMQEQAGCVIGRDYPKPIVVHEVVSKRNVERMKAAYARRSSSTTAQLEGGGGKKGIN 1 2 GAKRRTPAGPSVAELLTKKPKT* 0 >CRY64_allMis Alligator mississippiensis (alligator) blat 0 MKHSCIHWFRKGLRLHDNPALLAAMKDCSELYPIFILDPWFPKNMQVSVNRWRFLVESLKDLDESLKKLNSR 2 1 LFVVRGHPAEVFPGLFKAWKVTRLTFEVDTEPYSKQRDAEVVRLAAAHGVQVIQKVSHTLYDTDR 2 1 VIAENNGKVPLTYRQLQAVLAGLGPPKQPVLAPTLETLK 1 2 DCCTPGRDNRDPKYEIPTLEELGQDPKEAGPCLYPGGESEALSRLAFHMKRM 0 0 TWVCNFKKPDTEPNTLSPSTTVLSPYVKFGCLSVRTFWWKLAEIYRG 0 0 KKHSSPPVSLHGQLLWREFFYTAGAGIPNFNRMEGNPICVQVDWDDNPEYVKAWKE 0 0 GRTGYPFIDAIMTQLRTEGWIHHLARHAVACFLTRGDLWVSWEEGQK 0 0 VFEELLLDADWSLNAGNWQWLSASAFFHQFFRVYSPVAFGKKTDKSGAYIK 2 1 KYLPILRKFPAEYIYEPWKAPRSMQEQAGCIIGRDYPKPIVEHEALSKRNIMRMKAAYAQRSHSKAAQVEKESTKKGN 1 2 GGKRKLPAGPSVVELLTKKPKAKSS* 0 >CRY64_croPor Crocodylus porosus (crocodile) blat/genome 0 MKHSCIHWFRKGLRLHDNPALLAAMKDCSDLYPIFILDPWFPKNMQVSVNRWRFLIESLKDLDESLKKLNSR 2 1 LFVVRGHPAEVFPGLFKAWKVTRLTFEVDTEPYSKLRDAEVVRLAAAHGVQVIQKVSHTLYDTDR 2 1 IIAENNGKVPLTYRQLQAVLAGLGSPKQPMLAPTLETLK 1 2 DCCTPARDNHDPKYEIPTLEELGQDPKEAGPRLYPGGESEALSRLDFHMKRM 0 0 TWVCNFKKPDTEPNTLSPSTTVLSPYIKFGCLSVRTFWWKLAEIYRG 0 0 KKHSSPPVSLHGQLLWREFFYTAGAGIPNFNRMEGNPICVQVDWDDNLEYVKAWKE 0 0 GRTGYPFIDAIMTQLRTEGWIHHLARHAVACFLTRGDLWVSWEEGQ 0 0 VFEELLLDADWSLNAGNWQWLSASAFFHQFFRVYSPVAFGKKTDKSGAYIK 2 1 KYLPILRKFPAEYIYEPWKAPRSMQEQAGCIIGRDYPRPIVEHEAVSKRNIMRMKAAYAQRSRSHSKSAQVEKEGTKKGN 1 2 GGKRKLPAGPSVVELLTKKPKAKSS* 0 >CRY64_xenTro Xenopus tropicalis (frog) synteny: STS1 RPL27A CRY64 FOXRED1 SRPR PubMed:19715341 19345672 9016626 0 MKHNSIHWFRKGLRLHDNPALLAAMKDCAELYPIFILDPWFPRNMKVSVNRWRFLIEALKDLDENLKKINSR 2 1 LFVVRGKPTEVFPLLFKKWKVTRLTFEVDTEPYSRQRDADVEKLAAEHNVQVIQKVSNTLYAIDR 2 1 IIAENNGKPPLTYVRFQTVLALLGPPKRPVQVPTQENMK 1 2 DCCTLWKSSYNEKYGVPTLEELGQDSLKLGPRLYPGGESEALSRLELHMKRT 0 0 TWVCNFKKPETEPNSLTPSTTVLSPYVKFGCLSARTFWWRIADIYQG 0 0 KKHSDPPVSLHGQLLWREFFYTAGVGIPNFNKMEGNTVCVQVDWGNNKEHLQAWSE 0 0 GRTGYPFIDAIMTQLRTEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDADWSLNAGNWQWLSASTFFHQFFRVYSPVAFGKKTDKNGDYIK 2 1 KYLPILKKFPAEYIYEPWKAPRSLQERAGCIIGKDYPKPIVEHDVASKQNIQRMKAAYARRSGSTAEVDKDSGQSNKN 1 2 GAKRKVAGGPSVAELFKKNKSKKD* 0 >CRY64_ambMex Ambystoma mexicanum (axolotl) CO788545 fragment 224-370 TVRNPGMFEMHMKRTAWVCNFKKPDTEPNSLTASTTVLSPYVKFGCMSVRTFWWRLAEIY QGKKHSDPPVSLHGQLLWREFFYTTGAGIPNFNKMEGNSVCVQVDWDNNQENLRAWSEGR TGYPFIDAIMTQLRTRRLDPPLGASCCGVLPYERIHHLARHAVACFLTRGDLWKL >CRY64_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01024141 AHAT01024142 0 MMHRSIHWFRKGLRLHDNPALLASLRDCAELWPVFLLDPWFPKNARVSVNRWRFLLRALQDLDGNLRKLGSR 2 1 LFVVRGSPAEVFPRLFEQWKVTRLTFEVDTEPYARQRDAQVGKIAEEHGVEVIQKVSHTLYDTER 2 1 ILVENNGKAPLTYNRLQALLKTLGAPKRPVPPPTAEDMK 1 2 GVCTPCSERHDEEFGVPTLEELGQDPRTAGPELYPGGETEALSRLDRHMQRT 0 0 AWVCGFQKPNTEPNALSPSTTVLSPYLKFGCLSARTFWWRLTDVYRG 0 0 0 0 GCTGFPFIDAIMTQLRSEGWVHHLARHAVACFLTRGDLWISWQEGQK 0 0 VFEELLLDADWALNAGNWQWLSASAFFHQYYRVYSPIAFGKKTDKNGDYIR 2 1 KYLPVLKKFPSAYIYEPWKAPRSVQEQAGCIVGKDYPRPIVDHDVVSKKNIQRMKLAYARRAQLGGEQEGTGK 1 2 GMKRKGQSVADLLTKKQKRNSVEEKMTLSG* 0 >CRY64_danRer Danio rerio (zebrafish) BC044204 6-4 photolyase aka CRY5 synteny: FOXRED1 0 MSHNTIHWFRKGLRLHDNPALIAALKDCRHIYPLFLLDPWFPKNTRIGINRWRFLIEALKDLDSSLKKLNSR 2 1 LFVVRGSPTEVLPKLFKQWKITRLTFEVDTEPYSQSRDKEVMKLAKEYGVEVTPKISHTLYNIDR 2 1 IIDENNGKTPMTYIRLQSVVKAMGHPKKPIPAPTNEDMR 1 2 GVSTPLSDDHEEKFGIPTLEDLGLDTSSLGPHLFPGGEQEALRRLDEHMERT 0 0 NWVCKFEKPKTSPNSLIPSTTVLSPYVRFGCLSARIFWWRLADVYRG 0 0 KTHSDPPVSLHGQLLWREFFYTTAVGIPNFNKMEGNSACVQVDWDNNPEHLAAWRE 0 0 ARTGFPFIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDSDWSLNAGNWQWLSASTFFHQYFRVYSPIAFGKKTDKHGDYIK 2 1 KYLPVLKKFSTEYIYEPWKAPRSVQERAGCIVGKDYPRPIVDHEVVHKKNILRMKAAYAKRSPEDKTINK 1 2 GEKRKASPSIKEMFQKKAKR* 0 >CRY64_salSal Salmo salar (salmon) BT058852 0 MAHTCIHWFRKGLRLHDNPALVAALRDCKEIYPVFVLDPYSPNNVNIGINRWKFLIGALKDLDCSLRKLNSR 2 1 LFVVRGKTDEVFPKLFQKWKVTRLTYEYDTEPFSLRRDKEVGRLAEEHGVEIIYKVSHTLYNIDR 2 1 IIEENNGKAPLTYNRLQTLVSSIGPPKRPIPAPTCDDMK 1 2 DVKTPCSEKHEENYGISTLEKLYQDPESLTEELFPGGEQEALRRLDQHMARK 0 0 EWVCGFEKPQTSPNSLSPSTTVLSPYMTFGCLSARTFWWSLTDVYQG 0 0 KKHSQPPVSLHGQLLWREFFYTAGLGIPNFDKMEGNPVCTQVDWDSNFEYLAAWAE 0 0 ARTGFPFIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDGDWSLNAGNWQWLSASTFFHQFFRVYSPIAFGKKTDKNGDYIK 2 1 KYLPHLKKYPAQYIYEPWKAPRSVQEAAGCIVGKDYPRPIVEHEVISKKNIQRMKAAYAKRSPHSSEESP 1 2 GKKEKGRKHKAPSVVDMLMKKKL* 0 >CRY64_gadMor Gadus morhua (cod) CAEA01245832 CAEA01245833 EG640694 0 MAHVCIHWFRKGLRLHDNPALMSALRGCKEIYPLFILDPCIHNKESMGVNPWRFLMGSLQDLDGSLRKLNLR 2 1 LFVVRGKPQDILPRLFQKWGVSRMTYEYDTEPYSRSRDLKVSELAKEHGVEVVYKISHTLHDVDR 2 1 IIEENNGKAPLTYGRFQTVLKTLGPPKRPIPSPTVEDIK 1 2 DVKAPCVESHEEQQYGLPSLEELGHDLSCLQEAQFPGGEEEALRRLEESMERT 0 0 GWVCSFEKPQTAPNSLSPSTTVLSPYLTFGCLSARTFWWRLADVYQG 0 0 KKHSAPPVSLHGQLLWREFFYTASVGVPNFDRMLDNPVCTQIDWDTNPEYLSAWKE 0 0 ARTGFPFIDAIMTQLRREGWIHHLARHAVACFLTRGDLWISWEEGKK 0 0 VFEGLLLDGDWALNAGNWLWLSASAFFHQYFRVYSPIAFGKKTDKHGEYIK 2 1 KYLPVLKKFPVEYIYEPWKAPLSVQKAAGCIVGKDYPSPIVEHEVISKQNIQRMKTSYGKRSQGVSESPQPMKAEKRK 1 2 GPSVLDMMKNKKKK* 0 >CRY64_takRub Takifugu rubripes (fugu) chr13:3,905,361 gene span: 2373 bp 0 MTHTCIHWFRKGLRLHDNPGLMAALRDCKELYPVFILDPQLHNKSVGVNRCRFLIGALKDLDLSLRQLNTR 2 1 LFVVRGKPEEVFPKLFCQWKITKLTYEYDTEPLSLSRDKTVTRLAEEHGIDVVCKVSHTLFDINR 2 1 IIEENNGKTPLTYKSMQAIVKKLGPPKRPLSAPSMEDLK 1 2 DVNTPCSESHEKKYRIPTLEDFGHNLADLPEEQFPGGEQEALRRLEEHMKRT 0 0 AWVCNFEKPKTSPNSLSPSTTVLSPYVTFGCLSVRTFWWRLSDVYEG 0 0 KKHSAPPVSLHGQLLWREFFYTASVGISNFNKMVDNPVCTQVDWDINSEYLAAWRE 0 0 ARTGFPFIDAVMTQLRQQGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDGDWALNAGNWQWLSASAFFHQFFRVYSPVAFGKKTDKNGDYIK 2 1 KFLPHLKKFPAEYIFEPWKAPQSVQQAAGCIVGKDYPHPIVQHEVVSKKNIQRMKAAYAKRSANTAKSLSKIQ 1 2 GLKRKPSSSVDMLKKKKKNNTSEPLL* 0 >CRY64_tetNig Tetraodon nigroviridis (pufferfish) chr5:2421194 FRMD5 CRY64 SEC1L1 TSPAN3 tiny gene: span 2341 bp 0 MGHASIHWFRKGLRLHDNPALMAALRDCKELYPVFILDPHLHNKSVGINRCRFLIGALRDLDLSLRNLNSR 2 1 LFVVRGKPEEVFPKLFSQWKVTKLTYEYDTEPYSLSRDRTVTTLAEESGVQVVYRVSHTLYDTER 2 1 VLEENNGKPPLTYNSMQAIVKKLGPPKRPISAPSMDDLK 1 2 GVSTPCLEDHEKKYGIPTLEDLGHDPAGLPEEKFPGGEQEALRRLEDQMKKT 0 0 SWVCNFEKPQTSPNSLSPSTTVLSPYVTFGCLSARTFWWRLSDVYSG 0 0 KKHSAPPVSLHGQLLWREFFYTASVGIPNFNKMADNPVCTQVDWDTDSEYLAAWRE 0 0 ARTGFPFIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDGDWALNAGNWQWLSASAFFHQFFRVYSPVAFGKKTDKNGDYIK 2 1 KYLPHLKKFPAQYIYEPWKAPQSIQKAAGCIIGKDYPHPIVKHEEVSKKNIQRMKLAYARRSTSNAASPKKT 1 2 GVKRKGPSVVDLLKKKRKKI* 0 >CRY64_gasAcu Gasterosteus aculeatus (stickleback) chrII:15282042 RASGRF2 FRMD5 CRY64 SEC1L1 TSPAN3 ISL2 0 MAHTCVHWFRKGLRLHDNPALMAALRDCKELYPVFILDPHLHNNTCVGINRWRFLVGALKDLDCSLRGLNSR 2 1 LFVVRGKPEEVFPELFDRWKVTKLTYEYDTEPYSLSRDKTVATLAKEHGVEVIYKISHTLYDTER 2 1 IIEENSGKAPLTYNRMQVIVKTLGPPKRPVPAPTIEDMQ 1 2 GAMTPCSENHEKKYGIPTLEELGQDAAALGEEQFPGGEQEALRRLDKHMKRT 0 0 GWVCSFEKPQTSPNSLSPSTTVLSPYVTFGCLSARTFWWRLAEVYQG 0 0 KKHSSPPVSLHGQLLWREFFYTASVGISNFNKMEGNPVCTQVDWDTNPEYLAAWRE 0 0 ARTGFPFIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEELLLDGDWALNAGNWQWLSASAFFHQYFRVYSPVAFGKKTDKNGDYIK 2 1 KYLPLLKKFPAEYIYEPWKAPRSVQQAAGCIVGKDYPQPIAKHEVISKKNIQRMKLAYAKRSGDSAESANKSPVKRQ 1 2 GTKRKAPSVVDMLKKKDRRK* 0 >CRY64_oryLat Oryzias latipes (medaka) 0 MAHVCIHWFRKGLRLHDNPALMAALRDCKELYPLFILDPYLYDQNLAGINRLRFLISSLQDLDCSLRKLNSR 2 1 LFVVRGKPEEVLPKLFTKWNVTKLTYEYDTEPYSRSRDKNVTMLAEEQRIQVIYKISHTLYDIDR 2 1 IIEENNGKPPLTYNRLRDIVKALGSPKKPIPAPTVEDMK 1 2 NIAPFSEKHKPEYGIPSLEELGLDTSSLAEEIFPGGEQEALRRLDTYMQRP 0 0 GWVCSFEKPNTSPNSLSPSTTVLSPYVTFGCLSARTFWWRLAEVYQG 0 0 KKHSDPPVSLHGQLLWREFFYTASVGIPNFNKMTGNPACTQVDWDENQEYLAAWRE 0 0 ARTGFPFIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEKLLLDGDWALNAGNWQWLSASTFFHQFFRVYSPVAFGKKTDKNGDYIK 2 1 KYLPILKKFPPQYIYEPWKAPRSVQQAAGCIVGKDYPKPIIEHEVISKKNIQRMKQAYARRTSGSTESPTKKQ 1 2 GVKRKAPTVVDLIQKKQKRS* 0 >CRY64_oreNil Oreochromis niloticus (tilapia) XM_003437598 AERX01000034 0 MAHTCIHWFRKGLRLHDNPALMAALKDCKQLYPVFILDPYLQNKACVGINRWRFLIGALKDLDGSLRKLNSR 2 1 LFVVRGKPEDVLPKLFTKWKVTRLAYEYDTEPYSLQRDSKVTSLAKEHGVEVIYKVSHTLYNIDR 2 1 IIEENNGKPPLTYTKLQAIVKTIGPPKRPIPAPTMDDMK 1 2 DVKTPSSENHEKEYGIPTLEELGLDTAPLGEDLFPGGEQEALRRLDEHMKRT 0 0 KWVCSFEKPQTSPNSLSPSTTVLSPYVTFGCLSVRTFWWRLTEVYRG 0 0 NKHSDPPVSLHGQLLWREFFYTASLGIPNFNKMEGNSVCTQVDWDTNPDYLAAWRE 0 0 ARTGYPFIDAIMTQLRQEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEQLLLDGDWALNAGNWQWLSASTFFHQYFRVYSPVAFGKKTDKNGDYIR 2 1 KYLPLLKKFPAEYIYEPWKAPRSIQQAAGCIVGKDYPHPIVQHEVISKKNIQRMKLAYAKRSPDTTESPSKSK 1 2 GVKRKAPSIIEMIKKKAKVK* 0 >CRY64_braFlo Branchiostoma floridae (amphioxus) BW780666 FE555184 XM_002595028 fused exons 2-3 BW780666 fusion exons 2-3 odd splice phases exon 5-6, no split 8-9 short final exon 0 MANKVQHSTIHWFRKGLRFHDNPSLLHALRTSRHVYPVYVVDQNWMKEHDIRYGANWWRFVIQCLEELDTRLRKYGLRLFVVRGSAEDFFKEHFRKWKVTQLTHDVDTEPFYRIRDVAVRKIASDMGVEVVTHVAHTLYDIDR 2 1 TVERNGGTPPLTYRKFLKVYNEMGKPPKEKETATTEHFANCTLPPEALQDKKFDMVTLEELGMRCDFPAKFVGGETEALRRLEKSLEDK 0 0 DWVLKFEKPKTSPNSLLPSTTVLSPYITVGCLSARLFYHQLDKIYSK 0 0 AKNHSQPPVSLHGQLIWREFFYTAAAHTPNFNQMEGNRVCLQIPWNKNDEHLTRWRN 1 2 AQTGFPWIDAAMTQLRKEGWIHHLARHAVACFLTRGDLWISWEEGMK 0 0 VFEELLLDADWSLNAGNWMWLSASAFFHQYFRVYSPIVFGKKTDPKGTFIR 2 1 HYLPVLKNFPKEYIYEPWKAPRNVQEKAGCIVGKDYPRPIVDHKEASQRNLDIMRDVRKDQKETAAVTL 1 2 GYGK* 0 >CRY64_strPur Strongylocentrotus purpuratus (urchin) XM_001189626 extra 1st exon unwarranted MCGAPRSYVEIRDSEEHSRRHVARLQFQFQSDLP 12 K 0 MPHNSTIHWFRKGLRIHDNPALLTAIQGTKVFRPIFILDPHFIESEKVGINRWRFLLETLQDLDYSFRALGTR 2 1 LFVVRGNPTTVFPEIFKKWNVTRLTFEVDTEPYARRRDQEVIELAKKNDVEVITKVSHTLYDTER 2 1 TIKANKYKPPMTYQRMVGLLSEIGAPAIPELPPLMANFT 1 2 GVATPVKPDHDNEYGVPSLEDLGLDLEGLGPRLYPGGETEGLQRMDLHLARK 0 0 SWVCGFEKPKTSPNSLEPSTTVLSPYLKWGCLSPRKFYYAIKEVYAQQ 0 0 TNCTKPPVSLMGQLIWREFFYTVAAGTPNFHQMEENPICIQVPWDENPEFLAAWKE 0 0 GRTGYPYIDAIMTQLRTEGWIHHLARHSAACFLTRGDLWQSWVKGQE 0 0 VFDEWLLDADYSLNAANWMWLSSSAFFHQYYRVYSPVVFGKKTDKTGEYIR 2 1 KYIPALNKLPAEYIYEPWTAPRSVQEAAGCIIGRDYPRPIVDHSIVSKRNIGRMKDARACQPGKKA 1 2 EKRPAEPSKQDNNGKKVRKITSMLKKK* 0 >CRY64_lytVar Lytechinus variegatus (urchin) JI440523 AGCV01417941 0 MPYNSTIHWFRKGLRIHDNPALLTAIQGTKVFRPVFILDPNFLESGKVGINRWRFLLEALRDLDYSFRALGSR 2 1 LFVVRGNPTTVFPELFKKWNVTRLTFDVDTEPYARQRDQEVIELAKKSGVEVVTKVSHTLYDTER 2 1 TIKANKNKPPMTYQRMVGLLSEIGAPAVPELAPQISNFT 1 2 GVATPVKPDHDSVYGVPSMEDLGLDASGLGPRLYPGGETEGLQRMELHLARK 0 0 SWVCSFEKPKTSPNSLEPSTTVLSPYLKFGCLSPRKFYYAIKEVYAQ 0 0 TNCTKPPVSLMGQLIWREFFYTVAAGTPNFHQMEENPICLQVPWDDNPEFLAAWKE 0 0 GRTGYPYIDAIMTQLRNEGWIHHLARHSAACFLTRGDLWQSWVKGQE 0 0 VFDELLLDADYSLNAANWMWLSSSAFFHQYYRVYSPIVFGKKTDQNGDYIR 2 1 KYIPIMERFPAQYIYEPWTAPRSVQEAAGCIIGRDYPRPIVDHSVVSKRNIGRMKDARACQPGKSA 1 2 EKRPTDASNKNSNGKVRKITSMLKKK* 0 >CRY64_eucTri Eucidaris tribuloides (pencil_urchin) JI324408 fragment imputed introns 0 STTVLSPYLKFGCLSPRKFYYAIQEVYNQK 0 0 SNCTKPPVSLHGQLLWREFFYTVASGTPNFAQMEENPICLQVPWDNNTELLEAWKE 0 0 ARTGFPFIDAIMTQLKQEGWIHHLARHASACFLTRGDLWVSWVEGQK 0 0 VFEEWLLDADYSLNAANWMWLSSSAFFHQYYRVYSPVAFGKKTDKTGEYIR 2 1 KYLPILRRMPTEYIYEPWNAPRSVQEAAGCVVGRDYPHPIVDHSMVMKRNLQRMKDARECRPKET 1 2 TKRPLTES * 0 >CRY64_aplCal Aplysia californica (sea_hare) scaffold_427 0 MGKHRAIHWFRKGLRLHDNPALLKACEQATELYPIFVLDPWFASSPQCHVGCNRWRFLLQALQDLDQSLQSLNLR 2 1 LFVIRGQPEVEIEKLCQQWSITKLTYEADTEPYAVQRDAKVEARMEALGVEVIKCVSHTLFDVNL 2 1 TVKNNGGCAPLTYQKFQSVLSKMGTPAKALLSPTLTDLEGIK 1 2 GCKTPVSSSHDQYYGVPTLEELGKSSSDCGPLLFPGGETEALDRMERHLKKI 0 0 NWICNFEKPKTSPNSLSPSTTVLSPYLKFGCLSPRLFYHRLQQVYSGK 0 0 KHSQLQFLSTDNCCGENSFYTVGAVTPNFDRMKGNPGVTQIPWVQRLEHFESLDR 0 0 ARTGYPFIDAVMTQLRLEGWIHHLARHAVACFLTRGDLWVSWEQGMK 0 0 VFEELLLDADWSLNAGNWMWLSASAFFHQYFRVYSPVAFGKKTDPNGDYLR 2 1 KYLPVLKKFPAKYIFEPWLAPSSVQQAAGCIVGKDYPRPIVDHDRARQENIKKMAAAYAAAKSSNE 1 2 * 0 >CRY64_vilLie Villosa lienosa JR505030 transcript assembly mollusc 0 MTQRSVHWFRKGLRLHDNPALLAAFENSSHVWPIFILDPWFIKNAQVGVNRWRFLLQSLQDLDNNLKKLNSR 2 1 LFVIRGNPNEVLLELFKEWKVTKLTFEVDTEPYAQQRDGEIEELAKKHNVKVVKCVSHTLYDVQE 2 1 TIEKNGGKPPLTYQRLQTILFKMGPPPKPVKTLDSSVVM 1 2 DNKTTVSEDHDKKYCVPSLADLGKSHEACGPLLFSGGETEGLSRLERMLKKT 0 0 NWICTFEKPKTEPNSLTPSTTVLSPYLKFGCLSVRTFYYRLHEVYSQN 0 0 KKHTQPPVSLHGQLLWREFFYTAGAGTPNFDKMEGNPICVQIPWEKNRGHLEAWAN 0 0 GRTGYPFIDAVMTQLRKEGWIHHLARHAVACFLTRGDLWISWEEGQK 0 0 VFEEHLLDADWSINAGNWMWLSASSFFYQYYRVYSPISFGKKTDKNGDYIR 2 1 KYLPILSKFPEKYIYEPWTAPNDIQVKAGCIIGKDYPPPIVNHDKMRPINIKRMAEAYGKRFDSGKDNTNGGSIKKKI 1 2 SASEDDDEAPKKKKVKKSSNKMTDFLM* 0 >CRY64_droMel Drosophila melanogaster (fruitfly) 6-4 photolyase PDB:3CVW CG2488 uses 5-deazariboflavin 0 MDSQRSTLVHWFRKGLRLHDNPALSHIFTAANAAPGKYFVRPIFILDPGILDWMQVGANRWRFLQQTLEDLDNQLRKLNSRLFVVRGKPAEVFPRIFKSWRVEMLTFETDIEPYSVTRDA AVQKLAKAEGVRVETHCSHTIYNPELVIAKNLGKAPITYQKFLGIVEQLKVPKVLGVPEKLKNMPTPPKDEVEQKDSAAYDCPTMKQLVKRPEELGPNKFPG 1 2 GETEALRRMEESLKDEIWVARFEKPNTAPNSLEPSTTVLSPYLKFGCLSARLFNQKLKEIIKRQPKHSQPPVSLIGQLMWREFYYTVAAAEPNFDRMLGNVYCMQIPWQEHPDHLEAWTHGRT GYPFIDAIMRQLRQEGWIHHLARHAVACFLTRGDLWISWEEGQRVFEQLLLDQDWALNAGNWMWLSASAFFHQYFRVYSPVAFGKKTDPQGHYIRKYVPELSKYPAGCIYEPWKASLVDQRAYGCV LGTDYPHRIVKHEVVHKENIKRMGAAYKVNREVRTGKEEESSFEEKSETSTSGKRKVRRATGSAPKRKR* 0 >CRY64_danPle Danaus plexippus (butterfly) EF117813 PubMed:17244599 two novel exons 0 MTKVASVIHWFRLDLRLHDNLALRNAINE 0 0 AENRKQILRPIYVIDPDIKNRVGCNRLRFLFQSLKNLDTSLRKINTRLYVIKGKAIECLPKLFDEWHVKFLTL QVDIDADLVKQDEVIEEFCEANNIFVVKRMQHTVYDFNSVVKKNNGSIPLTYQKFLSLVSDVQVKDIIQISKGVSDECKASDYDSQGYDIPSLEEFGVNESELSECKYPGGESEGL KRLDVYMAKKQWVCNFEKPKSSPNSIEPSTTVLSPYISHGCLSAKLFYHKLKQVENGSKHTLPPVSLMGQLMWREFYYTAGSGTENFDKMVGNSVCTQIPWKKNDAHLKAWAEGKT GYPFVDAIMRQLKQEGWIHHLARHMVACFLTRGDLWISWEEGAKVFEDFLLDYDWSLNAGNWMWLSASAFFYKYYRVYSPVAFGKKTDKDGLYIRKYVPELKKYPSEFIYEPWKAP KGVQKTAGCIIGEGYPNRIVDHDKVHKDNIQKMNAAYKVNKEKKAMKRPRQ* 0 >CRY64_acyPis Acyrthosiphon pisum (aphid) XM_001945977 single exon 0 MDKNDADVGRETTVHWFRKGMRLHDNPAFKLSCEAKNSNGERYKLRPIYILDPYFRKYIRAGANRWRFLQQSLVDLDTTLRKLGTRLYVIRGLPHEVFPDLF AKWNVKLLTFELDTEPYARERDNQVEQLARKHGVKVEQKVSHTIYNTELVLRANGGSVPMTYQKFVSVVGSMPTPRRPIPAPDMLPSECLLDDDLNNPEFDVPTLDELLTLKGFNP AELKPCLYPGGEKEALRRLEEYMKNKTWVCKFEKPNTSPNSLKPSTTVLSPYMKFGCLSASHFYYRLKEVIGNSPHSKPPVSLIGQLYWREFYYTVGASTPNFDKMVGNSICCQVP WDNNPDALEAWTNGKTGYPFIDAIMRQLRDEGWIHHLARHAVACFLTRGDLWISWEKGLAVFEELLLDADWSMNAGNWMWLSASAFFHQFFRVYSPVAFGKKTDKSGDYIRKYIPE LAKYPDQYIYEPWSAPKSIQERAGCVVGVHYPKRVVVHEDVYKNNIAKMSLAYKSTKAGKSSNTKKSRDMSTSPDKKNIKKPKLK* 0 >CRY64_anoGam Anopheles gambiae (mosquito) XM_314748 0 MAKRETIVHWFRKGLRIHDNPALTVAVDKVRANPAKYCLRPIFVLDPGIRKWLRVGPNRWRFLQQTLANLDENLRSINSRL 0 0 YVVRGNPVEVFPKLFADWNVSLLTYEHDIEPYAVKRDSTVEEQARKHWVEVHIEKSHTIFDPEGIVKKNGGKPPLTYQRYATLASACKIPQPLPVPQKLPAKETSPEADKEERKNPSCYDPPTMEELDIEEASMGQC KFPGGESEALRRMNEILSRKAWVCKFEKPNTSPNSLEPSTTVLSPYLKFGCLSVRLFYSRIAETIKGQKHSQPPVSLIGQVMWREFYYCVAAATPNYDKMVGNGICTQIDWDTNKD YLEAWTHGRTGYPFIDAIMRQLRQEGWIHHLARHAVACFLTRGDLWISWEEGQRVFEELLLDADWALNAGNWMWLSASAFFHQYFRVYSPVAFGKKTDPEGKFIKKYVPELARFPA GIIYEPWKANLETQKKLGCIIGKDYPMRIVVHEDISKVNIQRMSAAYKRNKAQKDGGTEQDDSVSSGSGGTPGKKRPTSTGSTAKGKAAPKKRKIEANIEKFLKKK* 0 >CRY64_bomMor Bombyx mori (silkworm) AK381942 frameshift 0 MSKTATVIHWFRLDLRIHDNLALRNAINE 0 0 AENRKHLLRPIYFLDPNIKDKVGINRLRFLLQSLEDLDSNLKKLNTCLYVLRGKAVDLLPKLFDDWQVKYLTCQVDIDPEFVQQDEYIEDIAEKKGVFINKRVQHTVYDVHKVLRENN GAVPLTYQKFLSLVKSINVKEPIEISNVLSSHCKPIDIQSVNYSIPNLKELQIDEETLAPVKYHGGETEALKRLNLYMSKKEWVCKFEKPNSSPNSIEPSTTVLSPYISHGCLSAKLFYH KLKEVENGRQHTLPPVSLMGQLMWREFYYTAGTGVANFDKMVGNAICIQIPWTKNDAFLKAWAEGKTGYPFVDAIMRQLKQEGWIHHLARHMVACFLTRGDLWISWEEGAKIFEDYLLDY DWSLNAGNWMWLSASAFFYKYFRVYSPVAFGQKTDKEGVYIKKYVPELKKYPREYIYEPWKAPQSIQRNAGCIIGEHYPKRIVNHDTIHKENIQKMATAYKLNREKKAVKRPRS* 0 >CRY64_craMey Crateromorpha meyeri (sponge) PubMed:20121950 MTYNTLHIFTDDALRINDNNSLLSNLENTKNLYTVFYYGHIKKINHIPPHRVIFLLESLKNLKERLDEYGIPLYFIDEPMFQSLRMLISKWNINRVTTEP VVSIVGKRDQLSLRNFLSAFGVMLRTYNSSTLYDTVKVPVNITRSEFFRIITQMEPELSLPEELTEFLSKFSSFPDPFINKKIPSLSEFGITNETMSSID TKFRGGETAAILQLEKLIANRCEPNNLPKVAQLNQYDAISPAIKFGCISVRTIYNRVSKLEPKYNEVKNQIYDGLRNRDYCILVGGNCPNIDNQGSIYTY ILPWDVKQDASTRFQTGRTGYPFIDAAIAQLKREGFIHNSVKNILVRTLTCDLMWIGWHEGVRMFYKWSLDYNAAICALSWMHGSKSTWLLEEISISQIN PIEEAKEIDKDGDYIRKYLPELKDYPSEYIHTPWLAPLDQQIESECVIGQDYPYPNYCDVEERVQQCRKRLQIFYNIMPIAKKRRTLSLLKKRTRININNNDSVINIRAQLECKATKLSQ* 0
>CRY64A_triAdh Trichoplax adhaerens (placozoa) XM_002108524 ABGP01000049 no UIM domain affinity to CRY class MVCTVVWLRYDLRLIDNPALFHAAKRGHVVIIYILDQETPGKWKLGKASLWWLHHSLKSLQNDLLKFNIPLILRRGADPLTILRQVLLQSRADAVYWNRC YEPYAVKRDQSIEEALKVDDVTVGSFQAGLLYEPWAVKSAKDEPFKLYIMYWNRCLSSDQPRKPYDKPRFNNTLDIKVQTDTLDSWHLVSAKKDWKDELK IVIGCPGEDGAIRKLKEFIKYKLVDYRKGKETVWPSKTSQLSTHLHFGEISPFVVWHTAKESVHLRSVPSDSSQKFLTELGWRDFCFHLLHYYPDFPEKS YKLNFDETIWKVDKEKLKLWQQGKTGYPIVDAGMRELLNTGVISYRVRTIVASFLTKHLLIPWQNGAEWFWDTLVDADLALNSCSWQRITGCGADISSYF YIINPVTQGERFDSDGNYVRKWIPEIAKLSNDYIQQPWEAPAAVLKKAGIALGKTYPKCIVNHKKARDVALTKFAPLKRQEPMDNLKKLPNKKKMKK* 0 >CRY64B_triAdh Trichoplax adhaerens (placozoa) XM_002107723 ABGP01000051 anti-parallel tandem no UIM domain MSSSKIPCTLVWLRQDLRLIDNPALFHAAKRGQLITVYIFDEDCSGKRRTGQACLWWLYHSLKSLRNQLSQWNIPLIVKSGKDAFTILKELLDSYNADAI YWNRCYEPFEYNRDCSIEEKLKMENVTVETYQDRVLFEPWTIKTAKGGPVQIYVHFWNQCLSLPQPRKPYDKPKFKNQTGCQSKCDEPAVWQRLAKLSQD MPMKLQDFWHPGEEQAISKLKSFIKNSLNAYAEKSQCLRLNTTSNLSPYLHFGEISPFTVWHSVRHASSSAEPPAVKSESLKQYLRSLGWREYTCHLLFH YPDLPQKCFRSSFEQLKLQVDPVKLQSWQQGNTGYPVVDAGMRQLSVAGIMPNRLRMIVASFLIKHLLVPWQKGEEWFWSKLIDADLAQNAFNWQWAAGS GPNACPYFRVFNPITQGEKFDADGYYVRKWIPELAALPNAYIHKPWEAPSTVLKKANIVLGKTYPRPIVQHKAARELALATFGQTKRKQEPPTSNSNKRVKEE* 0
CRY7 UIM ubiquitin cryptochromes
>CRY7_xenTro Xenopus tropicalis (frog) XP_002938187 AAMC01077621 AAMC01077620 many transcripts CDK10+ CRYM+ GCSH- PDK1L2- BCMO1+ GL172982:273,091-396,180 1U3C 34% 3CVW 29% 0 MDLEPFERAQIDDVLQQLESGSVQADEFLCLVLSILGSSRTYSQFPAILQSLSRKEPAMYRELMDLHAEYFRK 0 0 EPADLETLGYETDLELAIALSLQEHNQLTDTASFASEVDPAPKISFADAAKLSHFSHKHNKKNSSSKTEITKLKDNVAAMNLYQERKRYHINGQEKTCISN CYNGQPEPEDCVLKSEDGEDVFHVETSRPRESKAKHSRRSRKKKKSAPSRGLVAMKPVLVWFRRDLRLHDNPALISALEHGVPVIPVFLWCINEETGQNFTLATGGAT KYWLHHALLKLNQSLIQRFGSHIIFRVARSCEEELVSLVHETGADTIIINAVYEPWLKERDDLISETLRRHGVELKKHHSYCLYEPDSVSTEGVGLR 1 2 GIGSVSHFMSCCKRNNSAPIGMPLDAPRCLPAPCNWPESDHLDTLELGKMPHRKDGTL 0 0 IDWAVTIRESWDFSEDGAYTCLANFLQD 1 2 GVKHYEKESGRADKPYTSHISPYLHFGQISPRTVLHEAYFTKKNVPKFLRKLAWRDLAYWLLILFPDMPSEPVRPAYK 0 0 SQRWSSDLNHLRAWQKGLTGYPLVDAAMRELWLTGWMCNYSRHVVASFLVAYLHIHWVHGYRWFQ 0 0 DTLLDADVAINAMMWQNGGMSGLDHWNFVMHPVDSALTCDPYGSYVRKWCPELAGLPDEYIHKPWKCAPSQLRRA 1 2 GVILGRNYPHRIVLDLEERREQSLKDVVEVRKKHLEYLDEVSGCDMVQIPDQLLALTLGHTSGEDEVVRNRTGSFLLPVITRKEFKYKTLQPDTKDNPYNTVLKGYV SRKRDETIAYMNERHFTASTINEGAQRHERIERTNRLMEGLPAPSDAKNKSRRTPKKDPFSIIPPSYLHLAN* 0 >CRY7_xenLae Xenopus laevis (frog) transcripts DC068968 EG576829 BU901325 0 MDLEAVERAHINDIVRQLETGIVQTDEFLCLVLSVLGNRRTFWHLPAIIQSLREKEPAMYRELMDLHAHYFRK 0 0 EPTDLETDLELAIALSMHEQNQLTDTASSEMSPTAKISFADAAKSSCYSHKYSEKTSSSKTEIDKLKHNVAAMHLSQETNRCQAIRQEKTLSSCYNGQPEPEDCVLKSVDCEEVFHVEASV RE QAKQSRRGPKKTKSAPSGVLVAMKPVLGWV EGTLRLH 1 2 0 0 1 2 VKHYEKESGRADKPYTSHISPYLHFGQISPRTVLHEAYFTKKNVPKFLRKLAWRDLAYWLLLLFPDMPSEPVRPAYK 0 0 SQRWSSDLNHLRAWQKGLTGYPLVDAAMRELWLTGWMCNYSRHVVASFLVAYLHIHWVHGYRWFQ 0 0 DTLLDADVAINAMMWQNGGMSGLDHWNFVMHPVDSALTCDPYGSYVRKWCPELAGLPDEYIHKPWKCAPSQLRRA 1 2 ILGQNYPHRIVLDLEERREQSLKDVVEVRKKHLEYLDEVSGSDMIPIPDQLLALTLGRPNGDDEVVRNRTSSFLLPVITRKEFKYKTLQPEAKDNPYSTVLKGYV GVSRKRDETIAYMNEKHFTASTIHEGAQRHERTERTSRLMEGLPAPSEPKNKSRRTPRKDPFSIIPPSYLHLAN* 0 >CRY7_ranCla Rana clamitans (green frog) GAEG01010186 GAEG01009609 fragments 91% frog DLLSETLKKHGVTFTRYHSYCLYEPYSVSTEGVGLRGIGSVSHFMGCCERNSSSPIGDPL EAPTSLPLPSCWPDSLELDRLDLAKMPRRKDGTVIDWAATIKKSWDFSEDGAYICLGNFL EDGIKHYDKESGRADKPYTSHISPYLHFGQISPRTVLHEAYFTKK SVPKFLRKLAWRDLSYWLLVLFPDMHVEPVRPAYKSQRWSSDRSHLRAWQKGMTGYPL VDAAMRELWLTGWMCNYSRHVVASFLVAYLHLHWIHGYRWFQDTLVDADVAINAMM wQNGGMSGLDHWNFVMHPVDAAMTCDPYGSYVRKWCPELAGLPDEYIHKPWKCPPSQLRRA GIVLGQNYPHRIVVDLEERREQSLRDVVEVRQKHSEYVDKVSGSDMVSVPDQLLAFTLGC ADGDDDVIKSNTCQFLLPVITRKEFKYKTLQPDTKDNPYNTVLKGYVSRKRDETIAYM NERHFTASTINEGAQLYERRERTARIMEGLPQTRDPKNKSRRTPTSDPFSIVPPAYFTI >CRY7_pseReg Pseudacris regilla (Pacific treefrog) GAEI01034610 fragment WSNDKNNLRAWQKGMTGYPLVDAAMRELWLTGWMCNYSRHIVASFLVAYLHLHWIHGYRW FQDTLVDADVAINAMMWQNGGMSGLDHWNFVMHPVDAALTCDPDGSYVKKWCPELAGLPD EYIHKPWKCAPSQLRRAGM >CRY7_latCha Latimeria chalumnae (coelocanth) AFYH01265207 pseudogene 0 DTASFNKELDFALAQSLQEMESTYSCQPEPEDCVLNSYDTDTVYSNTSK**AGPTYNFKKMKEERKKTSSLIRGPAIAVKLVLVWFRRDLHLYDNPAFVGAIELGMPVIPVFL*YLSEEDGQNYNMVVGRATKYWLHHALLQLNLASIQKFNSHIVFGKTESGLQE^KKLVNLVKETEADSLIVNTAHEPWLKERDDLIFQTLEKGVKCRHYHSYCLYEPYSVSTEGIGLK 1 >CRY7_lepOcu Lepisosteus oculatus (gar) AHAT01010533 AHAT01010534 0 MCDTGGTVTLLTQVQQMMAELQLGELDTEEFFCLTLSLLGHQNTQEQFLELIEPLALRHEKEHMCLTTIFLEYFTQ 0 0 VEEEEVEVALALSLQELGVSKQEKPQPSSHPGARESQRETPFQSGSTDVCVNVAGEASQPGAGQGVQALKATSSSGECDPRLEQLSSKGCQPPSRERAGGTPPGNTSQPDGSRSSRRRRTRKKASSVHRSPSMPRPVLVWIRR DLRLSDNPALVGSLELGAPVIPVFLWCPREEEGPGVTVAVGSASKYWLHHALLCFIQSLHKLGGHLVTRRVETTTQQALQSLVSETGADTLLANALYEPWLKERDELAFSALESQGVKCHLYHSYCLREPGSVCTEGVGLR 1 2 GIGSVSHFMSCCQHNPAPGLGSVLQAPATLPVPSQWPQSGPLDQLDLAKMPRRKDGTT 0 0 VDWAATIRSSWDFSEEGAHARLQDFLCD 1 2 0 0 AIRWSADRAHLKAWQRGRTGYPLVDAAMRQLWLTGWMNNYMRHVVASFLIAYLHLPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAAMTCDPDGSYVRKWCPELSGLPDELIHKPWKCPASVLRRA 1 2 GVTLGGDYPERIVADLEERRARSLRDVATARQQFAGEYVDRRSGCDLLPLPDKLVKEALGGGGGGEVVQRGGRFLLPVITRKEFQHQTLEPGAQTNPFDAVLRGYV SRQRDEAAAFLRERDFTASVMSEGGRRLERQERDRRVLEGLPPPPAQKSRARRTPRTDPFSVVPGGAPITPS* 0 >CRY7_danRer Danio rerio (zebrafish) ENSDART00000125725 no synteny to frog 0 MSAGEETKPVSEVRELLRELILGREDPQGFFCMCVSLLGDADTRRLFLDLIKPLSSEYEHLHSQLTSVFLEYFSK 0 0 DESEELELALTLSLYETKQIDPEDKLHNSDHQHHVPRSYKTYADVRTESSGADGSVVPENQPTAKCTLDTEVHKTRNAIKHINGDKITKSQKRRLRKKQHLLKNPSGPRPVVLWFRRDLRMWDNPALIGCLELGAPVIPVFLWNAMEEEGPGVTMSTGGAS KYWLHQALVSLKRSLEERGSHLVTLKAEPSSLTALQGLMDETGAASVVATALYEPWLKERDDALWETLEKRGVTCHIYHSYCLRDPYTVSTRGVGLR 1 2 GIGSVSHFMSCCQQNPAGGLGSPLDAPTTLPSPSAWPQGCPLADLGLARMPRRKDGTV 0 0 IDWAVDIRKTWDFSEEGAHTHLEAFLRD 1 2 GVYRYEKESCRADAPNTSCLSPYLHFGQLSARQVLWAARGARCKSPKFQRKLAWRDLAYWQISLFPDLPWESLRPPYK 0 0 ALRWSSDHAHLKAWQRGRTGYPLVDAAMRQLWQTGWMNNYMRHVVASFLIAYLHFPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPIDAALTCDPCGTFVRQWCPELKALPDDLIHKPWKCPTSMLRRA 1 2 GVVFGDSYPERIVIDLEERRAQSLQDVASVRRRFRQFVDQRSGCDLVPVPSRLVQDALGSMEDIVKHEKNGFLLPVITRMEFKHQSEDPDRQDNPYSAVLKGYV SRKRDETVAFLNERDFTASVMCESAQRRERLERDSCLLEGLPRPTASRGGARRTQTRDPYSKVPGGVAVPRK* 0 >CRY7_salSal Salmo salar (salmon) AGKD01006863 0 MPVSGGEDPMSQVRQMLRELLVGRENAEGFFCLCVSVLGHNDTRTHFLPLIQLLATDHSSLHTTLTSIYLEYFSK 0 0 DEDDELAVALALSLLEVKRQQQTDTKPLFPDPKLQGQTDTQATATYRPQNSSIQSQSVSPNGSSSQPPPAPLPKGASYAQLAAAGGRRQTQAQHQDSTSPRRSSGHGSSPQTERQRGPTKTQ KQTDIKWSNVTQDVCASIATSLTSNLNQTKDDVTVVECEVDQSEKPKRSKNRRQRRKGYGQQVVGVPRCPSAPPPVLLWFRRDLRLHDNPAVIGSLEAGGPVIPVFIWCPEEEEGPGVTVAMGGA 1 2 CKFWLHQALSCLSSALEHIGSHLVFLRPDEEREGIGSSLRALRSLVRETGAQTVLASALYEPWLRERDQVVVSALQKDRVEVNMVHSYCLRDPYTVTTEGVGLR 1 2 GIGSVSHFMSCCQMNPGPGLGVPLDPPISLPSPSVWPRGCPLEGLGLARMPCRKDGTT 0 0 IDWAANIRSSWDFSEEGAQSRLEAFLND 1 2 GVYRYEKESGRADAPNTSCLSPYLHFGQLSARWLLWDTKGARCRPPKFIRKLAWRDLAYWQLTLFPDLPWESLRPPYK 0 0 ALRWSNERGHLKAWQKGRTGYPLVDAAMRQLWLTGWMNNYMRHVVASFLIAYLHLPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAAMTCDPYGNYVRKWCTELAVLPDDLIHKPWKCPASMLRRA 1 2 GVVLGQSYPERVVTDLEERRSQSLQDVALVRRRFGEYVDPCSGCDLVPLPPRLVSEAMGVCQLTASIVL 1 2 CGQFLLPVITRMEFKHQSDDPDADAASNPYNAVLKGYVSRRRNETIAFLNQTDFTASVINEGTERRERQERDQRRMEGLPRPLAAQGRGKRTPAAKDRFSTVPGGVATSHR* 0 >CRY7_hapBur Haplochromis burtoni (chichlid) AFNZ01022319 0 MPPSGADAQATVKWWLGEVLEGREDPEGFFAMCVSILGQVETRSQFLSLIEPLSTGDRLLHTSLTSIYQEYFTK 0 0 TEDDELELALALSLLDMKGHPLQSPSQESQPLQSGDAQNQRGSVQLNSILKPEGSNCTNQTGSWNTESPPQTAREMEFQISTYPNKQTLGTGQTVCVSEQEGDVDQSQKPKRSKNRRQRRKCTGQHLVGLPRSPSAPPPVLLWFRKDLRLCDNPALVASLEVGAPVIPVFIWSPKEEEGTGITVAMGGA 1 2 GKYWLHQALLCFCSSLKHIGSRLVVLKANGEMTSTESSLKALKELIKETGARTVLANALYEPWLKERDDLVVSALQKQGVECRMFHSYCLRDPYSVTTDGVGLR 1 2 GIGSVSHFMSCCRQNPGTAVGVPLDPPVSLPTPAHWPKGLCLDELGLARMPRRKDGTT 0 0 VDWAVNIRKTWDFSEGGAQARLEAFLHD 1 2 GVYRYEKESGRADAPNTSCLSPYLHFGQLSPRWLMWDAKGARCRPLKFQRKLAWRDLAYWQLTLFPDLPWESIRPPYK 0 0 ALRWSSDRGHLKAWQRGRTGYPLVDAAMRQLWQTGWMNNYMRHVVASFLIAYLHLPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAAMTCDPYGTYVRKWCPELAELPDELIHKPWKCPASMLRRA 1 2 GVVFGQSYPERIITDLEERRNQSLQDVALVRREFEQYVDKRSGCDLVPLPPRLVSEALGLSHKDGAVVTQGKQFLLPVITRMEFKHQHEDPDADATSNPYNAVLKGYM SRKRDETIAFLNKRDFTASVMHEAVQRKERLNSDYRRMEGLSPSPSPRGRARRTPTGKDRFSIVPGGAVTSLK* 0 >CRY7_gasAcu Gasterosteus aculeatus (stickleback) DN725444 0 MPPSAGEDSKVFVRRMLVEVLAGREDPEGFFAMCVSVLGHRETSSRFLPLIQPLSTANGALHAALTSIHTDYFSK 0 0 TQDDELELALSLSLVEMDDHRLSSTSQESRPQRPADGPNKPTAVSQGSSRVPRAGVIASGKKRDSPQTGASVKANSPQTNNTPERQQNDEYETETPGASQAARVSRPVSKQDAFKESDAM MNDANLSPSEKPKRSKNRRQRRKGAGRQVVGLPRSPSAPPPVLLWFRRDLRLGDNPSLTGSLEVGAPVIPVFIWSPEEEEGPGITVAVGGA 1 2 CKYWLHQALSCFSASLERIGSHLVFLEANTSSLHTLKELVKETGARTVLANALYEPWLKERDDAVVSALQKDGVVCKTFHSYCLRDPYSVSTEGVGLR 1 2 GIGSVSHFMTCCRQNPGSALGAPLDSPVSLPTPARWPVGVPLNTLGLARMPRRKDGTT 0 0 IDWAANIRKSWDFSEEGASDRLEAFLND 1 2 GVYRYEKESGRADAPNTSCLSPYLHFGQLSPRWLLWDAKGARCRPPKFQRKLAWRDLAYWQLTMFPDLPWESLRPPYK 0 0 ALRWSSDGGHLKAWQRGRTGYPLVDAAMRQLWLTGWMNNYMRHVVASFLIAYLYLPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAAMTCDPYGSYVRKWCPELADLPDDLIHKPWKCPPSMLRRA 1 2 GVvFGQTYAERIVTDLEERRSRSLQDVALVRREFGQYVDKHSGCDLVPLPPRLVSQALGSAHRDAGLVAEGKQFLLPVITRMEFKHQQEDPDADAASNPYNAVLKGYV SRKRDETVAFLNERDFTASVMHEGVQRRERLESNYRRMEGLPQPPAPRGRARRTPTAKDKFSVAPSGAVSSVR* 0 >CRY7_oryLat Oryzias latipes (medaka) CRYM+ GCSH- two transcripts, very small introns 0 MPPSKSDQALVSEMLKEVLAGREDPEGFFAICLSVLGHRETYSVFQTLIKPLSTANTSVHSTLTLIYEEYFSK 0 0 TEDDDLELALALSLMEMEDQQPSTQHMQAEVRTNQSGPFQLNSASMSLEKSSIKPEDQVGRENDPVAKSGLKSKESPNSESRDIAPQRGTHENQAQTKQDAFKEPQNLAKDQTLNQPEEP KRSKSRRKRRKGAASLVGLPGSPSASPPVLLWFRRDLRLCDNPALNAALEMDAPVIPIFIWSPEEEEGPGVTVAAGGA 1 2 SKYWLHQALACICTSLNHIGSRLTFLKADGEEKCSLPTLKQLVKETGAKTLLANALYEPWLKERDDMVESTLQKEGVQCRIFPSYCLRDPYSVSTVGVGLR 1 2 GIGSVSHFMSCCKQNPGSAIGVPLDPPQSLPTPSCWPQGVPLEALGLARMPRRKDGTT 0 0 IDWAANIRKSWDFSEDGAHARLDNFLRD 1 2 GVYRYEKESGRADAPNTSSLSPYLHFGQLSPRWLLWDAKGARCRSPKFQRKLAWRDLAYWQLTLFPDLPWESIRPPYK 0 0 ALRWNTDRSHLKAWQRGKTGYPLVDAAMRQLWLTGWMNNYMRHVVASFLIAYLHLPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAALTCDPYGHYVRKWCPELADLPDELIHKPWKCPASMLRRA 1 2 KVVFGETFPERIVVDLEERRNQSLQDVASVRKEFGQFVDKRSGCDLVPLPPRLVAEALGLSQRDGAVGKQFLLPVITRMEFKYQQDDPDAASNPYNAVLKGYV SRKRDETIAFLNQRDFTASVMHEGTQRLERLERDYRRMEGLPPPPSTQGRARRTPTAKDRFSIVPGGAVTS* 0 >CRY7_oreNil Oreochromis niloticus (tilapia) 0 MPPSGADAQATVKWWLGEVLEGREDPEGFFAMCVSILGHRETRSQFLSLIEPLSTGDRLLHASLTSIYQEYFTK 0 0 TEDDELELALALSLLDMKGQPLQSPSQESQPLQSGDAQNQRGSVQMNSILKPEGSNCTNQTGSWKRESAPQTAREMEFQISTYPNKQTLGTDQTVCVSEQEGDVDQSQKPKRSKNRRQRRKCTGQHLVGLPRSPSATPPVLLWFRRDLRLCDNPALVASLEVGAPVIPVFIWSPEEEEGTGVTVAMGGA 1 2 gKYWLHQALLCFCSSLEHIGSHLVFLKANGEMTSAESSLKALKELIKDTGARTVLANALYEPWLKERDDLVVSTLQKQGVEFRMFHSFCLRDPYSVTTEGVGLR 1 2 GIGSVSHFMSCCRQNPGTAVGVPLDPPVSLPTPAHWPKGVCLDELGLARMPRRKDGTM 0 0 VDWAVNIRKTWDFSEGGAQARLEAFLHD 1 2 GVYRYEKESGRADAPNTSCLSPYLHFGQLSPRWLMWDAKGARCRPLKFQRKLAWRDLAYWQLTLFPDLPWESIRPPYK 0 0 ALRWSSDRGHLKAWQRGRTGYPLVDAAMRQLWQTGWMNNYMRHVVASFLIAYLHLPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAAMTCDPYGTYVRKWCPELADLPDELIHKPWKCPASMLRRA 1 2 GVVFGQSYPERIITDLEERRNQSLQDVALVRREFEQYVDKRSGCDLVPLPPRLVSEALGLSHKDGAVVTQGKQFLLPVITRMEFKHQQEDPDADATSNPYNAVLKGYV SRKRDETIAFLNERDFTASVMHEAIQRKERLNSDYRRMEGLPPSPSPRGRARRTPTAKDRFSIVPGGAVTSLK* 0 >CRY7_tetNig Tetraodon nigroviridis (fugu) 0 MPPSAAPDDESNKVWARRILREVLAGREDPEGFFALCLSVLGHQETRSQFLSYIEPLSTANSCLHSVLSSIHRGYFSK 0 0 TEDEDLELALALSLVEMKDCELSAPSQERQLRQPGGGQTGSGCVGPTSVSQTPRRGQICSAGADSREGNNSPEESGDLVTEVGALDPSGKPKRSKKRRRRRQGAGRQVVGLPPAPSAAAPVLLWFRRDLRLCDNPALTAALEVGAPVIPIFIWSPEEEEGPAGVTVATGGA 1 2 CKYWLHQALSCLCASLERNGSHLVFLRSAEAGGEGGSSLQTLKELVKETGARTVMANALYEPWLKDRDHAAAVALWKSGVECKMFHSYCLRDPYSVRTEGVGLR 1 2 GIGSVSHFLSCCRQNPGPALGAPLEPPTALPVPAHWPRGAPLEALGLARMPRRKDGTT 0 0 VDWAASIRKSWDFSEEGAHARLEAFLGD 1 2 GVYRYDKESGRADAPNTSCVSPYLHFGQLSPRWVLWDAKAARCRPPKFQRKLAWRDLAYWQLSLFPELPWESLRPPYK 0 0 ALRWSSDRRHLKAWQRGGTGYPLVDAAMRQLWLTGWMNNYMRHVVASFLIAYLHLPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAAMTCDPDGSYVRKWCPELSGLPDELIHKPWKCPASVLRRA 1 2 GVVLGQTYPERIVTDLEERRSRSLRDVALVRKQFGQYVDERSGCDLLPLPPRLVSEALGSSPQGGVEAAGGRPFLLPLITRMEFKHQQGDPDADAASNPYNAVLKGYV SRKRDEAVAFLHERDFTASVMQEGARRRERMESDQRRLEGLPRASEPRGRARRTPTAKDRFSVVPGGAVTPAR* 0 >CRY7_takRub Takifugu rubripes (fugu) 0 MPPSAAPDGDSTKVWVRRILREVLAGREDPEGFFALCLSVLGHQETRSQYPSHIEPLRAANSSMHSVLTSIYKEYFST 0 0 TEDDELELALALSLVETKDYELSAPVHEPQLKRPGGRRNESGSVMPTSEFQTQRRSPIQPAEAASGTRNNSPQTGPSPKVCSPRMAVEPEHNADAWTPGKARSASCESGHQVMEVGDVDQSEKPKRPKKRRQRRKCAAQQVVGLPRAPSAAAPVLLWFRRDLRLSDNPALVSALKVGAPVIPIFIWSPEEEEGPGVTVAMGGA 1 2 CKYWLHQALSCLRSSLENIGSHLVFLKSETRGSEGGASLRALQGLAKATGARTVVANALYEPWLKGRDDAVAAALQKSGVECEMFHSYCLRDPYSVSTEGVGLR 1 2 GIGSVSHFMSCCRQNPGAALGAPLDPPTTLPVPAHWPPGVSLETLGLARMPKRKDGTT 0 0 IDWAANIRKSWDFSEGGAHARLEAFLHD 1 2 GVYRYEKESGRADAPNTSCVSPYLHFGQLSPRWLLWDAKGARCRPPKFQRKLAWRDLAYWQLTLFPDLPWESLRPPYK 0 0 ALRWSTDRRHLEAWQRGRTGYPLVDAAMRQLWLTGWMNNYMRHVVASFLIAYLHLPWQEGYRWFQ 0 0 DTLVDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAAMTCDPCGQYVRKWCPELSGLPDELIHKPWRCPTSLLRRA 1 2 GVVFGQTYPERIVTDLEERRSQSLQDVALVRKEFGQYVDKRSGCDLVPLPPRLVSEALGSSHLDGGVATGGKQFLLPVITRMEFKHQQEDPDADAASNPYNAVLKGYV SRKRDETIAFLNERDFTASVMHEGAQRRERMESDQRRLEGLPRGPAPRGRVRRTPTAKDRFSVVPGGAVASLR* 0 >CRY7_gadMor Gadus morhua (cod) CAEA01536921 0 MNVKRTIAGAADNIASLSNTLQQLVLGGEDPGGFFGICVSSMGHSETLSVFPSLIQPLASVDPSQHSTLIAIYVEYFSK 0 0 DEEDELEVALALSLLDVKPQQKPACRPEPRPPVWQGGYPAKVGGSIPGGATKSKTTTLPGLSYAQSAATGGRGRRPSENLPKTGSGVPSSTPPTHPSRGRAPLPTDTHSTTSQTT RHKHVPGLPAPPTFHQSDRRQDAVNDDARDPLDTEGTCPDQSQKPKRSKNRRQRGKGCGQRAVVGLPHSPSATPPVLLWLRRDLRLHDNPALIGSLQAGAPVVPVFIWSPEEEEGPGVTMATGGA 1 2 cRYWLHQALACLQGALQLLGSHLVLLKADAPLWGALLGLVGETGARTVLATALYEPWLRERDQRVEAGLRRAGVAWRMVHSYCLRDPYSVSTQGVGLR 1 2 GLGSVSHFISCCEQNPGPALGPSLDPPVSMPTPSQWPPGDPLDALGLARMPRRKDGTV 0 0 IDWAANIRASWDFSEQGAHARLEAFLQD 1 2 GVYRYEKESGRADSPNTSCLSPYLHWGQISPRWLLWDAKSARCRPQKFQRKLAWRDLAYWQLSLFPALPWESLRPAYK 0 0 ALRWSSDRGHLKAWQRGRTGYPLVDAAMRQLWLTGWVNNYMRHVVASFLIAYLHLPWQEGYLWFQ 0 0 DTLLDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAALTCDPGGSYVRKWCPELAALPDDIIHKPWRCPASILRRA 1 2 GVAFGHTYPEPIVTDLEARRGQSLQDVAQVRSTLSEYVDERSGCDLVPLPPRLVAEALGSAGGDQLRTG GEQFLLPVITRMEFKHRRGDPGEDAASNPYDAVLKGYVNRKRDETVAFLNQRDFSASVMNEGAQRRERLDSDMRILEGLPRAPPGRGEPGQLPRPRAKPRRARPTPAAKGQSA* 0 >CRY7_xipMac Xiphophorus maculatus (platyfish) AGAJ01012112 0 MPHSAAGGQAAVRQMLREVLVGREDPEGFFAMCVSILGHQETRLQFLSIIKPLSTANIRLHSVLSFIYQGYFSK 0 0 MEDDELELALALSLLDMKDQQISTPNQMTPHPAVGGNASSVLCNSNAKTQGNSCTQQEDVPGREKSAFVAQSMSGSKENFRENQTYKSTHVEGVDNKSLGTSQALYVPDSSTSFSITNTASVSGQVMKKEDLSKKPRCSKNRRQRRKVAAQQIVGLKRSPSAPPPVLLWFRRDLRLCDNPALIRCLELGAPVIPVFIWSPEEEEGSGVTLAMGGA 1 2 cKYWLHQALSCFCASLGRIGSHLIFLKASGDENKDGSSLHTLKTLVKETGAQTVMANALYEPWLKERDDVVFSALHKEGVECKMFHSYCLRDPFSVSTEGVGLR 1 2 GIGSVSHFMNCCRQNPGASIGVPLDPPVSLPSPTRWPQGVPLDKLGLARMPRRKDGTI 0 0 IDWAENIRKSWDFSEDGAHARLEAFLHD 1 2 GVYRYEKESGRADAPNTSCLSPYLHFGQLSPRWLLWDAKGARCRPPKFQRKLAWRDLAYWQLTLFPDLPWESLRPPYK 0 0 ALRWSTDCRHLKAWQKGKTGYPLVDAAMRQLWLTGWMNNYMRHVVASFLIAYLHLPWQEGYRWFQ 0 0 DTLMDADVAIDAMMWQNGGMCGLDHWNFVMHPVDAAMTCDPYGSYVRKWCPELADLPDELIHKPWKCPASILRRT 1 2 GVVFGQTYPERIVTNLEEQRSKSLQDVALVRKQFEQYVDKRSGCDLVPLPKRLVSEALGVSHWDGAVVTEGKQFLLPVITRMEFKHQLEDPDADAASNPYNAVLKGYV SRKRDETIAFLNETDFTASVMYEGAHRQERLERDYRRMEGLPPAPSNRGRARRTSTAKDKFSVVPGGAIASRN* 0 >CRY7_rudPhi Ruditapes philippinarum (clam) JO112203 gonad transcript missing first half of antenna domain note filter feeder 0 RDMINHYNNPIIFRKSTNYTSDIFDIIKQSGAQTLVMNDVYEPYLKERDDKICKALKSKGIKCERFHSYLLHQPGAIKTEALCMR 1 2 GMGSVTHFMECCRQSDPEPLGIPVDPPNILPVASYRPECIPVSELGLGKLPKRKDGTE 0 0 IDWAKVIRESWDFSEDGAWEALQLFLTE 1 2 GIRNYEKESSRGDKLNTCRISPYLHFGQISPRTVLNEGRYLKSPKFLRKLAWRDLSYWLLSVFPDLPHAPTRPQYQ 0 0 YQRWSNNKVHLKAWQKGNTGYPLVDASMRQLWLTGWMNNYMRHVVASFLLSYLHITWVEGYLWFQ 0 0 DTLLDADVAINAMMWQNGGMSGLDQWNFVMHPVDAAMTCDPKGDYVRKWIPELAKMPEEFIHKPWKCPPSILRRA 1 2 GVQLGVTYPHRIITDLEEAREQSLRDVVDVRQKFEELIDPRTGNDLVKLPNGVIIPVITRKEFKYKTRNPETRDNPHNAVLRGYRSRKRDELVEYLNERDFMASTMKECSMRHERGQKSPQAFVLK* 0 >CRY7_craGig Crassostrea gigas (oyster) HS138673 EIAALPNDFIHQPWKCPPSILRRCGIKLGETYPYRVISDLEGAREQSLTDVVNVRKKHPEFVDRRTGNDLMPLPDGLCVPVITRKEFKYKLHHPEAKDNPHTAVLRGYRSRKRDEAIAFANERDFMASATNESVKLTERRLKATQYEAL* 0
DASH cryptochrome sequences
>DASH_galGal Gallus gallus (chicken) pseudogene in syntenic position with numerous indels, frameshifts, internal stops
0 0
0 2
1 MVVRKGKLEDMVHDVITSLGCVGVVAFHEE 0
0 2
1 IQPDVYTNF*KAVESEVKTWPALQRADQLKPLAGG.EEGCIPTTEDLGQK 1
2 DPGTDPQTAF-LGFNETAAL 0
0 LLFASYNETWSGLIGVD*STKFAPW 2
1 LVLGCLSP*YIFSQIGKYVKERAANHST*W 2
1 AVFELLWWDCF*FVALKYGRRILSLK 1
2 0
0 LGLDWRMGAEWFEYLL 0
0 VDYNVFSN*STWLYSAGIGNHPKDNRKFNVIKQGPDYDGN 0
0 GDCIQLRVTELQGMKEADIHSPWALNSATLSQVGVTLGKTYPQVQSDSP^EWSRQIRQRA 0
0 GRSPHPKGRRGPAHSPLQHKERGTDCFSLKK* 0
>DASH_melGal Meleagris gallopavo (turkey) ADDD01036185 pseudogene in syntenic position
0 0
0 2
1 0
0 2
1 DVYTNL*KAVESEVKVWPALQRADQLKPLA-GSGRKMYPYNKGFG 1
2 0
0 LFASYKEIWSGLTGADNSTKFAPW 2
1 LVLDCLSP*CIFSQVQKYIKERAANQSTFW 2
1 VFELLWWDCF*FVPLKYGRRIFSL 1
2 0
0 LKVDYNVPNN*VT*LYSADIGNHPKENRKFNMIKEGPNDDG 0
0 VPELQGIKGPDIHSPWALNSASVFQVGGTLVETYPQPAV 0
0* 0
>DASH_anaPla Anas platyrhynchos (duck) scaffold1769
0 0
0 VLHWAASNADYLIPLYCFDPRHYAGTHCYGFPKTGPHRLRFLLESVKDLRETLKKKGS 2
1 TLVVRKGKPEDVVRDLITQLGCVGVVAFHEE 0
0 ATQEELDVEKELCQVCEQHGVKVQPFWAATLYHRDDLPFGPIAR 2
1 LPDVYTQFRKAVEAEAKVRPALQLPDQLKPLAPGVEEGCIPTMEDLGQK 1
2 DPGTDPRTAFPCCGGETQALARLQYYFWDT 0
0 NLVASYKETRNGLIGMDYSTKFAPW 2
1 LALGCISPRYIYEQLRKYEKERGANDSTYW 2
1 VLFELLWRDYFRFVALKHGRRIFSVR 1
2 GLQSKEVPWKKDLQLFDCWK 0
0 EGKTGVPFVDANMRELAATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEHLL 0
0 VDYDVCSNYGNWLYSAGVGNDPRDKRKFNVIKQGLDYDGN 0
0 GDFVRLWLPELQGIAGADAHAPWALSSTALRQAGIILGQTYPQPLVRAPEWSRHINQRL 0
0 GRSPHPRGRRGPADAPAQRKDRGIDFYFSRKKDE* 0
>DASH_taeGut Taeniopygia guttata (finch) ABQF01044665 ABQF01044669 ABQF01044671 synteny: ACAA1 DASH MYD66 OXSR1
0 MSGTAGTAICLLRCDLRAHDNQ 0
0 VLHWAQHNADFVIPLYCFDPRHYLGTHCYRLPKTGPHRLRFLLESVKDLRETLKKKGS 2
1 TLVVRKGKPEDVVCDLITQLGSVTAVVFHEE 0
0 ATQEELDVEKGLCQVCRQHGVKIQTFWGSTLYHRDDLPFRPIDR 2
1 LPDVYTHFPKGLESGAKVRPTLRMADQLKPLAPGLEEGSIPTMEDFGQK 1
2 DPVADPRTAFPCSGGETQALMRLQYYFWDT 0
0 NLVASYKETRNGLVGMDYSTKFAPW 2
1 LALGCISPRYIYEQIQKYERERTANESTYW 2
1 VLFELLWRDYFRFVALKYGRRIFSLR 1
2 GLQSKDIPWKKDLQLFSCWQ 0
0 EGKTGVPFVDANMRELSATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0
0 VDYDVCSNYGNWLYSAGIGNDPRDNRKFNMIKQGLDYDGN 0
0 GDYVRLWVPELQGIKGADIHTPWALSSAALSQAGVTLGETYPQPVVTAPEWSRHIHRRP 0
0 GGSPHPRGRRGPAQRKDRGIDFYFSRKKDAC* 0
>DASH_melUnd Melopsittacus undulatus (budgerigar) AGAI01061648 homopolymer frameshift exon 6
0 msgtSARTAICLLRCDLRAHDNQ 0
0 VLHWAQSNAEFVVPLYCFDPRHYLRTHWYGFPKTGPYRLRFLLESVKDLRETLKKKGS 2
1 TLVVRKGKPEDVVRDLITQLGSVSAVVFHEE 0
0 ATQEELDVENELCQVCSQHGVKVYTFWASTLYHRDDLPFRPIAR 2
1 LPDVYTHFRKAVESEAKVRPTLRMADQLQPLAPGVEEGCIPTMEDFGQK 1
2 DPDTDPRTAFPGSGGETQALMRLQYYFWDT 0
0 NLVASYKEVRNGLVGMDYSTKFAPW 2
1 LALGCISPRYIYEQIKKYERERTANQSTYW 2
1 VLFELLWRDYFRFVALKYGRRIFSLR 1
2 GLQSKEVPWKKDLQLFNCWK 0
0 EGKTGVPFVDANMRELAATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0
0 VDYDVCSNYGNWLYSAGVGNDPRDNRKFNMVKQGLDYDGN 0
0 GDYVRLWVPELQGIKGADIHTPWALSSAALSQAGVTLGETYPQPIVTAVEWSRHISQRP 0
0 GRSLHPRGSRRPAHSPVQHKDRGIDFYFSRKKDA* 0
>DASH_chrPic Chrysemys picta (turtle) AHGY01416292 AHGY01416294
0 MSGAAPRTVICLLRSDLRYHDSE 0
0 VLRWAQNNADYVIPLYCFDPRQYLRTYYYGFPKTGPHRLKFLLESVKDLRQTLKKKGS 2
1 NLVVRKGNPEDVVHDLITQLGSVTAVAFHEE 0
0 ATKEELDVEKALQRVCVQHGVKIQTFWASTLYHRDDLPFRHISR 2
1 LPDVYTQFRKAVEAQAKVRPTVQMPDQLKPLAPGVEDGCIPVPEEFGQK 1
2 DPLTDPRTAFPCCGGETQALMRLQHYFWDT 0
0 NLVASYKETRNGLIGIDYSTKFAPW 2
1 LALGCISPRYIYEQIRKYEKERTANQSTYW 2
1 VIFELLWRDYFRFVALKYGTRIFLLK 1
2 GLQDKEVPWKKDPKLFDAWK 0
0 EGRTGVPFVDANMRELAATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDSN 0
0 GDYVRLWVPEIQCVKGGEIHTLWALSNAALSQAGIALGETYPLPVVLAREWSRHINQKP 0
0 NKRDHPPRGRKDPSDTPRQHKDRGMDFYFSRNKDLS* 0
>DASH_anoCar Anolis carolinensis (lizard) XM_003221869 14 exons
0 MASSLPRTALCLLRNDLRCHDNE 0
0 VLHWAQSHADRIVPLYCFDPRHYAQTYHYNFPKTGPFRLRFLLESVKDLRETLKKKGS 2
1 NLVVRKGKPEDVVRDLIIQLGSVASVSFHEE 0
0 ATKEELDVEKALIRVCTEHGVEVQTFWGSTLYHRQDLPFKHISQ 2
1 LPDVYTQFRKAVESQARVRPVLKMEGQMKSLPPEIEEGNIPSLEDFGMT 1
2 EPLRDPRSAFLCSGGETQALMRLQHYFWET 0
0 NLVASYKETRNGLIGLDYSTKFAPW 2
1 LALGCISPRYIYDQIQKYEKERTANQSTYW 2
1 VLFELAWRDYFRFVALKYGRKIFFPR 1
2 GLQDKDISWKKDPKLFDAWK 0
0 EGRTGVPFVDANMRELATTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFESLL 0
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQALDYDGN 0
0 GDYARLWVPELQCIRGGDVHTPWALSNGALSQAGVSLGSSYPVPIIKAPEWGRHINQKP 0
0 SNQGPKGRKGPSHTPKQHKNRGVDFYFSHKKDLS* 0
>DASH_xenTro Xenopus tropicalis (frog) XM_002938001 BQ392047 CX471210 Pmid: 15147276 synteny: ACAA1 DASH MYD66 transcripts AL790297 CR419606 etc
0 MAARARVIICLLRNDLRLHDNE 0
0 VLHWAHRNADQIVPLYCFDPRHYGGTHYFNFPKTGPHRLKFLLESVQDLRNTLKERGS 2
1 NLLLRRGKPEEIIAGLVKQLGNVSAVTLHEE 0
0 ATKEETDVESAVRRVCTQLGVRYQTFWGSTLYHREDLPFRHISS 2
1 LPDVYTQFRKAAETQGKVRSTFQMPDRLKPLPSGLEEGSVPTHQDFDQQ 1
2 DPLTDPRSAFPCCGGETQALQRLHHYFWET 0
0 NLVASYKDTRNGLIGIDYSTKFAPW 2
1 LALGCISPRYIYEQIRKYEKERTANQSTYW 2
1 VIFELLWRDYFRFVALKYGRRIFFLR 1
2 GLQDKDVPWKKDPKLFDAWK 0
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGIDWRLGAEWFEYLL 0
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDAG 0
0 GDYIRLWVPELQQIKGGDAHTPWALSTASLAHSNVSLGETYPYPIVMAPEWSRHINQKPS 0
0 GSWETSARRGKGPSHTPKQHRNRGIDFYFSRNKNV* 0
>DASH_xenLae Xenopus laevis (frog) NM_001090969 BU916216 BX850012 PMID: 15147276 match to xenTro: 483/524 (92%)
0 MCVPSRVIICLLRNDLRLHDNE 0
0 VLHWAHRNADQIVPLYCFDPRHYVGTHYFNFPKTGPHRLKFLLESVRDLRITLKKKGS 2
1 NLLLRRGKPEEVIEDLVKQLGNVSAVTLHEE 0
0 ATKEETDVESAVKQACTRLGIKYQTFWGSTLYHREDLPFRHISS 2
1 LPDVYTQFRKAVETQGKVRPTFQMPDKLKPLPSGLEEGSVPSHEDFDQQ 1
2 DPLTDPRTAFPCSGGESQALQRLEHYFWET 0
0 NLVASYKDTRNGLIGLDYSTKFAPW 2
1 LALGCVSPRYIYEQIGKYEKERTANQSTYW 2
1 VIFELLWRDYFRFVALKYGRRIFFLR 1
2 GLQDKDIPWKRDPKLFDAWK 0
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGIDWRMGAEWFEYLL 0
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDSG 0
0 GDYIRLWVPELQQIKGGDAHTPWALSNASLAHANLSLGETYPYPIVMAPEWSRHINQKPA 0
0 GSWEKSARRGKGPSHTPKQHKNRGIDFYFSRNKDV* 0
>DASH_hymCut Hymenochirus curtipes (frog) fragment
1 QMPKLKPLPDGLEEGKVPTYEEFDQE 1
2 DPLTDPRTAFPCSGGETQALQRLHHYFWDT 0
0 NLVPPYKDTRNGLIGLDYSTKLAPW 2
1 LALGCVSPRFIYEQIKKYEQERTANQSTYW 0
0 VIFELLWRDYFRFVALKYGRRIFFLR 1
2 GLAHTNV* 0
>DASH_ambMex Ambystoma mexicanum (axolotl) JV207726 TSA 73% duck
0 MSVQARTIICLLRNDLRFHDNE 0
0 VLLWALKNAERIVPLYCFDPRHYVGTHNYNFPKTGPHRLKFLLESVKDLRETLKKKGS 2
1 NLLVRKGKPEDVIQELLKQLGSVSAVAFHNE 0
0 VTKEELDVEAAVKQVCLGPGIKIQTFWGSTLYHRDDLPFRHISR 2
1 LPDVYTQFRKGVESQGTVRPTLQMPQKVPPLPSGLEEGTIPTASDFGQE 1
2 GPLADPRSAFPCSGGETQARSRLQHYFWDT 0
0 NLVASYKDTRNGLIGLDYSTKFAPW 2
1 LALGCISPRYIYEQIQKYEKERTANQSTYW 2
1 VIFELLWRDYFRFVALKYGRKIFFLN 1
2 GLQDKEVPWKKDIQLFDAWK 0
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDCN 0
0 GEYIRLWVPELQGVSGGDAHTPWALSSASLANANVSLGETYPLPIITAPEWGRHINQKP 0
0 NDRGSTGRRTRGQPYTPKQHKNRGVDFYFSHNKDLH* 0
>DASH_latCha Latimeria chalumnae (coelocanth) AFYH01055296 AFYH01281932 probable pseudogene
0 0
0 2
1 0
0 2
1 1
2 0
0 2
1 2
1 VIFELLWRDYFRSVALKYGRRIFLLE 1
2 GLQEKEIPWNKRPELFDVWK 0
0 ynerGVPFVDANMRKLTLTGFISN---QNVASFLTKDLGLDWRMGAEWFEYLL 0
0 IVHDVCSNYGNWLYSAGIGNDTRENRKFNVVKEGLD*DSN 0
0 0
0 * 0
>DASH_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01010414 AHAT01010415
0 MSTIRTIICLLRNDLRFHDNE 0
0 vfHWAQGNAEQIVPLYCFDPRHYLGTHCYNFPKTGPFRLRFLLESVKDLRDTLKKNGS 2
1 NLLVKRGKPENVVSDLIKQLGSVTAVAFHEE 0
0 VTKEEQDVETEVTRVCAQFKVRVHTCWGSTLYHREDLPFNHIAR 2
1 LPDVYTQFRKAVETQSQVRPTIRMPEQLKPFPHNLEEGPIPTPEDLGQP 1
2 DAVSDPRTAFPCIGGETQALARLKHYFWDT 0
0 NLVASYKETRNGLIGMDYSTKFAPW 2
1 LALGCISPRYIYEQIKQYEKERTANQSTYW 2
1 VIFELLWRDYFRFVAVKYGNRIFTVK 1
2 GLQQKSVSWKKDVKLFDAWK 0
0 EGKTGVPFVDANMRELALTGFMSNRGRQNVASFLTKDLGIDWRMGAEWFEFLL 0
0 VDYDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0
0 GDYVRLWVPELRGLKGGDVHAPWVLSSASLADAEVSLNQTYPVPMVVAPEWSRHTTSKA 0
0 SGGGLSQKGKRGPSHTPKQHRDRGIDFYFSRNNKKLQ* 0
>DASH_danRer Danio rerio (zebrafish) NM_205686 chr24
0 MSASRTVICLLRNDLRLHDNE 0
0 VFHWAQRNAEHIIPLYCFDPRHYQGTYHYNFPKTGPFRLRFLLDSVKDLRALLKKHGS 2
1 TLLVRQGKPEDVVCELIKQLGSVSTVAFHEE 0
0 VASEEKSVEEKLKEICCQNKVRVQTFWGSTLYHRDDLPFSHIGG 2
1 LPDVYTQFRKAVEAQGRVRPVLSTPEQVKSPPSGLEEGPIPTFDSLGQT 1
2 EPLDDCRSAFPCRGGETEALARLKHYFWDT 0
0 NAVATYKETRNGMIGVDFSTKFSPW 2
1 LALGCISPRYIYEQIKKYEVERTANQSTYW 2
1 VIFELLWRDYFKFVALKYGNRIFYMN 1
2 GLQDKHVPWKTDMKMFDAWK 0
0 EGRTGVPFVDANMRELALTGFMSNRGRQNVASFLTKDLGLDWRLGAEWFEYLL 0
0 VDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0
0 GDYVRQWVPELRGIKGGDVHTPWTLSNSALSHAQVSLNQTYPCPIITAPEWSRHVNNKS 0
0 SGSSSSKGRKGSSYTARQHKDRGIDFYFSKNKHF* 0
>DASH_oreNil Oreochromis niloticus (tilapa) GL831138
0 MSSSRTVICLLRNDLRLHDNE 0
0 LFHWAQRNAEHIVPLYCFDPTHYVGTYNYSLPKTGPFRLRFLLEGIRDLRNTLINKGS 2
1 NLVVRRGKPEEVVADLIRQLGSVSSVAFHEE 0
0 VTSEELNVEKRVKDVCAQMKVKVHTCWGSTLYHRDDLPFPHMSR 2
1 LPDVYTQFRKAVESEGRVRPVFSTPEKLNPLPPGLEEGAIPTAEDLQQT 1
2 EPVTDPRSAFPCGGGESQALARLKHYFWDT 0
0 DAVATYKETRNGLIGVDYSTKFSPW 2
1 LAMGCISPRYIYHQIKQYEKERTANQSTYW 2
1 VIFELLWRDYFRFVSVKYGNRIFQVK 1
2 GLQDKSVPWKKDMKLFDAWK 0
0 EGRTGVPFVDANMRELAATGFMSNRGRQNVASFLTKDLGLDWRMGAEWFESLL 0
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0
0 GDYVRGWVPELQNIRGGDVHTPWALSTAVLAHAHVTLGDTYPAPIVTAPEWSRHVNKKA 0
0 GGTGPSPRGKKGPSHTPKQHRDRGIDFYFSRSKNL* 0
>DASH_anoFim Anoplopoma fimbria (sablefish) JO682130 JO685461
0 MSASRTVICLLRNDLRLHDNE 0
0 LFHWAQRNAEYIVPLYCFDPRHYVGTYNYNLPKTGPFRLRFLLESIRDLRNTMLTKGS 2
1 NLVVRRGKPEEVVADLIKQLGSVSTVAFHEE 0
0 VTSEELNVEKRVKDVCAQMNVKVHTCWGSTLYHRDDLPFHHMSR 2
1 LPDVYTQFRKAVETQSRVRPLFPTPEQLKPLPKGLEEGAIPTAEDLEQT 1
2 EPLADPRSAFPCgGGEsqALARLKHYFWDT 0
0 DAVATYKETRNGLIGADYSTKFAPW 2
1 LAMGCISPRYIYHQIQQYERERTANQSTYW 2
1 VIFELLWRDYFKFVGVKYGNRLFQIK 1
2 GLQEKSVPWKKDMRLFNSWK 0
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAELFEYLL 0
0 VDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDGN 0
0 GDYVRQWVPELQGIKGADVHTPWTLSTASLSHAHVSLGETYPTPIVMAPEWSRHANKKP 0
0 SGTGPSPRGKKGASHTPKQHRDRGIDFYFSKSKNL* 0
>DASH_takRub Takifugu rubripes (fugu) CAAB02011066
0 MSGPRTVICLLRNDLRLFDNE 0
0 LFHWAQRNADHIVPLYCFDPRHYMGTYHYNLPKTGPFRLRFLLESIKDLRNTLLNKGS 2
1 NLIVRRGKPEEVVASLIKQLGSVSTVAFHEE 0
0 VTSEELDVEKRVKDVCAQMKVNVHTCWGSTLYHRDDLPFHHISR 2
1 LPDVYTQFRKAVESQCRVRPVFPPPEHLKPLPQGLEEGTILTAEDLEQK 1
2 EPVADPRSAFPCSGGESQALARLKHYFWDT 0
0 DAVAVYKETRNGLIGVDYSTKFSPW 2
1 LALGCISPRYIYHQIKQYESERTANQSTYW 2
1 VIFELLWRDYFRFVAVKYGTKLFQVN 1
2 GLQDKSVSWRKDMKLFNAWK 0
0 EGKTGVPFVDANMRELATTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0
0 VDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDNN 0
0 GEYVRLWVPELQRIMGADVHTPWTLSSAILSHAHLSLGETYPTPIIVAPEWSRHFNKKM 0
0 SGAGPSPRGRKGPSHTPKQHRDRGIDFYFSRSKNL* 0
>DASH_gasAcu Gasterosteus aculeatus (stickleback) chrXXI DN658701
0 MSTSRTVICLLRNDLRLHDNE 0
0 LFHWAQRNAEYIVPLYCFDPRHYVGTYNYNLPKTGPFRLHFLLEGVKDLRNTMLSKGS 2
1 NLVVRRGKPEEVVADLIKRLGSVSTVAFHEE 0
0 VTSEELSVEKRVKDVCARMKVKVHTCWGSTLYHRDDVPFHHISR 2
1 LPDVYTQFRKAVETQSRVRPLFPTPEQLKPLPEGLEEGAIPTAEDLEQT 1
2 GPVTDPRSAFPCSGGESRALARLKHYFWDT 0
0 DAVATYKETRNGLIGMDYSTKFAPW 2
1 LAMGCISPRYIYHQIQQYEKERTANQSTYW 2
1 VIFELLWRDYFKFVAVKYGNRLFQIN 1
2 GLQEKSVPWKKDMTLFNAWK 0
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAELFEYLL 0
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDGN 0
0 GDYVRRWVPELQGIRGADVHAPWTLSTASLSHAGVSLGETYPNPVVVAPEWSRHVNKKP 0
0 SGARPSPRGKKGPSHTPKQHGDRGIDFYFSRSKNL* 0
>DASH_oryLat Oryzias latipes (medaka) FS553506
0 MASSRIVICLLRNDLRLHDNE 0
0 LFFWAQKNADHIVPLYCFDPRHYVGTYNFNFPKTGPFRLRFLLDSVRDLRNTLLSKGS 2
1 NLVVRRGKPEEVVADLIKQLGSVSSVAFHEE 0
0 VASEELNVEKKVKEVCAQMEVKVHTCWGSTLFHRDDLPFPHMAR 2
1 LPDVYTEFRKAVESKSRVRPVFPTPDRLNSLPPGLEGGAIPTAEDLEQT 1
2 EPETDPRSAFPCSGGESQALARLKHYFWDT 0
0 DAVATYKETRNGLIGVDYSTKFSPW 2
1 LAMGCISPRYIYHQIKKYEQERTANQSTYW 2
1 VIFELLWRDYFKFVGVKYGNKMFFIK 1
2 GLQDKSLPWKRDTKLFDAWK 0
0 EGRTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQALDYDSN 0
0 GDYVRQWIPELTAVRGGDVHTPWNLSSAALSRARVSLGETYPTPIVTAPEWGRHFNKKP 0
0 SGPGPSQKGKKGPSHTPKQHRDRGIDFYFSKSKNL* 0
>DASH_dicLab Dicentrarchus labrax (seabass) CBN81995
0 MSTFRTIICLLRNDLRFQDNE 0
0 LFHWAQRNAEYIVPLYCFDPRHYVGTYNYNLPKTGPFRLRFLLDSIRDLRNTLLSKGS 2
1 NLVVRQGKPEEVVADLIKQLGSVSAVAFHEE 0
0 VTSEELNVEKGVKDVCAQMKVKVHTCWGSTLYHRDDLPFHHISR 2
1 LPDVYTQFRKAVETQSRVRPVFPTPEQLKPLPSGLEEGAIPTAEDLQQT 1
2 EPLTDPRSAFPCSGGESQVLARLKHYFWDT 0
0 DAVATYKETRNGLIGVDYSTKFAPW 2
1 LAMGCISPRYIYHQIKQYEKERTANQSTYW 2
1 VIFELLWRDYFKFVGVKYGNRLFQVK 1
2 GLQDKSIPWKKDMKLFNAWK 0
0 EGQTGVPFVDANMRELAMTGFMSNRGRQNVASFLTKDLGLDWRMGAEWFEYLL 0
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDSN 0
0 GDYVRQWVPELQGIKGADVHTPWTLSTAALSHAHVSLGETYPNPIVIAPEWSRHVNKKP 0
0 SGTGPSQRGKRGPPHTPKQHRDRGIDFYFSRSKNL* 0
>DASH_patPec Patiria pectinifera (starfish) HP101597
0 MAGRMKIIICLLRNDLRYRDNE 0
0 VLHWAHQNADILPLYCLDPRHYHGTWHFSFPKTGPHRLKFLLESLRDLKETLRQVGS 2
1 NLVISRGKPETVIQDLIRQLGQDNVKAVALQQE 0
0 ATREELDVEAAIEKSCQVPVHKFWGSTLYHKDDIPFRIRD 2
1 LPNVFTEFRKKTENQSDVRPTLPMPSKLKPLPPGIDPGHAPSVEEMGMK 1
2 DPASDERTAFPYSGGEKQALARIQHYFWDY 0
0 DYVSSYKETRNGLIGGDYSTKFSSW 2
1 LALGCLSPRKIYEEIRKYEKERRANQSTYW 2
1 VVFELLWRDYFKFVALKFGDRLFYLS 1
2 GLLGNHREWRENKALFDAWR 0
0 EGRTGVPFVDANMRELAATGFMSNRGRQNVASFLTKDLKLDWRLGAEWFEHLL 0
0 IDHDVCSNYGNWLYCAGVGNDPRQDRKFNMIKQGLDYDAQ 0
0 GDYVRLWVPELQGIKGGAVHTPWILSSSMLIQANVRLGHTYPLPVVKAPEWNKHMRP 0
0 QERGKSGKGPGQRQQTRGIDFYFSSGKQGKFKK* 0
>DASH_strPur Strongylocentrotus purpuratus (urchin) SPU_027694 JT098657 last exon compositionally simple expansion
0 MAGRMKIIICLLRNDLRYRDNE 0
0 VLFWAHKNATNVIPLYCFDPRHYKGTHQFGFPKTGPHRLKFLLESVRDLRTTLQSVGS 2
1 GLVVRSGNPEDVVPQLIQQFGEGEVAAVALQEE 0
0 ATREELDVEAGLHQSCSKLGVNVKKFWGSTLYHREDVPFSPG 2
1 VPNVYTEFRKKVENRSHVRPPINMPKPLKGLPEGVEEGDIPVFSSFGLK 1
2 DPGSDSRTAFPFQGGETTGLARIEDYFWKS 0
0 DCIAKYKETRNGLIGSEYSTKFAAW 2
1 LAHGCVSPRQIHAEVKRYEKERTENQSTYW 2
1 VIFELIWRDYFKFVALKFGDKMFYLS 1
2 GLLGVHKPWNKNKQWFDAWC 0
0 KGQTGVPFVDANIREMAATGFMSNRGRQNVASFLTKDLVLDWRLGAEWFEQML 0
0 VDHDVTSNYGNWLYSAGVGNDPRQDRKFNMIKQGLDYDPN 0
0 GDYIRLWVPELSGIKGGSIHMPWTLSSAQLNQAGVSLGETYPSPIVTAPEWSRHSKRP 0
0 QSGRGGAGGSQGGGSQGGGRNQSGKGQRRNQGPPRGQQRGLDFYFSNPSQK* 0
>DASH_aplCal Aplysia californica (sea_hare) scaffold_151:75,790-145,485 mollusk
0 MSSFSSSPVNTVIYLFRNDLRVHDNE 0
0 ALLLANQKGTRLLPLYCFEPRHYSGTYHFGFPKTGNHRLSFLLDSVKDLQKNLKSRGS 2
1 DLVIRKGKPEEVIPDLLRTLNVTRDAAVVFQKE 0
0 VMEEEVKVEKAIEAHVNVPVNTTWGHTLYHVEDLPFQPI 2
1 LPDVYTQFRKKVEDNTTIRKCVHVPDALKPLPEGIEVGSLPSYEDLGVS 1
2 EPEKDSRAAFPFLGGETTALERLHSYLWGTDNVSTYKETRNGMIGSDYSTKFSPW 2
1 LAHGCLSPRKIYWEIKKYEKERTSNQSTYWVIFELIWRDYFRFVGLKYGNKLFHSG 1
2 GIKGDRVEWKVNKEQFKAWQ 1
2 EGRTGVPYVDANMRELMATGFMSNRGRQ 0
0 NVASFLTKDLHLDWRLGAEWFESML 0
0 IDHDVCSNYGNWLYSAGIGNDPRENRKFNVVKQGLDYDAE 0
0 GDYVRLWVPELAGIKTGSVHCVWTLSPAVLESGDVSLGQTYPLPLVKAPEWSRHVNRP 0
0 GSSGGGRDKGPGSARGRGQNKGSHRGQGHQSQQAGQKRGLDFYFSSSKPH* 0
>DASH_vilLie Villosa lienosa (mussel) JR504188 transcript assembly mollusk
0 MSKTLTVICLLRNDLRIHDNE 0
0 VLYWANKYADYVLPVYCFDPRHFKGTHHFNFPKTGPHRLRFLLESIKDLRKNLQSCGS 2
1 GLVVKQGEPEVVILDLIQKLGKDKVHALAYQTE 0
0 ATKEELDVEETLRAKCGVKIQTFWGHTLFHRDDLPFGVGQ 2
1 LPDVYTQFRKQVEGECVVRDTIDMPRTFKPLPPGIVLDEIPTAEALGVK 1
2 EAVSDPRSVFPWSGGETSALERLNQYLWNTDKVATYKETRNGMIGADYSTKFSSW 2
1 LALGCLSPRVIYWEIKRYEKERASNQSTYWVVFELLWRDYFRFVALKYGNRIFYLS 1
2 GIQGKYIEWKQDTKLFDAWR 0
0 DGKTGVPYVDANMRELATTGFMSNRGRQ 0
0 NVASFLTKDLKLDWRLGAEWFESML 0
0 IDHDVCSNYGNWLYSAGIGNDPREDRKFNMVKQGLDYDPN 0
0 GDYVRLWIPELAGVKDGSIHTVWTLNSGALSRAGVSLGKTYPHPILIAPEWNRHMGRT 0
0 KPGMGRGTGSQPNKLKGIDFYFSSGTQRQ* 0
>DASH_rudPhi Ruditapes philippinarum (clam) JO106851 mollusk fragment
1 YYEIKRYEQERVANNSTYWVIFELLWRDYFRYVALKYGNRLFYLS 1
2 GIQGKQVPWKQDRELFRAWK 0
0 EGRTGVPYVDANMRELAATGFMSNRGRQ 0
0 NVASFLTKDLKLDWRLGAEWFESML 0
0 IDHDVCSNYGNWLYSAGLGNDPREDRKFNMIKQGLDYDPE 0
0 GDYVRLWVPELQNVSGGNVHTVWTMSNNALAKLGISLGETYPNPITVAPEWSRHYGKS 0
0 GGASARGGGGGRGRGGYSGGGQSHGHSGRKESAGQKRGIDFYFKGS* 0
>DASH_dapPul Daphnia pulex (water_flea) ACJG01000530 FE360630 FE400207 FE361270 crustacean mostly re-intronated
0 MSNRVAICLFRNDLRYHDNE 0
0 VlALAHKSADFVLPLYCFDPRHFEGTHHYKFPKTGIFRTQFLLESVEDFRQTLVKRGSNLMIVHSKPEEALLKIFKSLTGLKVTLILQTEVTKEETDVEKcLQKICQEIKASYINCWG
STLYHKGDLPFQINHVPDSYTGFRKDVEEKLRIRPEISMPDKMKPVPTFAHEIPWGNLPTIEALNSTKPIPNSSSAFPFNGGETAALLRLKSYLWDTNAVAQYKETRNGLIGSDYSTKFSS
WLSHGCLSPRRIHWELEKYELQRTKNQSTYWVRFELLWRDYFKFVSMKYGDRIFYPNGMKGRRQQWKKDMELFKAWQ 1
2 MGKTGVPFVDANMRELLATGWMSNRGRQNVASFLVKDLLLDWRLGAEWFESLLLDHDVCSNYGNWNYVA 1
2 GIGNDPRENRKFNMIKQSMDYDLEGNYIRMWVPELREIPGSKIHSPWMLSSGALSAAKIRLGDNYPNPVVVAPEWSRHQKGGK 0
0 DFGQGNPKGGTQRGIDFYFKNPGGQK* 0
>DASH_celPug Celuca pugilator (fiddler_crab) JO491098 distal fragment
0 WVLFEMIWRDYFKFVCMKFGDRVFYPSGIMGKKTVWKQNHELFKKWK 1
2 EGRTGVPFVDANMRELRETGWMSNRGRQNVASFLIKDMGLDWRLGAEWFESQLVDHDVCSNYGNWNYSA 1
2 GIGNDPRENRKFNMIKQAFDYDPEVTCAVLVLSWLG
>DASH_bemTab Bemisia tabaci (whitefly) EZ942653 HP660316 Hemiptera but phylogenetic isolation and best-blast suggest fungal contamination
MASPKILIYLLRRDLRVHDNPIFHKLTSMSSQANAPFTHLLPLYVFPAHQIEVSGFLSSSDEKSPYPEARSKVGKFWRCGQLRAKFLVESVWDLKQNLETIGSGLEIRVGMLHDVVKQL
VEGFKSKGVQVKGLWMTSEEGYEEKAEERQVRKIIVNAGGDFHLWKDEKYFIDDDDIPFDDPQKYPDVFTKYRNTVEPLREAPRKVLPTPKKLPPLPQNIPPQAHPFKIPGNLKDLIAAL
QKPLDAGLGLKNPPQMPSAGASSAVPFAGGATSGQKRLKHLIESGAMTRYKDTRNGMVGTDYSSKLSLWLALGSLTAREVHSALIDFEEGKTDVGKGAEGYGKGENKGTTHMRFELLWRD
YMRLCTRKYGSRLFLVGGFRNARNIQWKHDNSIMQRWLEGTTGIGLVDAAQRELFLTGFTSNRARQNVASFLTKHLEQDWRLGAEWYECNLVDYDVSSNWGNWQYTAGVGNDPREDRKFN
PVKQASDYDPKAEFVKAWIPEVRELQPEEAWQCWKASYAAKQKPGLRGNIMAERPLAKISFTPRSGGNDGHRGGGRSRGRAQWF* 0
>DASH_acrMil Acropora millepora (coral) cnidarian JT000937 transcript single exon based on A. digitifera BACK01017766
MDAQTSNSIAIYLIRNDLRVHDNECLSWAQQNADFVIPLFCFDKEIFGHGAETWHFKFPKTAIHRARFILDSVVDLKDSLKKGGSDLLLRSEQTMKEAVL
GVIQLCRQQSIQNLSLVYQREIAKEEKDVEAELLELCKKENVAVKSFWGLTLYHVEDLPFASVRHLPDTYTEFRKSVEARCRVRPMIPAPNRLKAIPDFI
TSDNMGGIPTLTELVKDQGTTPDARSAFPFCGGETAAIERMNNYLWTTDNVSKYKETRNGLIGAEYSTKLSPWLAVGALSPRKIYECVKQYEKERTANQS
TYWVLFELLWRDYFKFVCFKFGDSVFYLSGIMKKVGLSWKQDMSSFDKWRFGQTGVPFVDANMRELLYTGWMSNRGRQNVASFLVKDLGLDWRLGAEWFE
SLLVDHDVCSNYGNWNYSAGIGNDPRENRKFNMIKQAFDYDADGDFVRLWVPELAGLKGAKVHIPWTLTSSELKVAGITLGESYPRPMVNPPEWKRHTSK
VKGPGNSTRRHNRGIDFYFKSPKDTNPSKKRH*
>DASH_nemVec Nematostella vectensis (sea_anemone) XP_001623243 ABAV01026885
0 MATSKTQKTVICLLRNDLRYHDNEALLWAHKNADFVLPLYCFDPDHYKTTWRFSLPKTAQYRAKFLLESVTDLRSTLQIHGSNLIIRQCQPLEAVTKLTE
LLKPVAPVTSIVFQQEVTYEELNVEKALVEFCKKSGIHMHTIWGSTLFHKDDIPYKAKTVPDTYTQFRKGVENQSTVRNLIDMPKNLKPLPPVKGELGTI
PDLKSLLNDSEIKEVDQRSAFPFMGGEQEALSRLGSYLWGTDSVAKYKETRNGLLGENYSTKLSPWLANGSLSPRMVYHRIKQYEEERVANHSTYWVLFE
LIWRDYFKFVCLKYGDRVFYRSGIMGKSLPWKHDKMTFKLWCEGKTGVPFVDANMRELKETGWMSNRGRQNVASFLIK 0
0 DLGLDWRYGAEWFESLLLDHDVCSNYGNWNYAAGIGNDPREGRKFNMVKQGLDYDPDGDYIKTWVPELAKIPGAKVHVPWTLKPGELR 2
1 SSGVELGVTYPNPIVIAPEWARHTSKTISGKKPSNFNPRQKGVDFYFKSANNGPSGSR* 0
>DASH_hydMag Hydra magnipapillata (cnidarian) XM_002166508 single exon ABRM01055505
MAGFSSRKTIIYWIRNDLRLHDNECLNFACRDGSHVLPLYCFDPDHYKLTHHFGFLKTGIHRAKLILDAVSGLRNSLQKKGSNLVISKHRPKEAIEKLIL
ECEQTAPVTSIVFQKEITKEETDVEASVFKVCDSKGIKVLSFWGSTLYHIEDIPFQINQIPNTYTLFRKGVEKCPVRKLNPIPDKISSLPNLKNFPLGEV
PLLNELGFDKNEYEIDVRSVFPFNGSEVDAIKRLNEYLWGNNAILTYKETRNGLLGKDYSTKFSPWLALGSLSPRMIYHKIKDFEEQKKIKENDSTYWVV
FELIWRDYFKFICSKHGNKVFFKGGLKGGKGPWTWKQDLNLFMKWANGDTGIPFVDANMKELLATGWMSNRGRQNVASFLVKDLNLDWRMGAEWFESLLI
DSDVCSNYGNWNYIAGIGNDPRSNRKFNVIKQAADYDEEGDYVRTWLPVLKKLGICHTPWRLSASELNAGGVDLGKNYPHPIIVAPEWAKHALKMNSKNC
FHEHRGKQNGQKRSIDFYFKSGKTGE* 0
>DASH_monBre Monosiga brevicollis (choanoflagellate) XP_001745157 ABFJ01000402
0 MAKAGNPRPVVVWFRNDLRVHDNEVLLQAAK 0
0 ASHNHVVPVYCFDIRQ 0
0 YSLVITHRSRRCGQFPKCGRPRARFLIESVDDLRTRLQELGSGLVVRTGLPEEEVARVAAQVGATQVFAHQEVCSEEVAAEHRLKRQLEVPLSLHWGAVTLCHLDDLDFGPRCKHLPSVFTQFRKRVEADMHVRPVVAAPARLAPLPSDLELGSIPTVEDLCPGQH 0
0 EPDERAVLPFKGGETAARARLQYYLWESNLLAS 2
1 YKDTRNGLVGGDYSSKFSPWLAHGNLTARWIYHE 0
0 VKRYEQERTENTSTYWLIFELLWRDYFRFVALQHGTAIFKP 1
2 GGVQHKDVPWRHSPADFEAWQNGQTGFPFIDANMRELAKTGFMSNRGRQNVASFLTK 0
0 DLQIDWRLGAEWFETLLLDHDPCSNY 1
2 GNWNYAAGVGNDPRQGRHFNVIKQ 1
2 AKTYDPTAEYVHLWVPELRGLDAPQAHVPFK 0
0 LSSEELAQANIQLGSTYPRPVVDRLDGGRLPLKVRDETAYGPHVQLPQQGLIKMILFDTLFTGLSW* 0
>DASH_araTha Arabidopsis thaliana (cress) NM_122394 AFMZ01019177 aka:CRY3 PDB:2VTB
0 MNDHIHRVPALTEEEIDSVAIKTFERYALPSSSSVKRKGKGVTILWFRNDLRVLDNDALYKAWSSSDTILPVYCLDPRLFHTTHFFNFPKTG 1
2 ALRGGFLMECLVDLRKNLMKRGLNLLIRSGKPEEILPSLAKDFGART 0
0 VFAHKETCSEEVDVERLVNQGLKRVGNSTKLELIWGSTMYHKDDLPFDVFDLPDVYTQFRK 0
0 SVEAKCSIRSSTRIPLSLGPTPSVDDWGDVPTLEKLGVEPQE 0
0 VTRGMRFVGGESAGVGRVFEYFWKK 0
0 DLLKVYKETRNGMLGPDYSTKFSPWLAFGCISPRFIYEE 0
0 VQRYEKERVANNSTYW 2
1 VLFELIWRDYFRFLSIKCGNSLFHL 1
2 GGPRNVQGKWSQDQKLFESWRDAKTG 2
1 YPLIDANMKELSTTGFMSNRGRQIVCSFLVRDMGLDWRMGAEWFETCLLDYDPCSNYGNWTYGA 1
2 GVGNDPREDRYFSIPKQ 0
0 AQNYDPEGEYVAFWLQQLRRLPKEKRHWPGRLMYMDTVVPLKHGNGPMAGGSKSGGGFRGSHSGRRSRHNGP* 0
>DASH_phaTri Phaeodactylum tricornutum (diatom) XM_002178853 CPF2
MSSSRSKQSFWFLIFVHTVLILTFAAGAATAMRGASVSTVTKATSRASSQVVLHWFRHGDLRLLDNPALIHSSKTAESCVPVFCFDDSVYGNDNRTPDTRAP
HSNDRGQLKCGPRRAQFVLDSVQDLRRSLQSRGSALYVAHGKPAQVFQRLVDAWPAVPAADTAAPNGSLLTIVCQREVVREENDAVRAVQSVLRRRFPQAKVQQIWGSTMYELDDL
PFATDLANMPDTFTPFRNKVEKNCQIGTPLPVPKQLSLPENFPSALKQGLEYLPTLKELGYTDAQIQQVETHDERGVLHFMVAKQPDSHDCLKDYFETRNGMLGPNYSTKFSPWLA
HGNVSPRYVAAQCRKYEEERVENKSTYWVVFELLWRDFCKFFATKHGDAIFYPYGTTERTDRHKPWSTFGRNLQAWQEGRTGYPLVDANMRELVATGFMSNRGRQNVASFLAINLN
HDWRCGGDFFESHLLDYDVYSNWVNWCAAAGMTGGRLNRFNISKQSKDYDQHGDYVRHWLPELAKVPNEFVHEPWKMTSFQQMEYECKLGVDYPNPIVPPSRPNPHTDRNNRGRGG
HQSKGNNRHGPPDKSRKANASGSNRHQKYEMKSLQPGSFRVKES* 0
>DASH_thaPse Thalassiosira pseudonana (diatom) XM_002291289
MASSATQYKLNIHLFRQTDLRLHDNPALCHTVDLSLGTSSSDAASTAPTGILPVFVFDTTRIYGSNAKSRLGNNLKCGPRRARFAIEAVADLRSNLEKRGSG
LVVAVGSPESILARIATAAVESSGDKGALINVVCQEEVCSEELSVDKAIRSSLAKTMKSTSKFTFETVWGSTMYDPETLPFDGGVFGVPDIFSPFRKDVEAKCQIVNDANNKCSLK
YMPTLSDLGYNAEDIESVSSVDSRTALPVNYRGGETFALRRVKDYIWDKDLLKLYFDTRNGMIGSDYSTKFAPWLALGNVSPRYIARECSRYEEQRVQNKSTYWVVLELLIRDHFK
FFAKKHGDKVFYLTGNIGKQRGSERKWGMDRKKIQAWQDGMTGFPLVDANMRELAATGFMSNRGRQNVASFLTIDLNIDWRYGGEYFEETLLDYDVHSNWGNWCSAAGMTGGRLNR
FNIVKQSKDYDFGGEYVRLWCPELKNVPDKHIHSPWEMSGEMMEECGVKIGHGGDYPSPIVDPGAGPKAVANSNSRGGGRGDQGKYNINTNKGRGQRQDMKSLKTGSYRFAASESESKYYE* 0
Cryptochrome CPD photolyases
>CPD_monDom Monodelphis domestica (opossum) NP_001028149:wrong OPC1 PubMed:7937136 synteny: TNK1 MUC4 CPD KIAA0226 FYTTD1 0 MAPKKRSLGDGDEQEKTETQENKTKRKPLQKHQFSKSNVVQKEEENKREGEKKGGAEGLQEVVMQSRLQTASSVLEFRFNKQRVRLISQDCHLQDHSQAFVYWMSRDQRVQ 1 2 DNWAFLYAQRLALKQKLPLHVCFCLAPCFLGATIRHYDFMLRGLEEVAE 0 0 ECEKLHIPFHLLLGLPKDVLPAFVQAHSIGGIVTDFSPLLHHTQWVKDVQDGLPKQVPFVQ 0 0 VDAHNIVPCWIASDKQEYGARTIRHKIHDRLPHFLTEFPPVICHPYPSNIQAE 0 0 PVDWNACRAGLQVDRSVKEVSWAKPGTASGLTMLQSFISQRLPYFGSDRNNPNKDALSNLSPWFHF 1 2 GQVSVQRAILEVQKHRSRYPDSVANFVEEAVVRRELADNFCFYNKNYDKLE 1 2 GAYDWAQTTLRLHAKDKRPHLYSLEQLESGKTHDPLWNAAQ 0 0 MQMVQEGKMHGFLRMYWAKKILEWTRSPEEALEFAIYLNDRFQLDGRDPNGYV 1 2 GCMWSICGIHDQGWAEREIFGKIRYMNYAGCKRKFDVAEFERKYSPAD* 0 >CPD_sarHar Sarcophilus harrisii (tasmanian_devil) AEFK01107967 0 MAPKKRSHGTGSEPEKTDSQESKAKRKPLQKHPSSKNNVVQPEREDKKEGQKRAGAEGLQEAVRQARLRTAPSVLEFRFNKQRVRLISQDCHLQDHSQAFVYWMARDQRVQ 1 2 DNWAFLYAQRLALKQKLPLHVCFCLAPCFLGATLRHYDFMLRGLEEVAE 0 0 ECEKLCIPFHLLLGLPKDVLPPFVQKHSIGGIVTDFSPLVHHTQWVKDVQDALPRQVPFVQ 0 0 VDAHNIVPCWVASDKQEYGARTIRHKIHDRLPHFLTEFPPIICHPYPSSIQAE 0 0 PVDWNACRAGLQVDRSVKEVSWAKPGTAFGLTMLQSFIAERLPYFGSDRNNPNKDVLSNLSPWFHF 1 2 GQVSVQRAILEVQKHRGRYPDSVANFVEEAVVRRELADNFCFYNKNYDKLE 1 2 GAYDWAQTTLRLHAKDKRPHLYSLEQLESGKTHDPLWNAAQ 0 0 MQMVKEGKMHGFLRMYWAKKILEWTHSPEEALEFAIYLNDRFQLDGRDPNGYV 1 2 GCMWSICGIHDQGWAEREIFGKIRYMNYAGCKRKFDVAEFERKYSPVD* 0 >CPD_potTri Potorous tridactylus (rat_kangaroo) D26020 PubMed:7813451 0 MDSKKRSHSTGGEAENMESQESKAKRKPLQKHQFSKSNVVQKEEKDKTEGEEKGAEGLQEVVRQSRLRTAPSVLEFRFNKQRVRLISQDCHLQDQSQAFVYWMSRDQRVQ 1 2 DNWAFLYAQRLALKQKLPLHVCFCLAPCFLGATIRHYDFMLRGLEEVAE 0 0 ECEKLCIPFHLLLGLPKDVLPAFVQTHGIGGIVTDFSPLLHHTQWVKDVQDALPRQVPFVQ 0 0 VDAHNIVPCWVASDKQEYGARTIRHKIHDRLPHFLTEFPPVICHPYTSNVQAE 0 0 PVDWNGCRAGLQVDRSVKEVSWAKPGTASGLTMLQSFIAERLPYFGSDRNNPNKDALSNLSPWFHF 1 2 GQVSVQRAILEVQKHRSRYPDSVTNFVEEAVVRRELADNFCFYNKNYDKLE 1 2 GAYDWAQTTLRLHAKDKRPHLYSLEQLESGKTHDPLWNAAQ 0 0 MQTVKEGKMHGFLRMYWAKKILEWTRSPEEALEFAIYLNDRFQLDGWDPNGYV 1 2 GCMWSICGIHDQGWAEREIFGKIRYMNYAGCKRKFDVAEFERKISPAD* 0 >CPD_ornAna Ornithorhynchus anatinus (platypus) 0 1 2 0 0 ECEKLCIPFHLLLGLPKDVLPAFVQTHGIGGIVTDFSPLLHHTQWVKDVQDALPRQVPFVQ 0 0 VADNWAFLYAQRLALQRKLPLHVCFCLVPSFLGATIRHFGFMLGGLKEVAK 0 0 1 2 GQVSVQRAILEVRKQRHRYPDSVASFVEEAMVRRELADNFCFYNENYDKVE 1 2 DTLWNAAQ 0 0 LQMVQEGKMHGFLRMYWAKKILEWTRSPEEALEFAIHLNDRFQLDGRDPNGYV 1 2 GCMWSICGIHDQGWAEREVFGKIRYMNYAGCKRKFDVAQFEHKYSPRE* 0 >CPD_taeGut Taeniopygia guttata (finch) XM_002190577 0 1 2 DNWAFLYAQRLALKQELPLHVCFCLVPKFLGATIRHYRFMLRGLQEVAE 0 0 ECAELNIPFHLLLGYAKDVLPTFVVEHGVGGLVTDFSPLRLPRQWVEDVKERLPEDVPFAQ 0 0 VDAHNIVPCWVASPKQEYSARTIRGKIHAQLPEFLTEFPPVVPHPHLPSCPAE 0 0 PIAWEACYSSLQVDHTVKEVEWATPGTAAGMAVLKSFIAERLKSFSTHRNDPNKAALSNLSPWLHF 1 2 GQVSTQRAILEVQKQRRNYKDSVDAFVEEAVVRRELAENFCYYNENYDSVQ 1 2 GAYDWAQTTLKLHAKDKRPYLYSLQELEQGTTHDPLWNAAQ 0 0 LQMVQEGKMHGFLRMYWAKKILEWTRSPEEALQFAIYLNDRYELDGRDPNGYV 1 2 GCLWSICGIHDQGWAERPIFGKIRYMNYAGCKRKFDVDQFERRYTPTHSQ* 0 >CPD_melUnd Melopsittacus undulatus (budgerigar) AGAI01046895 0 MLRRQRKRKASAPEEIPRVSRKRTEQEEELHEARRRAAPSVREFKFNKKRVRLISQASDLKDGGRCILYWMSRDQRLQ 1 2 DNWAFLYAQRLALKQELPLCVCFCLVPKFLEATIRHYGFMLRGLQEVAE 0 0 ECAELNIAFHLLRGYAKDVLPAFVEEHGVGGLVTDFSPLRLPWQWVQDVREWLPEDVPFAQ 0 0 VDAHNIVPCWVASPKQEYSAYTIRRKIQAQLPEFLTEFPPVVRHPYSPSCPAE 0 0 PIAWEACYSSLEVDRTVKEVEWATPGTAAGLAVLQSFIRERLKYFGSHRNDPNKAALSNLSPWFHF 1 2 GQVSTQRAILEVQKHQHKYKESVDMFVEEAIVRRELAENFCYYNENYDSVQ 1 2 GAYPWAQTTLKLHAKDERPFLYKLQELEQGTTHDPLWNAAQ 0 0 LQMVREGKMHGFLRMYWAKKILEWTHSPEEALQFAIYLNDRYELDGMDPNGYV 1 2 GCLWSICGIHDHGWTERAVFGKIRYMNYAGCKRKFDVRQFERRYAPCTLSQ* 0 >CPD_galGal Gallus gallus (chicken) XM_422729 0 MPRGNGKGRKERDAGREEEEAVGTLEAAVREARRRTAPSVRDFRYNKQRARLVSRGSELKEGAECILYWMCRDQRVQ 1 2 DNWAFLYAQRLALKQELPLRVCFCLVPAFLDATIRHYGFMLRGLREVAK 0 0 ECAELDIPFHVLLGCPKDVLPSFVVEHGVGGLVTDFCPLRVPRQWVEEVKERLPEDVPFAQ 0 0 VDAHNIVPCWVASPKQEYSARTIRAKIHSQLPEFLTEFPPVIRHPHPPPNPPE 0 0 PIAWDACYSSLQVDRTVTEVAWATPGTAAGLAMLQSFITERLKSFGSQRNDPNKAALSNLSPWFHF 1 2 GQVSTQRAILEVQKHRRVYKESVDAFVEEAVVRRELAENFCYYNENYDSVR 1 2 GAYDWAQSTLKLHAKDKRPFLYKLPQLEQATTHDPLWNAAQ 0 0 LQMVREGKMHGFLRMYWAKKILEWTRSPEEALQFAIYLNDRYELDGMDPNGYV 1 2 GCLWSICGIHDQGWKERDVFGKIRYMNYAGCKRKFDVDQFERRYAHCK* 0 >CPD_melGal Meleagris gallopavo (turkey) XM_003209143 0 MPRGSGKGRKERDAGREKEEAEAGAVGSLEAAVREARSRAAPSVRDFRYNKQRARLVSRGSELKEGAECILYWMCRDQRVQ 1 2 DNWAFLYAQRLALKQELPLRVCFCLVPTFLDATIRHYGFMLRGLREVAE 0 0 ECAELDIPFHLLLGCPKDVLPSFVVEHSVGGLVTDFCPLRVPRQWVEEVKERLPEDVPFAQ 0 0 VDAHNIVPCWVTSPKQEYSARTIRAKIHSKLPEFLTEFPPVIRHPHPPPNPPE 0 0 PIAWDACYSSLQVDHTVTEVAWATPGTAAGLAMLQSFITKRLKSFGSQRNDPNKAALSNLSPWFHF 1 2 GQVSTQRAILEVQKHRCAYKESVDAFVEEAVVRRELAENFCYYNENYDSVR 1 2 GAYDWAQSTLKLHAKDKRPFLYELPQLEQATTHDPLWNAAQ 0 0 LQMVQEGKMHGFLRMYWAKKILEWTRSPEEALQFAIYLNDRYELDGMDPNGYV 1 2 GCLWSICGIHDQGWKERDVFGKIRYMNYAGCKRKFDVDQFESRYTHCK* 0 >CPD_allMis Alligator mississippiensis (alligator) genome/blat 0 MFPKKQRQAAGADTEPSGSRKRGRQDAEVVPGAESLAEAVKQARRRAALSVREFKFNKKRVRLLSQESHLQDGALGILYWMSRDQRVQ 1 2 DNWAFLYAQRLALKQQLPLHVCFCLVPKFLDATIRHFGFLLKGLQEVAE 0 0 ECGELGIPFHLLQGYAKDVLPAFVTRHGIGGVVTDFSPLRVPLQWVEDVKERLPQGVPLVQ 0 0 VDAHNIVPCWVASNKQEYGARTIRPKIHAQLPEFLTEFPPIIQHPFPAAAPAQ 0 0 PIDWDACYAGLEVDRTVPEVKWATPGTAAGLAVLQAFVTRRLPGYSEYRNNPNKAALSNLSPWFHF 1 2 GQVSVQRAVLEVQKSRDRHRESVNSFVEEAVIRRELADNFCFYNKSYDQLE 1 2 GAHDWARDTLILHARDKREYLYDLHQLEQGKTHDQLWNAAQ 0 0 IQMVREGKMHGFLRMYWAKKILEWTRSPEDALRFAIYLNDRFELDGRDPNGYV 1 2 GCMWSICGTHDQGWAERNVFGKIRYMNYAGCKRKFNVGQFEHKYRP* 0 >CPD_chrPic Chrysemys picta (turtle) AHGY01112360 incomplete 0 MSQKSRNRQARGGEEISGDAARPGAAAAPRGQLKGKRKVLAEPGGAEVLTVEKRRKEGEAAEGGGAGSLGEAVRQARLQVAPSVQDFKYNKKRVRLVSQGSDLKDAAQGVVYWMSRDQRVQ 1 2 DNWAFLYAQRLALKQQLPLHVCFCLVPKFLEATIRHFGFMLKGLQEVAE 0 0 ECQELDIPFHLLIGFAKDVLPAFVTGRGLGGVVTDFSPLRVPMQWVEDVKERLPGDVPFVQ 0 0 VDAHNIVPCWVASDKQEYGARTIRRKIHDQLSEFLTEFPPLIKHPYPSTAPAE 0 0 PIDWNACRASLQVDCSVKEVTWATPGTAAGLAVLESFIGERLKSFGTDRNNPNR------------ 1 2 ---------------------SVDGFIEEAVIRRELADNFCYYNRNYDKVE 1 2 GAYDWAKTSLKLHAQDKRSHLYELKQLEEGKTHDPLWNAAQ 0 0 LQMVHEGKMHGFLRMYWAKKILEWTRSPEEALEFTIYLNDRYELDGRDPSGYV 1 2 GCMWSICGIHDQGWAERAVFGKIRYMNYAGCKRKFDVGQFERRY-------* 0 >CPD_anoCar Anolis carolinensis (lizard) XM_003226963 0 MPQKRKKRKEPASGDCENQVEGDGGTSFGSESGDKKTEGKPEEKKPKKTAAVAKGPPLTEAVKASRLKMAPSIAEFHFNKKRVRMISENSDLKEGAQGILYWMSRDQRVQ 1 2 DNWAFLYAQRLALKQNLPLHVAFCLVPKFLEATIRQFGFMLKGLKEVAE 0 0 ECQELSIPFHLLIGFAKDTLPPFVAKHNIAGVVTDFAPLRVPMQWVQEVQERLPPDVPFVQ 0 0 VDAHNVVPCWVASEKQEYGARTIRRKIHDRLCEFLTGFPPVIQHPHQAASQAE 0 0 PIDWDACSASLEVDRSVPEVEWAKPGSAAGLLVLGEFVQERLKFFSSDRNNPNRAALSNLSPWFHF 1 2 GQVSVQRAILEVRKHRTRCRESVEAFIEEALIRRELADNFCFYNPNYDKVE 1 2 GAYEWAKTTLKLHATDKRPHLYTLKQLEEGKTHDQLWNAAQ 0 0 LQMVHEGKMHGFLRMYWAKKILEWTNSPEEALKFSIYLNDRYELDGRDPNGYV 1 2 GCMWSVCGIHDQGWAERAVFGKIRYMNYAGCTKKFDVGQFERKYNPRKIAY* 0 >CPD_pytMol Python molurus (python) 0 YNKKRVRLVSQAPELKEGAQGILYWMSRDQRVQ 1 2 DNWAFLYAQRLALKQRLPLHVCFCLVPKFLDATIRHFGFMLRGLEEVAE 0 0 ECRTLDIPFHLLIGYAKDTLPPFVAQRGIGGVVTDFSPLRLPMKWIQEVLEQLPSDVPFAQ 0 0 VDAHNIVPCWVASDKQEYGARTIRRKIHDQLAEFLTEFPPVIKHPFPVGLQEE 0 0 PISWDACYSSLQVDCSVKEVDWAKPGAASGLKVLEEFVKERLKHFASGRNNPNQAALSNLSPWFRF 1 2 GQVAVQRAILEVYKHRTRYKESVEGFIEEAVVRRELADNFCFYNPNYDKIE 1 2 GAYDWAKTTLKIHAKDKRPHLYTLKQLEEGKTHDPLWNAAQ 0 0 LQMVQDGKMHGFLRMYWAKKILEWTRSPEEALQFSIYLNDRYELDGRDPNGYV 1 2 GCMWSICGIHDQGWAERAVFGKIRYMNYAGCKRKFDVGQFERKYGAHSFTQ* 0 >CPD_xenTro Xenopus tropicalis (frog) NP_001135721 0 MDTKAKSKTITKSPGKRKTPGKVLSETSSGESSDVDAKKKKKMVEQTEEKGVGGAVKISRVEAAPSVSEFKFNKKRVRLvSTEADLKDDAQGIVYWMSRDQRVQ 1 2 DNWAFLYAQRLALKQKLPLHVTFCLVPKFLDATIRHYGFMVKGLQEVAE 0 0 ECKELNIPFHLLIGYAKDILPNFVKKHAIGGVVTDFSPLRVPLQWVEDVSKRLPKDVPLVQ 0 0 VDAHNIVPCWVASNKQEYGARTIRKKIHDQLSQFLTEFPPVIKHPYEPKLEAE 0 0 PVDWDNCYNSLEVDRTIGEVEWAKPGTNAGFNVLQSFISERLRHFNSDRNNPNQNALSNLSPWFHF 1 2 GQLSVQRAILEVQKYRSKCKDSVDSFVEEAVVRRELADNFCFYNKNYDKIE 1 2 GAYDWAKNTLKDHAKDKRTHLYTLEKLEAGKTHDPLWNAAQ 0 0 LQMVHEGKMHGFLRMYWAKKILEWTSSPEEALHFSLYLNDRYELDGRDPNGYV 1 2 GCMWSICGIHDQGWAERAVFGKIRYMNYQGCKRKFDVAQFERRYHPKKFSQ* 0 >CPD_ambMex Ambystoma mexicanum (axolotl) JK977035 fragment 81% allMis EECKRLNIPFHLLIGFAKDVLPGFIKEHAIGGVVTDFSPLRVPMQWVQDVKELLPEDVPF VQVDAHNIVPCWVASVKQEYGARTIRNKIHDKVSEFLTEFPPVLVHPHESKFQGEPIDWD ACIANLQVDRTVGEVDWAKPGTKAGMGVLKSFIAERLKFFGTDRNNPNKNALSNLSPWFH FGQVSVQRAILEVRKYRSRFKESVEGFIEEAMVRRELADNFCFYNKKYDTVEGAYDWAKNT >CPD_lepOcu Lepisosteus oculatus (spotted_gar) AHAT01034265 0 MSGRSPPSPQGGAAGKGAAKRKLQGGEGTGEKKAKKTAPEQGAGWLAEAVAQSRRQAGLSVQGFGFDKKRMRVLSKASAMKGGSEGVLYWMSRDQRVQ 1 2 DNWALLYAQQLALEEQLPLHVCFCLVPCFLDATIRQFGFLLRGLEEVAK 0 0 ECSALGLEFHLLRGSAGEVLPQFVQKQGLGAVVTDFAPLRLPLQWVEEVKERLPQDIPLIQ 0 0 VDAHNVVPCWVASDKQEYSAKTIRKKITSKLPDFLKEFPLVAEHPHRASKPAK 0 0 IIDWEQARSSLAVDRSVKEVSWARPGAAAGLEVLESFIDHRLKDFATQRNNPNGEALSNLSPWLHF 1 2 GQVSAQRVVLEVRRHGRRWPQSVDVFVEEVVVRRELADNFCFYNKKYDRVD 1 2 GAYEWAQKTLREHAKDRRPYLYSRGELEASRTHDKLWNAAQ 0 0 FQMVSEGKMHGFLRMYWAKKILEWTASPEEALATAIYLNDRYELDGRDPNGYV 1 2 GCMWSICGIHDQGWKEREIFGKVRYMNYAGCTRKFNVPKFEIKYNPKRCDDEDGNRKRGGD* 0 >CPD_danRer Danio rerio (zebrafish) NM_201064 0 MSANKNNLKRQMKSTISAGGKQPKLTGEKGKESGWLLKEVAELRRAAQGCEFNKKRLRYLSDTQKVKQSSQGVLYWMSRDQRIQ 1 2 DNWALIYSQQLALAEKLPLHICFCLVPKFLDATYRQYAFMLKGLQEVVK 0 0 ECKSLDIEFHLLSGEPVHNLPAFVKSWNIGAVVTDFNPLRISLQWIDTVKKHLPSDIPFIQ 0 0 VDAHNVVPCWEASTKLEYGARTIRGKITKHLQEFLTDMPLVDTHPYCASRAAK 0 0 IIDWEEVLSSLEVDHNVCEVEWAQPGTTGGMFMLESFIDQRLHIFATHRNNPNSDAVSHLSPWIHA 1 2 GQLSAQRVVMQVKRKKNASESVASFIEEIVVRRELADNFCFYNQNYDNIS 1 2 GAYDWAKKTLQEHAKDRRQYLYTKKELESAETHDQLWNAAQ 0 0 RQLLLEGKMHGFLRMYWAKKILEWTASPEEALSIAIYLNDRLSLDGCDPNGYV 1 2 GCMWSICGIHDQGWAERPIFGKIRYMNYAGCKRKFDVEQFERKYAAIKKNPNLNAKISV* 0 >CPD_petMar Petromyzon marinus (lamprey) rough revised sequence 0 MCLSRLTIVFKLSSWLVNVVGSGNFEEQISKTRAALASSVLEFRFNKKRVRAVSNADDMAANASGILYWMSRDQRVQ 1 2 DNWALLYAQRLALKQRLPLYVCFCLVPKFMDATIRHFGFMLRGLQEVEK 0 0 EFRSLDISFHLLKGYAKDVLPPFVNEKNIGAVVTDFLPLRVPRQWVKDLKKALPKNVPLAQ 0 0 VDAHNVVPCWVTSDKLEYGARTIRRKIHDKLPQFFTEFPPAIKHTYPAKSLPE 0 0 ETDWEAAWQSLEVDRSVGEVAWAQPGTSAGLEMLNSFINERLKLFSTDRNNPTAMHSATSSPVIHF 1 2 GQISVQRCALEVSKLKPKYRESVDMYIEEAIVRRELGDNFCFYNKNYDSLD 1 2 GAYDWAKRSLDAHTQDNREYIYSLQQLEEAKTHDPLWNAAQ 0 0 LQMVVEGKMHGFMRMYWAKKILEWTSSPEEALRIAIYLNDRYQLDGSDPNGYV 1 2 GCMWSICGTHDQGWAERAVFGKIRYMNYAGCKRKFDVGAYESRYSSKLGF* 0 >CPD_braFlo Branchiostoma floridae (amphioxus) XP_002586934 FE570347 fixed frameshift exon 4 0 MSSGDKRKSSGSGDSPAKKAKVETGRSSAQDGAGSEDFLDQIAERRQNVCESVAGFKFNKKRCRQMSDVGDLKQNCGGIMYWMSRDQRVQ 1 2 DNWALLFAQRLAMKHKVPLYVCFCLVPKFLDANIRHYGFMLKGLEEVEK 0 0 ELTDLDVSFHLLTGYAVGFLPGFVDEKNIGGVVTDFSPLRTPLKWVHDLAGKLPSDVPFVQ 0 0 VDAHNIVPCWEASPKQEYGARTIRGKIHNQLSSYLTEFPPVSQHPHPPKDKPE 0 0 PTDWAAARASLKVDMTVAEVTWATPGTTGGLRMLESFCKQRLKNFGTDRNNPNKRALSNLSPWIHF 1 2 GQISVQRCILTVKKYRSKYKESVDAYIEEAIIRRELSDNFCFYNHDKYDSIK 1 2 GASAWAQKTLKDHAGDKREYLYSREQLEQAKTHDDLWNAAQ 0 0 DKYSLDGRDPNGYV 1 2 GCMWSICGIHDQGWAERPVFGKIRFMNYNGCKRKFDVAQFVTKYKAKGYLKK* 0 >CPD_strPur Strongylocentrotus purpuratus (urchin) JT122393 JT102939 FJ812411 0 MITFSVQGLKLGHHCQCWKVVFLSRFISSRSYSSIMGKDGQTAKRSSEGQDNGETSAAKKVKKEEDGAEVQDLQTKVASLRNASGSSILNFKFNKKRVKILSDTMDIAEDNRGIVYWMSRDQRVQ 1 2 DNWALLFAQRLAMKQEVPLHVCFCLVPKFLEGTIRHFNFVLEGLKEVSQ 0 0 ECKELNIPFHLLIGYAKDILPNFVKKHAIGGVVTDFSPLRVPLQWVEDVSKRLPKDVPLVQ 0 0 VDAHNVVPCWEASNKLEYGARTIRPKITKQLSTYLTEFPPVICHPQKAKAKAE 0 0 PIDWEGAYASLEVDQTVKPVDWAKPGTSEGMKMLDSFVKERLRYFSSARNDPTKSVCSNLSPWIHF 1 2 GQLSSQRAALIVRLYRSRFSESVAAFIEESIIRRELSDNFCFYNDKYDSIE 1 2 GTNDWAKKTLKDHAKDKRDYVYSRETLERAKTYDQLWNSAQ 0 0 KQMVREGKMHGFLRMYWAKKILEWTTSPEEALEIAIYLNDRYSLDGRDPNGYV 1 2 GCMWSICGIHDQGWGERPVFGKIRFMNYKGCKRKFDVDTFVNRYKKYNL* 0 >CPD_aplCal Aplysia californica (sea_hare) scaffold_446:238,174 0 MESDSPSKSEAGGDTGDFIERINKQRTSVCASVDKFKFNKNRVRVLSKEEDFPDDSDGVVYWMSRDQRVQ 1 2 DNWAFLYAQRLALKLEAPLHVCFCLVPKFLEATIRQFGFLLKGLAQVEK 0 0 ECRELNIAFHLLLGEASKVLPQFVKDNSIGGVVTDFSPLRTPAKWVDELKEALPEDVPLCQ 0 0 VDAHNIVPCWLASPKLEYGARTIRNKIHNQLGSFLTEFPPVSRHPHPPSKMPP 0 0 VTDWKAADGYLQVDRSVAEVTWAKPGTDPGLRMLESFCEKRLKSFATSRNDPNKQALSNLSPWIHF 1 2 GQISVQRCILMVKKYRSKSSEGVNAFIEEAVIRRELADNFCFYNEKYDSVK 1 2 GAYDWAQKTLKIHSGDKRPYLYDREKLEQGKTHDDLWNAAQ 0 0 IQMVTEGKMHGFLRMYWAKKILEWTPSPAVALTEAIYLNDKYNLDGRDPNGYV 1 2 GCMWSICGIHDQGWAERDVFGKIRYMNYKGCQRKFDVANFVRKYGKKKA* 0 >CPD_vilLie Villosa lienosa (mussel) JR505029 transcript assembly mollusc 0 MADGKRSKRKGVDEPYAEKKKLKGNEAQDTEQDFLSKIKYARLNVCKSVEEFKYNKKRVRIVSRCEEFPDACDGIVYWMARDQRVQ 1 2 DNWAFLYAQKLALKMQVPLHVCFCLVPKFLDANIRHYGFMLKGLQEVEE 0 0 ECRSLGISFNLLIGYAKDVLPDYVKKNKIGGVVTDFSPLKLPLQWVKDVGDALPKSVPFCQ 0 0 VDAHNIVPCWEASPKLEYSARTIRNKIHDKLSQFLTEFPPVIYHPHTAKSKSE 0 0 PLDWKAAEASLQVDRSISEVTWAKPGTIGGLLMLESFCKERLKYFASDRNNPNKNALSSLSPWIHF 1 2 GQISVQRSILMVRLYRGRFKDGVEAFIEEAVVRRELADNFCYYNENYDKVE 1 2 GAYDWAQKTLKDHEKDKRPYLYTKEQLEKARTHDDLWNSAQ 0 0 IQMVSEGKMHGFLRMYWAKKILEWTKTPQEALEISIYLNDKYNLDGRDPNGYV 1 2 GCMWSICGIHDQGWAERDVFGKIRYMNYAGCKRKFDVSLFERKYKNKNSDLMEKHKEG* 0 >CPD_droMel Drosophila melanogaster (fruitfly) thymidine dimer photolyase CG11205 uses 5-deazariboflavin 0 MFTLASYWRESFKI 0 0 VLPLQAMKRTKAQKAGPSKKAAKNEKASSEPKSDQESSDEEASTSKALLVSKPDYQNFEQFLTHLEHQRVCTAANIQEFSFRKKRVRVLSKTEDVKESSL GGVVYWMSRDGRVQDNWALLFAQRLALKLELPLTVVFCLVPKFLNATIRHYKFMMGGLQEVEQQCRALDIPFHLLMGSAVEKLPQFVKSKDIGAVVCDFA PLRLPRQWVEDVGKALPKSVPLVQVDAHNVVPLWVASDKQEYAARTIRNKINSKLGEYLSEFPPVVRHPHGTGCKNVNTVDWSAAYASLQCDMEVDEVQW AKPGYKAACQQLYEFCSRRLRHFNDKRNDPTADALSGLSPWLHFGHISAQRCALEVQRFRGQHKASADAFCEEAIVRRELADNFCFYNEHYDSLKGLSSWAYQTLDAHRKDKRDPCYSLEELEK 0 0 SLTYDDLWNSAQLQLVREGKMHGFLRMYWAKKILEWTATPEHALEYAILLNDKYSLDGRDPNGYVGCMWSIGGVHDMGWKERAIFGKVRYMNYQGCRRKFDVNAFVMRYGGKVHKKK* 0 >CPD_nasVit Nasonia vitripennis (wasp) XM_001603235 trimmed N-terminal LVKQIEEQRNQTADSVMTFKFNKKRVRILTDSDEVSSESKGIVYW MFRDARVHDNWAMLFAQKIALKNKVPLHVCFCILPKFLDATIRHYKFLLEALEEVEKDCKELNINFHLLHGEPNTAIINFVEKYKMGAVIADFFPLRLPLFWLEDIKKKLPKKIPL CQVDAHNIVPCWVASEKLEYAARTIRNKINSKLDEFLTEFPPVIKHPHTSDQKFDKINWDTALDDVLVDKSVDKITWAKAGYKGGIAELDKFLKIRLRIYDEKRNNPIFNALSNLS PWFHFGMISVQRCILEAKKYKSQYNKSVEAFMEEAIVRRELSDNFCFYNEHYDSLKGAYDWARETLNQHRNDKRDYIYTLNELENGLTHDDLWNAAEIQLVKEGKIHGFLRMYWAK KILEWTETPDDALKWSIYLNDKYSMDGRDPNGYVGCMWSICGVHDQGWRERPVFGKIRYMNYKGCERKFDVQAFVMKYGAKVQLTKDENKRKGKKK* 0 >CPD_bomImp Bombus impatiens (bumble_bee) XM_003488984 MDELNPSKRKKVSESIMTFNFNKQRIKHLSELNDVKENCKGILYWMFRDIRTQDNWALLFAQKIAVKRNVPLHICFCIMPSFLNASMRYYKFLLKGLMEIEE ECKQLNLNFHLLHGEPNESILKFVKTYNMGAVIVDFYPLKLPMLWIDNVQKNLPEDIPIYQIDAHNIVPCWYASSKQEFAAKTIRNKINTKLQEFLTEFPPVIKHPYLTKEKFENN NWDITLQDVEASKPTTEITWAKPGYRNGIKELENFIQNHLQKYGDERNNPLSNVISNLSPWFHFGMISVQRCILEIQEYKGLYKKSVESFMEEAIIRRELSDNFCFYNEKYDLVEG AYPWAIKTLNKHRKDTRKYVYSLSQLENSKTHDDLWNACQNQMIITGKMHGFLRMYWAKKLLEWTEIPEIALEWANYLNNKYSIDGCDPNGYVGCMWSICGVHDHGWPERDIFGKI RYMNYNGCKRKFNVAEFVMKWGKKKASELI* 0 >CPD_apiMel Apis mellifera (bee) XM_003250426 MEELNPFKRRKTSESIMTFKFNKKRIKRLNNLNDIKENCNGILYWMFRDIRIQDNWALLFAQKAALKNNVPLHICFCLISNFLNASIRYYKFLLKGLEEIEK ECKKLNINFHLLHGEPNINILKFIKIYNMGAIITDFYPLKLPMLWIDNVKKNLPKDIPICQVDAHNIVPCWYASSKQEFAAKTIRNKINTKLEEFLTEFPPVIKHPYTTKEKFEKN NWKIALQNVEVDKSVKEITWAKPGYENGIKELENFLQNRLKKYGDERNNPLSNAISNLSPWFHFGMISVQRCILEIKEYKKLYKKSVESFMEEAIIRKELSDNFCFYNEKYDLIEG AYPWAIETLNKHRKDKRKYIYFLNHLENSETHDDLWNACQNQMVTIGKMHGFLRMYWAKKILEWTETPEIALKWANYLNNKYSIDGCDPNGYVGCMWSICGVHDHGWPERDIFGKI RYMNYEGCKRKFNIAEFVMKWGKKKTNELI* 0 >CPD_anoGam Anopheles gambiae (mosquito) XM_313925 trimmed N-terminal YVAMLKADRKATAKSILDFDFNKKRVRILSDA KTIEDGKQGVLYWMSRDARVQDNWAFLFAQKLALKNDLPLHVCFNLVPKFLDATIRHFKFMLKGLEEVAEECRRLNIQFHLLRGSAGENVPAFVKKHKIGGVVCDFSPLRVPMKWV DEVRKALPMEIPLCQVDAHNIVPVWVTSDKLEYAARTIRTKVNKNLPTYLTPFPPLVKHPFTANFEADPINWTQVLDTLQVDRTVEAVEWATPGYAGGVKTLQTFVEKRLGKFNDK RNDPTENALSNLSPWFHFGQIAVQRAVLTVKKHGKRYSESVASFCEEAIVRRELSDNFCFHNKNYDNLQGAYDWARKTLDDHRKDKRVYCYSREELETAKTHDDLWNSAQLQMVKE GKMHGFLRMYWAKKILEWTKTPEEALETAIYLNDRYSLDGRDPNGYVGCMWSIAGIHDQGWKEREIFGKIRYMNYDGCKRKFNVNAFVVRYGGKVHRRK* 0 >CPD_aedAeg Aedes aegypti (mosquito) XM_001653905 trimmed N-terminal FVEKFKTIRKETAKSILDFDFKKKRVRILSDAKEVEENKKGVIYWMSRDARVQDNWAFLFAQKLALKNELPLHV CFSLVPKFLDATIRHYKFMLKGLEEVAKECESLNINFHMLTGMAKDTIPKFVKTHNIGAVVCDFSPLRVPMKWVEDVRKSLPAEVPLCQVDAHNIVPLWVTSEKQEYAARTIRNKV NNNLNTYLTQFPPVIKHPHKASFKADSIDWVKLLDTIEVDRTVDEVDWAVPGYTGGIGVLQSFVEKRLRKFNAKRNDPTDDALSNLSPWFHFGQISVQRCVLAVKKYGKGYSEGVA AFCEESIVRRELSDNFCYYNKNYDNLKGAYDWAQKTLDDHRKDKRTYVYSRDQLEQARTHDDLWNSAQIQMVKEGKMHGFLRMYWAKKILEWTKTPEEALETAIYLNDRYQLDGRD PNGYVGCMWSIAGIHDQGWREREVFGKIRYMNYEGCKRKFDVAAFVARYGGKVYKK* 0 >CPD_acyPis Acyrthosiphon pisum (aphid) XM_001949116 trimmed N-terminal FLNDVAAERNKTASSIMEFKFNKKRVRVLSEQKEVPEWAEGVIYWTFRDERIHDNWALLYAQKLAIKNKVSLHITFC RLKQFLDCSLRHYKHIFQGLEELETECKSLDIQFHFLIGCAADILPDFVKKHKLGAIIVDFMPVREHISWTKQLADRIGSEVPVIQVDAHNIVPCWVASDKQEYSARTIRNKINNK LPEFLTEFPPVIKHPFRSTFKAKPTNWDEADKTLEVDRSVVSVPGLKAGFKAGMSELEQFLKKRLPKYSTDRNNPVKDGLSKLSPWLHFGQISAQRCILEVSKLSKQYPDSVAAYR EEAIVRRELSDNFCFYNPKYDKIEGAPNWAQTSLNEHRKDKRMFVYTREELESSRTHDDLWNSAQIQLVKEGKMHGFLRMYWAKKILEWTDTPDRALADAIYLNDKYSMDGRDPSG FVGCMWSICGIHDQGWKEREIFGKIRYMNYAGCKRKFDINAFIARYGGMVHKYTKK* 0 >CPD_nemVec Nematostella vectensis (anemone) ABAV01006764 XM_001636204 bad BACK01030119 0 MASDEPAAKRRKQEVTGPNKSELDQLDDIVKTYKEKRMSVCNSVTDFKFNKKRVRMLSKEGSISENQCGGVVYWMWRDQRVQ 1 2 DNWALLYAQRLALKQQAPLHVCFCLPSNFLDATLRQYNFMIKGLQEVEK 0 0 ELKELEISFHLLLGDPGKVLPEFVKSAGIGGIVVDFCPLRLPTQWVNDVVKAVPKDVPVCQ 0 0 VDAHNIVPCWHASPKLEYGARTIRPKIHKVLTEFLTEFPPVIKHSHVSGEKTK 0 0 TTDWDAVDTFIEVDRSVGEVDWAKPGTAEGLFMLESFCKDRLKYFHSSRNDPTKRALSNLSPWFHT 1 2 GQISPQRAILRVRDFRSKFRESVESFIEECIIRRELSDNFCYYNNEKYDSI 1 2 EGTNEWARKTLNDHAKDKREYLYARGKLEKAQTHDDLWNAAQ 0 0 RQLVREGKLHGFLRMYWAKKILEWTDSPETALSDAIYFNDKYALDGIDPNGYV 1 2 GCMWSVCGIHDQGWAERPVFGKIRYMNYKGCQRKFAVAEFVKRYKP* 0 >CPD_acrDig Acropora digitifera (coral) BACK01030119 cnidarian one intron missing 0 MASQPPAKRRKEESSENGDDILASCMAKRAQVCASVADFKFNKKRVRLLSKSTDISDDCKAIAYWMWRDQRVQ 1 2 DNWALLYAQRMALKQEVPLVVCFCLPSKYLHSTFRQYSFMIKGLQQVEK 0 0 ELASLN-INFHMLLGEPNIVLPDFVKSENIGGIIADFTPLRKPMKWLNDVMDKLPKNVPVCQ 0 0 VDSHNIIPCWQASPKLEYGARTIRPKIHNQMREFLTEFPPVIKHPHAGKASIK 0 0 TDWKAADEFIEVDRNVQEVNWAEPGTKAGLEMLESFCKDRLKFFATDRNNPTKEALSDLSPWFHA 1 2 GQISVQRAILRVREFRGSSKDSVETFIEEAVVRRELADNFCFYNQEHYDSI KGAKDWAQKTLNDHAADKREYLYTREQLEKGQSHDDLWNAAQ 0 0 LQLVNEGKMHGFLRMYWAKKILEWTESAEEGLEISLYLNDKYSLDGTDPNGYV 1 2 GCMWSVCGIHDQGWAERPVFGKIRYMNYNGCKRKFDVGEFVRKYGATTSKEADGPAKKRKKAK* 0 >CPD_ampQue Amphimedon queenslandica (sponge) ACUQ01006132 XM_003388698 bad 0 MQCKQKNKAWSSKLTATPILCKKIKMAAGSQAAGAARGAALQESLLIAGSSFNMKRCRLITKPTAGKSSIVKGPVLYWMSRDQRVQ 1 2 DNWGLVYSQELANKHGVPLLVAFTLVPKFLDATWRQYSFMMSGLQEVEK 0 0 ELLKLKIPFHLLLGKAQSCLPSFIAKESVSVVVCDFSPLRVPLGWVKETGAELDKIKVPLVQ 0 0 VDAHNIVPVWLASDKQEYAARTLRNKIHKFLPEFLTEFPLVTLHTHNSKLTMK 0 0 STNWIKARESLEVDMTVSEVSWATPGTNAGLKVLDDFCTKRLKFFAAQRNDPNKDSLSNLSPWYHF 1 2 GQVGVQRAILKVKSYSSKHSESVSAYIEEAVVRRELADNFCYYNPHYDSI 1 2 YHYCDLFFFFSRAHKKDKREYIYTQEQFESSSTHDPLWNAAQ 0 0 LQMVQEGKMHGFLRMYWAKKILEWTSSPEEGLRIAIYLNDKYEIDGRDPNGYV 1 2 GCMWSICGIHDQGWAERSVFGKIRYMNYQGCKRKFDVGQFERKYNKKRKLGEQ* 0 >CPD_monBre Monosiga brevicollis (choanflagellate) ABFJ01000652 related intronation but numerous differences 0 0 0 LYWMSRDQRVQ DNWAFLYAQRLALKQRLPLHVCFCLVPKFLDATIRHFGFMLRGLEQ 0 0 VETHLRKKHIPFHLLTGYAKDVLPKFAEEQEACAVVCDMSPLRVPMAWVKETGSKLKDMNVPLYQ 0 0 VDAHNIVPVWHASPKQEYAARTIRNKIHQKLDTCLQPFPELESNSNSVQLPDT VDWKKARESLEINWDVKEVDWLKPGYEGGMKMLEEFINERLHRYADDRNDPNLDALSNLSPYYHFGQ 0 0 ISVQRVVLELRSKQRGKYAEGVKAYIEEAVVRRELSDNFCFYNHRYDSVE GASAWAQETLDVHSKDKR 2 1 EHLYTRKQLENAETADDLWNASQLQLVQEGKMHGFLRMYWAKKILEWTESPEKALEDAIYLNDKYELDGRDPN 1 2 GYVGCMWSICGIHDQGWGERPVFGKIR 21 YMNYKGCKRKFDIAAFVKRYPPAAKNAARGKDGR* 0 >CPD_salSpp Salpingoeca species (choanflagellate) ACSY01000967 different intronation still 0 VYWMSRDQRAK DNWALLYARSLARSARVPLVVVFSLVPKFLDATIRHYGFMLRGLHQ TAKHLHEKLVPFHLLQGSAATTVPAFAAQHEAAAVICDMSPLRVPLRWVKDVGQALEAQNVPLLQ VDAHNIVPVWVTSQKQEYAARTIRPKIHKHLDTYLQPFPELDANDKDTLGDMELPPVFDLEAQFDMLEVDTSVKE 0 0 VDWIEPGYEQGMAAAEAFGRDRAKKFDELRNNPNEDVCSNLSPYFHFGQ ISAAAVVLLLKSKYSKKAAKGVQTFIEEAVVRRE 0 0 LSDNFCFYNRRYDSIDGAAVWARDTLDTHRHDKR 2 1 EYVYTREQLEQGKTHDDLWNAAQLQ 0 0 MVERGKMHGFLRMYWAKKILEWTATPEDALQTALFLNDRYELDGRDPN 1 2 GYVGCMWSIAGIHDQGWAERAVFGKIR 21 YMNYKGCKRKFDVARFVSRFPQAVANAA* 0 >CPD_araTha Arabidopsis thaliana (cress) NM_179320 AFMZ01000529 PHR1 GC-AG splice exon 6-7 0 MASTVSVQPGRIRILKKGSWQPLDQTVGPVVYWMFRDQRLKDNWALIHAVDLANRTNAPVAVVFNLFDQFLDAKARQLGFMLKGLRQLHHQIDSLQIPFFLLQ 0 0 GDAKETIPNFLTECGASHLVTDFSPLREIRRCKDEVVKRTSDSLAIHEVDAHNVVPMWAASSKLEYSARTIRGKINKLLPDYLIEFPKLEPPKKKWTGMMDKKLVDWDSLIDKVVR 2 1 EGAEVPEIEWCVPGEDAGIEVLMGNKDGFLTKRLKNYSTDRNNPIKPKALSGLSPYLHFGQVSAQRCALEARKVRSTSPQ 0 0 AVDTFLEELIVRRELSDNFCYYQPHYDSLKGAWEWARKSLMDHASDKREHIYS 2 1 LEQLEKGLTADP 0 0 LWNASQLEMVYQGKMHGFMR 2 1 MYWAKKILEWTKGPEEALSISIYLNNK 0 0 YEIDGRDPSGYVGCMWSICGVHDQ 0 0 GWKERPVFGKIRYMNYAGCKRKFNVDSYISYVKSLVSVTKKKRKAEEQLTRDSVDPKITIV* 0 >CPD_orySat Oryza sativa (rice) B096003 BACJ01049170 aka:PhrII,Class II PMID:22170053 PDB:3UMV 0 MPPTSVSPPRTAPGPANPSPAHPSRVRVIHPGGGKPGGPVVYWMLRDQRLADNWALLHAAGLAAASASPLAVAFALFPRPFLLSARRRQLGFLLRGLRRLAADAAARHLPFFLFTG 0 0 GPAEIPALVQRLGASTLVADFSPLRPVREALDAVVGDLRREAPGVAVHQ 0 0 VDAHNVVPVWTASAKMEYSAKTFRGKVSKVMDEYLVEFPELPAVVPWDREQPEGVDWDALIARVCS 2 1 EAENVPEIDWCEPGEEAAIEALLGSKDGFLTKRIKSYETDRNDPTKPRALSGLSPYLHFGHISAQRCALEAKKCRHLSPK 0 0 SVDAFLEELVVRRELADNFCYYQPQYDSLSGAWEWARKTLMDHAADKREHIYT 2 1 REQLENAKTHDP 0 0 LWNASQLEMVHHGKMHGFMR 2 1 MYWAKKILEWTSGPEEALSTAIYLNDK 0 0 YEIDGRDPSGYVGCMWSICGLHDQ 0 0 GWKERPVFGKIRYMNYAGCKRKFDVDAYISYVKRLAGQSKKRNAEESPNPVVKLSKSQH* 0
Cryptochromes from cnidarians, ctenophore, trichoplax, sponge, choanoflagellate and diatom
This collection of cryptochromes and photolyases serves as deeper outgroups to bilateran sequences. Some do not classify clearly; others represent clade-specific gene family expansions of litle general interest. Those with minimal or novel introns may represent retroprocessed genes that were subsequently re-intronated at new locations. Those clearly orthologous to DASH and CPD have been moved to those sections. As the remaining sequences are classified, they will be moved to their corresponding orthologous class above to the extent that is useful.
The Arabidopsis cryptochromes/photolyases have been re-named according to contemporary classification but with common synonyms provided in their fasta header. To collect all the available sequences for this or any other species, simply use web browser search (eg '_araTha') or sort the summary page of fasta headers by species.
The phylogenetic position of ctenophores is uncertain (sister to cnidarians vs earlier branching). The genome assembly of Mnemiopsis contains only CRY64 and an ancient duplicated pair of CPD cryptochromes. While these bear no special relationship to cnidarian counterparts (clustering instead with invertebrate representatives), that might be attributable to rapid evolution of cnidarians rather than a non-sistered topology. Use of maximal likelihood cannot resolve this because it requires modeling evolutionary processes over a vast span (600 myrs of geologic time but tens of billions of branch length time) and so the result would be no more reliable than its untestable assumptions.
>CRY64_mneLei Mnemiopsis leidyi (sea_walnut) AGCP01007162 ctenophore no introns 49% identity to closest refSeq CPD MGKIALHWFRHGLRLHDNTPLQKAIAGCTSLIPLYILDTEYFKPGKVGINRMGFLLDSLKALDTDLRAIGSRLYVAKGDPEEVIGKFIKEHKIGAVSFER DTEPYNKVMDGKIINLTKELNVEAFPLWGHTMFDPEYLLALNNGDAPLTMTSFLRLMSEAGDPPKPIDPPKSLPPPPADSLVCKDVFIFQGVPTLSDLTE YEFNPKDYTTWFVAGEKEGIRVMNEFLAQKRRVSTFEKPKTDPTALQPDTTALSPYITRGSLSSRTFYHGLKDTLKGMKSSKPPVSLKGQLYWREMAYLI GFSVPNFNQMEGNPICKQIPWLTGDDAKALLDKWEMGQTGFPAVDAVMNQLRTEGWMHHLARHLVACFLTRGDLWVTWELGRDVFEKHLVDADWSINNFS WHWLSCSAFFHQYFRCYSPIAFFKKTDPNGNYIKKHVPILAKFPDKYIYEPWTAPKGIQIACGCIIGKDYPKPMVEHQFVCQENKSRMKKAYDTSKKRDS GEPPAKKLKQNNLSKYRTK* >CPD1_mneLei Mnemiopsis leidyi (sea_walnut) AGCP01013608 ctenophore no introns 54% identity to closest refSeq CPD MASDKEPAPKRQKTEEIKFNMSRLKQLHGGQGAPSNVTAVAYYMHRDQRVQDNWALIYAQSLATRHSAPLHVVTLITTSHPEQRGATFRHLQFCFDGLKE VSEELSELNIAFHLLIDQGGKVGGGKVAKWMKECEVDCLVTDFSPLRQHRSLLAQLTKSTDLPDSATVYQVDAHNIVPVEVTSDKQEYAARTIRNKIMSK LGKYLTEFPPVTKHPFGDAAMATRHFAEETGTSAKLKEDWDSVLQNLKLDFSVKPSELYKGGTKAGMSCLEDFVERRIKRYSDKRNDPTEDAISDLSPWL HMGNLSAQRAVLYVKKHASSYSSVFIEEAVVRRELSDNFCFYNQNYDSIKGAANWAQETLKAHRDDKREYVYTRQQFLEAKTHDNLWNAAQKQLVREGKM HGFMRMYWAKKILEWTASPEEALAEALYFNDHFSLDGNDANGFVGCMWSICGVHDQGWRERSVFGKIRYMNYNGCKRKFDIEKYIAKYS* >CPD2_mneLei Mnemiopsis leidyi (sea_walnut) AGCP01006228 ctenophore no introns 49% identity to closest refSeq CPD MNYLSFNVKRCRLLGGNDNIISKSSGIAYYMHRDQRVQDNWAFVYSQDLAVKHNLPLYVLAGFNVKHPENPEGTRRAIDFTIGGLKEVEKECTELGIQFH MFKDHHVPMFERILKFIKETSVRCIVADFSPLHPHRNQMNELAKRLDSSSCLLQVDAHNIVPVWEASKKDEPRATEMRNQIMPQFDEYFTEFPRIKNHPT KVLDPLPAVNWEEIRNSVTVDETVQSCEWAKPGTSHGMQHLKRFLTEGLHIYNEKRNIPTVDAISNLSPWFHSGQVSAQRAVIEVKKFEEQYPKSVYKFV DEAVVWSEMSDNFCFYNDNYDKLSGAPQWAQDTLNEHRSDKREYVYTKEQFEKAQTHDNLWNAAQIQLRDEGKMHGFMRMYWAKKILEWSENPDEAIAIA LYLNDHYSLDGADSNGFAGVMWSIAGVHDPPKWGERPVYGKIRYMSYNGCKGKFDIFEYIKKYQSGDAASAPQGGLHKYYKGADPKQKGQGSGRVQGSWG GGKQ* >CRY1A_acrMil Acropora millepora (coral) EF202589 MSLNLKSVEDNNSAVSAEKSQGKLKAKHAIHWVRKDLRLHDNPSLLEAVKGSDTVRIIYVLDTKVDHATGIGLNLWRFLLQSLEDVDDSLRKLNSRLFVV RGQPADVFPRLFREWKTSFLTFEEDSEPFGREKDAAIRLLAQESGVEVAVGRSHTLYDPQLIIKHNSGTAPLTYKKFLAIVRSLGNPQHPCATLDVHLLG GCSTPVSEDHEEKFGVPSLKELGLDVAKLSTEIWHGGETEALIRLDRHLERKAWIASFEKPKVTPNSLFPSPTGLSPYLRFGCLSPRLFYHRLSELYRKV KCKDPPISLYGQLLWREFFFTVAANNPNFDQMDENPVCLQIPWTANPEWLKKWEQGQTGFPWIDAIMIQLKQEGXIHHLARHAVGCFLTRGDLWISWEEG MKVFERWLLDAEWSLNAGNWMWLSCSAFFQQFFNCICPVGFGRKLDPNGDYVRKYLPVLKGFPAKYIHAPWTAPENVQRAARCIIGKDYPRPIVDHHKVS TANLEKMRNVFKALLRYKESTVVATSEKSDGSKDKQAKLKQQMLNLEDKENR* 0 >CRY1B_acrMil Acropora millepora (coral) EF202590 ICVQRLKDLCFSLDILGHTSLHWFRKDLRLHDNPSLRECLRNSKVFYGVYFLPPSEAKQGSVSLNRWGFLLESLRDLDTSLVECGSRLFVIRGNPVEMLP NLFKKWNINQMSFEVDSEPYSNSRDLVISHLAKENGIEVISRVSHTLYDPRILRGLSSGIVPLLFDEFKESVLQKRQPEKPVPKVGRKLFGACVTPVGSD HQQCYGVPRLDEIGGKFSKSGSVCSELYKGGESEALKRMETALQWMVKNDFTEPEITIHSLLXSATHLSPYLRCGCLSPRLLHQRITEEYIKSKGVSPPS ELFVKLLWREFFFVVGGQIPDAHEMINNPISLEIPWEDSVEYLERWKKGTTGFPWIDAIMRQLRTEGWIHNIARRAVASFLTRGCLWVNWEEGFKVFDEF QLDAERSLNVGNWLWVSTSTFVKGPVPWFCPVGVGKKIDPTGEYVRKYVPEIRNLPTEYIFEPWLTPRDLQRSYGCVIGRDYPAPIVNHVKQRAICVQRM QELAFKLASKIRWRQVLNVI* 0 >CRY1A_nemVec Nematostella vectensis (anemone) XM_001623096 MSGMDKFTTDSCNLVKAKKTSHSIHWLRKGLRIHDNPALRDAVLNWGTFRVVYILDTKSVASSNIGLNLWRFLLQALEDLDDSLRKLNSRLFVIRGQPAD VFPRLFREWGITRLTFEEDSEPFGKERDSAICMLAREAGVEVASHRSHTLYHLQGIIDRNGGTPPLTYKKFLSVIEGIAPPDPPVPHIDPSAQELGHTPL SDNHDELYGVPTMEELGLETSKLFVEVWHGGETEALKRLDRHLERKAWIASFGKPKVTSDSLMASPSGVSPYLRFGCLSPRLFYYRLMDLYRKVKGGPPP TSLYGQLLWREFFFVVSTNNPNFDKMESNPICLYIKWRTDKQVASDLQKWTDAQTGFPWIDAIMSQLKQEGWIHYLARVAIASFLTRGCLWISWEAGMKV FAEHLLDADWSSNAGNWMWWSYSAFSQQFFNPPCPVEFSKSLDASGDYIRKYLPVLKGFPARYIHEPWKAPLNVQKMAKCIIGKSYPSPMVDHEKAVSKN MEEMTSVYKSVIQYRISVLSSEEKQIEETKSQFQMPETSMH* 0 >CRY1B_nemVec Nematostella vectensis (anemone) XM_001623096 MSGMDKFTTDSCNLVKAKKTSHSIHWLRKGLRIHDNPALRDAVLNWGTFRVVYILDTKSVASSNIGLNLWRFLLQALEDLDDSLRKLNSRLFVIRGQPAD VFPRLFREWGITRLTFEEDSEPFGKERDSAICMLAREAGVEVASHRSHTLYHLQGIIDRNGGTPPLTYKKFLSVIEGIAPPDPPVPHIDPSAQELGHTPL SDNHDELYGVPTMEELGLETSKLFVEVWHGGETEALKRLDRHLERKAWIASFGKPKVTSDSLMASPSGVSPYLRFGCLSPRLFYYRLMDLYRKVKGGPPP TSLYGQLLWREFFFVVSTNNPNFDKMESNPICLYIKWRTDKQVASDLQKWTDAQTGFPWIDAIMSQLKQEGWIHYLARVAIASFLTRGCLWISWEAGMKV FAEHLLDADWSSNAGNWMWWSYSAFSQQFFNPPCPVEFSKSLDASGDYIRKYLPVLKGFPARYIHEPWKAPLNVQKMAKCIIGKSYPSPMVDHEKAVSKN MEEMTSVYKSVIQYRISVLSSEEKQIEETKSQFQMPETSMH* 0 >CRY1C_nemVec Nematostella vectensis (anemone) XM_001630979 MVNSVHWFRKGLRLHDNEPLRRAIEGSDTFRGIFFLDKAAVKNAKVSPNGWRFLIESLRDLDQSLQKYNSRLFIIQGQPIDVFPKLIKQWNISKLTFEYD SEPFPRQRDLAVKRIAEKAGVDVIVCSSHTLYDIECIEQILEMNEGKPPVLFKDFEAIVRKLGPPENPVESITRKSFMNCICPITSTHSEKFGVPSLAEL GIKPADVTCGDMWVGGEREALNRLGILEDKILGSNSSKNCDGSSVLFPSRTQLSPYLRFGCLSPKLYYKKLTGAYVKMQFSPPPMSLYRQLIWREFFFTL ASKNPNMDQVNDNPLCIQISWEENKDHFLCWKQGNTGFPWIDAIMRQLQKEGWIHHLARLSVGCFLTRGCLWISWEEGLRAFEELQLDAEWSMNSGSWLW LSCSAFFHGQIPWFCPVEVGKKLDPSGSYIRKYVPELRSMPLKYLYEPWTAPDDIQRAAMCVIGEDYPHPIVDHIEQRKACLQRLKTVCRHIQTTSFA* 0 >CRY1D_nemVec Nematostella vectensis (anemone) XM_001632799 MKERILFSSDCPSLGISSMHWFRKDLRLHDNPALLESFKNCQAFYGVYFLDPASVQRSNLSPNRWWFLLESLRDLDYNLRSLGSRLLVVRGQPVQEMPKL LDQWNIKRLTLEYDSEPPAKQRDAVVTHLAKNLGVEVIQRVSHTLYDVETVLETNDGKLPMTFDEMAKTAEQLGPPCPPCQTVDKTVFGACLTPVGPDHA DKYGVPLLSEFGMKELKEATAKKYWTGGEPEALRRLSAALKKCAENDFEERGWTIDEMFSNDAHLSPYMRFGCLSPRLYYQQLALTYMKEKKSIPPATLF TGLVRRELFLHVASHNADLDKMLDNPLSVQFPWEENKEGLERWKEGKTGFPWIDAIMRQLREEGWIHHLARQAVGCFLTRGCLWVSWEEGFKAFDELQLD AEWSLNASNWLWLSCSSYVHGAVPWYCPVEVGKKVDPTGDYIKRYVPEVRGLPSEYVCEPWNAPLSVQKACRCVVGEDYPSPIVDHMEQRMICVQRMQQLSIDLGAKGERVTS* 0 >CRY1E_nemVec Nematostella vectensis (anemone) XM_001632800 GCTSVHWFRKDLRLHDNPSLLASLDNCSTFFPIYVLDMESARASKISANRWNFLCECLEALDRQLRVLGSRLFVIRGRAIDVLPRLFHEWSVNRLTFERE SEPAGRQRDTVIQMLAENANVQLLQHNAHLLYDTDEVLETNDGKLPMTFDEMAKTAEQLGPPCPPCQTVDKTVFGACITPVGLDHAEKYGVPQLSDFGGK ELGRATAKLFWKGGELEAMRRLNLALQKKKFIKPPLSADTLLASDRALSPYMRFGCLSPCYILDRLTTEYQRTMGSKPPETLYTNLLWREFFFATASTNP DHHRMMGNPLALQIPWEQHPEALSLWKQGKTGFPWIDAIMRQLREEGWIHHLARQAVGCFLTRGCLWVSWEEGFKAFDELQLDAEWSLNAGNWLWLSCSS YVHGAVPWYCPVEVGKQVDPTGEYIRTFVPELRRLPTKFLHEPWKAPSSVQREAGCIVGKDYPQPMVNHLERRVVCVQKMQNFTQTLALLNG* 0 >CRY64_nemVec Nematostella vectensis (anemone) XP_001636303 ABAV01006592 last exon uncertain 0 MASKKTHQTVHWFRNGLRLHDNPALKEAFETSQTVRPLYVLDPDVLKNGNIGVVRWRFILESLADLDNNLKKLNSR 2 1 LFVVRGRPSEVFPKLFKEWKISKLTFEVDTTEPARKQDAEVLKIANKLGVDIEQRVSHTLYDLDR 2 1 IVNKNGGTAPLTYKKFQSIVSSLGPPAAPLPAIDKKMVAGCNTPTTANHDKIYGVPTLE EIGQEVPEESREVLYPGGETEALERLEVYMKKE 0 0 DWVCKFEKPNTAPNSIEPSTTVLSPYMTYGCLSARLFYTRLAEIYAK 0 0 KKKHSQPPVSLHGQLLWREFFTTAAYKTPNFNKMVGNSLCLQVPWDQNDEYLAAWAE 0 0 ARTGYPYIDAIMTQLRREGWVHHLARHSVACFLTRGDLWISWEEGLK 0 0 VFERLLIDHDWNLNAGNWMWLSASSFFHAYFRVYSPVAFGKKTDPNGDYIR 2 1 KYIPKLSRYPPKYIYEPWNAPLAVQEKAGCVIGRDYPCPIVDHNKVVTRNMSRMKEARSLKYGKTDMGVDKRE 1 2 EASTPSKRKSKTTTDKPAKKTKKITDFLADSSE* 0 >CRY2_ampQue Amphimedon queenslandica (sponge) XM_003386521 MDSYSNGFYEEEERTLTVFNNVDGASEEEQDSFFDSYSASASSSPSSLCSSNWHSERLPVSLHWFRSFSLRLHDNSALMASLLRPQTQFRAVFVLDPWFT ETDKKFGANRLKFLLECLHDLSNQLEALGLKLYVAQGQTTAVLAGLCQEWGVTHLSYQKSQEPRSKVEERTISEMASMMNIEVEEFHNHTLYSPTELIKL NEGKPILTFKDFRTLLSKLKPPAVPISSPSTKQKYNEDNIPMFKVQTFQIPSLASLGCYDVELGTGPWVGGETEALKRLDNYCTVRSKPFDNFLDCLFDK TSLSPYVRFGCLSVRHFWHYVRQLASTDKSKMTLVQEVSSKLLQREFYFLVSSQVPNFDTDTQNPICIQLPWEFDADGLRRWRSGMTGYPWIDAAIRQML KEGWIHNLLRESIATFLTRGDLWISWLKGAESFEMYMLDYEPSVSCGCWMKSSSTAFITGTIEHYNPISYGQQLDPNGDYIRHFVPEVKNLPKEYIFCPW MAPIEVQREAGCIVGTDYPHPVVDHMTAGVICMERLKSFMQSLSSHQTNP* 0 >CRY_ampQue Amphimedon queenslandica (sponge) XM_003386534 METLQGSYDGGGSGFFFPPAGSPRSAMSRHQYPAPEKGHLPSQPPTVLHWFRLDSLRLSDNPAFHHAVSSGKRLKAVVILDPWFNSNNKSGPSANVWRFL LESLHDLDSRLQKRPYNTRLNIYLGQPTVVLSALFHKWNVSELTFQASQTSLESKKHDELIKFAADSQNVKTTSFYSHTLYNPEDILRANNGRFPHSYKE LRRLLPLVGRPREPFPEPDPVLVLLRQNSPEDQDLEEPENRIPSLQDLGFGLEETLYTNSWVGGETEGLSRLSNFCSRRSTQPNEPITWLISKDSLGPYI RFGCLSVRQLFSQLRQYASTSSKGQVLFAELTKNLLMREFAYMVGLTVPKFDTMKDNPLCIQLPWEENERFFYAWKEGRTGYPWIDAAMRQIVRDGWSHY STRQSIAVFLTRGYLWISWEKGLEFFQEHMLDFELPVSTVCWMQSSCSGFFVDFIESYDPCYIGKQMDSDGHFIKLYVPELQDFPSDYIHTPWLAPLHVQ QQANCIIGKDYPKPLVDLCSQGELCCQRVRSILTALRELYGDNP* 0 >CRY_subDom Suberites domuncula (sponge) FN421335 MHCNYPLATQTTFQGQSIPDTTVHWFRLDALRLHDNPAFVDAVKTDGNFKAVFIIDPWFNANYNNGGPQVNVWRFLLEALHDLDSRLQKKPYCARLNVLY GQPTMILPELYKKWNVKKITFQASQVSSESMKHDGIIKILSEQQNVQAVSYFSHTLYDPANVIALNNGRVPLSYKEFRRLMPLMGKPASPIPEPHPMSLC MKAPPSELVPEPEGKIPKLQDLGLSDEFALYTNSWVGGETEALSRLSSFCSRRAAIPNEPVHWLMSKDTLSPYIKFGCLSVRQLFSQLLQFASTSSKGQE LFELLTKNLLLREFAFLVGSSSPKFDVMEGNSLCIQLPWESNNVFIQAFRNGQTGYPWIDAIIRQIRQDGWAHFLARQSIAVFLTRGYLWISWVLGKEFF QEFMIDFELPVSSVCWMQSSCSGFFCTQIESYDPCLVGKQIDTDGHYIKTYVPELKDFPSEYVHQPWKCSSLHTEASWLCDREQYPKPIIDVCKQGELCCKRVQSIMKALADVYGVE* 0 >CRY_aphVas Aphrocallistes vastus (sponge) PubMed:14499587 MESLSITEFSPSAPSPISSSSSPDEETDRDIVAKDIEQDVEPQGKVLLHIFNNRHLRLKDNTALYQAMAQNPDKFYAVYIFDGFDSKPVAPVRWQFLIDC LEDLKEQLNGFGLELYCFRGETIDVLATLVQAWKVKLLSINMDPDVNFTFFNEKIVKMCTINAVQLYNDMDSHRLLYLPPKYKSAIPMSKFRVLLAEAIT AKQNNLESEAKIQDITPPLNPEQLSDLGNKPRLDSPLPSEIPKLNALFTEEEIAKLNFIFQGGERRTEDYLNEYREARLRDVSGDEDASPIAAKAMGISP HLRFGCITPRHLFNFLVKTIKDANYSRIKINKVLAGIMARDFALQVSQLQTIPERIISLNKICLPIPWDKNNNEIVEKLTDAQTGFPFFDAAITQLKTEG YVINEVSEALATFVTNSLLWVSWEEGQNFFSQHLICFDLAMSTHSWLEASGSTMVTGRQKSYQDPLLFVSKKLDPNGEYIKRYLPKFINFPIEFIHKPGN ASLEAQQAANCVIDIDYPKPLFEYECRNGICCKRLRVFMEVVDSAAKATKLPHVIENCSGKFP* 0 >CRY1A_araTha Arabidopsis thaliana (cress) NM_116961 AFNC01018176 aka:CRY1,HY4 PDB:2VTB 0 MSGSVSGCGSGGCS 2 1 IVWFRRDLRVEDNPALAAAVRAGPVIALFVWAPEEEGHYHPGRVSRWWLKNSLAQLDSSLRSLGTCLITKRSTDSVASLLDVVKSTGASQIFFNHLY 1 2 DPLSLVRDHRAKDVLTAQGIAVRSFNADLLYEPWEVTDELGRPFSMFAAFWERCLSMPYDPESPLLPPKKIIS 1 2 GDVSKCVADPLVFEDDSEKGSNALLARAWSPGWSNGDKALTTFINGPLLEYSKNRRKADSATTSFLSPHLHFGEVSVRKVFHLVRIKQVAWANEGNEAGEESVNLFLKSIGLREYS RYISFNHPYSHERPLLGHLKFFPWAVDENYFKAWRQGRTGYPLVDAGMRELWATGWLHDRIRVVVSSFFVKVLQLPWRWGMKYFWDTLLDADLESDALGW QYITGTLPDSREFDRIDNPQFEGYKFDPNGEYVRRWLPELSRLPTDWIHHPWNAPESVLQAAGIELGSNYPLPIVGLDEAKARLHEALSQMWQLEAASRA AIENGSEEGLGDSAEVEEAPIEFPRDITMEETEPTRLNPNRRYEDQMVPSITSSLIRPEEDEESSLNLRNSVGDSRAEVPRNMVNTNQAQQRRAEPASNQ VTAMIPEFNIRIVAESTEDSTAESSSSGRRERSGGIVPEWSPGYSEQFPSEENGIGGGSTTSSYLQNHHEILNWRRLSQTG* 0 >CRY1B_araTha Arabidopsis thaliana (cress) CRY2 PHH1 NM_100320 AFNB01000167 no antennal chromophore 0 MKMDKKTIVWFRRDLRIEDNPALAAAAHEGSVFPVFIWCPEEEGQFYPGRASRWWMKQSLAHLSQSLKALGSDLTLIKTHNTISAILDCIRVTGATKVVFNHLY 1 2 DPVSLVRDHTVKEKLVERGISVQSYNGDLLYEPWEIYCEKGKPFTSFNSYWKKCLDMSIESVMLPPPWRLMPITA 1 2 AAEAIWACSIEELGLENEAEKPSNALLTRAWSPGWSNADKLLNEFIEKQLIDYAKNSKKVVGNSTSLLSPYLHFGEISVRHVFQCARMKQIIWARDKNSEGEESADLFLRGIGLREYSRYI CFNFPFTHEQSLLSHLRFFPWDADVDKFKAWRQGRTGYPLVDAGMRELWATGWMHNRIRVIVSSFAVKFLLLPWKWGMKYFWDTLLDADLECDILGWQYISGSIPDGHELDRLDNPA 0 0 LQGAKYDPEGEYIRQWLPELARLPTEWIHHPWDAPLTVLKASGVELGTNYAKPIVDIDTARELLAKAISRTREAQIMIGAAPDEIVADSFEALGAN TIKEPGLCPSVSSNDQQVPSAVRYNGSKRVKPEEEEERDMKKSRGFDERELFSTAESSSSSSVFFVSQSCSLASEGKNLEGIQDSSDQITTSLGKNGCK* 0 >CRY1C_araTha Arabidopsis thaliana (cress) NM_001035626 AFNC01013058 aka:UVR3,CRY3 PDB:3FY4 0 MQRFCVCSPSSYRLNPITSMATGSGSLIWFRKGLRVHDNPALEYASKGSEFMYPVFVIDPHYMESDPSAFSPGSSRAGVNRIRFLLESLKDLDSSLKKLGSRLLVFKGEPGEVLVRCLQE 0 0 WKVKRLCFEYDTDPYYQALDVKVK 0 0 DYASSTGVEVFSPVSHTLFNPAHIIEK 0 0 NGGKPPLSYQSFLKVAGEPSCAKSELVMSYSSLPPIGDIGNLGISEVPSLEELGYKDDEQ 0 0 ADWTPFRGGESEALKRLTKSISDK 2 1 AWVANFEKPKGDPSAFLKPATTVMSPYLK 0 0 FGCLSSRYFYQCLQNIYKDVKKHTSPPVSLLGQ 0 0 LLWREFFYTTAFGTPNFDKMKGNRICKQ 0 0 IPWNEDHAMLAAWRDGKTGYPWIDAIMVQ 0 0 LLKWGWMHHLARHCVACFLTRGDL 0 0 FIHWEQGRDVFERLLIDSDWAINNGNWMWLSCSSFFYQALSPFCFSF* 0 >CRY_phaTri Phaeodactylum tricornutum (diatom) XM_002180059 PMID:19424294 MAKSEEKKHDVAIHWFRNGLRFHDNPCLLDACQKSESLLPIYVVDPEFPFAQTAGCRAGTIRANFLLESINEVDEKLRKMGSQLVVVLGKSHEVLPEIVATT QAKALFYEQEAAAPVREQDAETIQAIKNRLKRDGKNYECKFEAYATHTLHPMERYLAQCKDHTAPSTYGSFTKIFNKMSVAKEVNEVKEVPSLPNKSVKLLEKSFAEALRMPTLKD LGYAAAADDMKNSGKGGYAFAGGENAAIELLAKNMARSQWVATFEKPKTSPNDATRPSTTALSPYVKHGCISPRRFYHELSKVYSKYNSKETSKPPVSLHGQLMWRDFNYLVGYST PNFDKMIDNPIARQIPWDDDPDLLLAWKMSKTGYPYIDAIMTQLRETGWIHHLARHSVACFLTRGDLWQSWEDGATVFEEYLIDADWSINNFNWQWLSCTAHFYQYFRCYSPIAFG KKTDPNGDYIRKWLPQFKDMPAKYIYEPWEAPIELQKKVGVIVGENYPHPIVDHKLVSKNNMSRMKEAYDAQKNREPMPANESHSSGLNRDTSPKRQRRN* 0 >CRY_thaPse Thalassiosira pseudonana (diatom) XM_002291108 MSAALKSTDQDTVVAMHWFRKGLRVHDNPALLHALAITKDTSGPIYPVYIVDPNCYQLLKCSVLRARFLLECISDLDKSLRERGSRLYVATGDPLEVLPELW KEWGVTHVTHEADETGEPYAVARDEGVRSVAKKNGVQVMEFRSETLRPLGNVPGGYVANVGGAVNSVPSTMSSFQGLLGRIDGGNIPLPLDAPTKEEFPQLSDDDDSNKYLPLEHP SDCEDASLTGIVKGGETLALAQLQSTVTARPDWTASFEKPKTSCTEVSTPSTTVLSPYLSLGCLSPRKAWHAVADANRKASSKVNKTKPPVSLHGQLLWRDFNNLIAHSANAQSPG SWGRMQDNPYCRNVPWSSDPKLLKAWKDGKTGFPWIDACMAQLRTEGWIHHLGRHAVACFLTRGDLWQSWEMGADHFEGELLDADYALNNFNWLWLSCSGFFYQYFRCYSPVAFQK KNDVNGNYIRKWVPELAGLPAKYIYEPWKAPSSVLNAAGIKLGDNYPNPIVDHAFVSKENMSKMSLAYDMHKDGEKQEGSKKAAATSSKNKSSDPAKKKQKQMTLK* 0
The 4Fe-4S photolyases and primases
A small subset of available sequences are shown as these suffice for GenBank probes. PFES stands for photolyase with a 4Fe-4S redox cluster. These are entirely prokaryotic.
>PFES_agrTum Agrobacterium tumefaciens (bacteria) NP_355900 aka: PhrB MSQLVLILGDQLSPSIAALDGVDKKQDTIVLCEVMAEASYVGHHKKKIAFIFSAMRHFAEELRGEGYRVRYTRIDDADNAGSFTGEVKRAIDDLTPSRIC VTEPGEWRVRSEMDGFAGAFGIQVDIRSDRRFLSSHGEFRNWAAGRKSLTMEYFYREMRRKTGLLMNGEQPVGGRWNFDAENRQPARPDLLRPKHPVFAP DKITKEVIDTVERLFPDNFGKLENFGFAVTRTDAERALSAFIDDFLCNFGATQDAMLQDDPNLNHSLLSFYINCGLLDALDVCKAAERAYHEGGAPLNAV EGFIRQIIGWREYMRGIYWLAGPDYVDSNFFENDRSLPVFYWTGKTHMNCMAKVITETIENAYAHHIQRLMITGNFALLAGIDPKAVHRWYLEVYADAYE WVELPNVIGMSQFADGGFLGTKPYAASGNYINRMSDYCDCRYDPKERLGDNACPFNALYWDFLARNREKLKSNHRLAQPYATWARMSEDVRHDLRAKAAAFLRKLD* 0 >PFES_rhoSph Rhodobacter sphaeroides (bacteria) CP000144 PDB|3ZXS PMID:22290493 6,7-dimethyl-8-ribityl-lumazine antenna aka CryPro 4Fe-4S photolyase MRGSHHHHHHGIRMLTRLILVLGDQLSDDLPALRAADPAADLVVMAEVMEEGTYVPHHPQKIALILAAMRKFARRLQERGFRVAYSRLDDPDTGPSIGAE LLRRAAETGAREAVATRPGDWRLIEALEAMPLPVRFLPDDRFLCPADEFARWTEGRKQLRMEWFYREMRRRTGLLMEGDEPAGGKWNFDTENRKPAAPDL LRPRPLRFEPDAEVRAVLDLVEARFPRHFGRLRPFHWATDRAEALRALDHFIRESLPRFGDEQDAMLADDPFLSHALLSSSMNLGLLGPMEVCRRAETEW REGRAPLNAVEGFIRQILGWREYVRGIWTLSGPDYIRSNGLGHSAALPPLYWGKPTRMACLSAAVAQTRDLAYAHHIQRLMVTGNFALLAGVDPAEVHEW YLSVYIDALEWVEAPNTIGMSQFADHGLLGSKPYVSSGAYIDRMSDYCRGCAYAVKDRTGPRACPFNLLYWHFLNRHRARFERNPRMVQMYRTWDRMEET HRARVLTEAEAFLGRLHAGEPV* 0 >PFES_metMah Methanohalophilus mahii (Euryarchaeota) CP001994 4Fe-4S photolyase MRHYAEKLRNRGADITYIKTAELEKSLSRWIKKKGIDELNIAEPANITLKEYLGKLNIDCKIVFVDNKQFIWSIPEFNTWASSRKNLIMEDFYRTGRKNSEI LLEKDGKPSGGKWNLDRENRKLPPKNGFQKKPPQHIKFSPDKITKEIIAEVERSEYPTYGKGKDFNLAVTHEDAQKALDFFIEEKLSNFGPYQDIMLTGDNVLWHSILSPYLNLGL LHPLNVIKKAELAYYQKNLPLNSIEGFIRQILGWREYMHCIYKYTGDKYLKSNWFDHERELPDIYWYPERTSMNCMASVIEEVLNTGYAHHIQRLMILSNFALLAEVNPAKVKNWF HAAFIDAYDWVMQPNVIGMGQFADGGILATKPYISSANYINKMSDYCQNCTYNHNHRTGEDACPFNYLYWAFLHKNNEKLRDIGRMKLILKNLDRINKKELKQIMTHADDFLKSLK* 0 >PFES_natPha Natronomonas pharaonis (Euryarchaeota) CR936257 4Fe-4S photolyase MTVLVLGDCLTEFGPLASDARSTDERVLCIEARAFARRKPYHPHKLTLVFSAMRHFRDRLREAGYTVDYRRVETFAEGLDAHFAAHPEDHIVTVRRTAHGAT DRLQRLVANRGGTVEFVADPRFHCSREEFDAWADGDPPYRHESFYRHMRRETGYLMDGDEPVGGEWNFDDENREFPGPEYVPPEPPQFEPDETTREVREWVDATFGEDGYDDAPYG GAWADPEPFSWPVTREGALQALEAFIEERLPTFGPYQDAMLGDEWAMNHALLSSSLNLGLLSPSEVIEAALAAFEEGSVSIASVEGFLRQVLGWREFVRHAYRRTPGMAAANQLGA AEPLPEFFWTGDTDMACVADAVDGVRTRGYAHHIERLMVLSNFATLYGVEPSRLNEWFHAAFVDAYHWVTTPNVVGMGTFGTDTLSTKPYVASANYIDRMSDHCSGCPYYKTKTTG DGACPFNALYWDFLGRNESQLRSNHRMGLVYSHYDDKSDGEREAIADRAETLRQRARNGTL* 0 >PRIM2_homSap Homo sapiens (human) primase large subunit 4Fe-4S pdb|3L9Q,3Q36 MEFSGRKWRKLRLAGDQRNASYPHCLQFYLQPPSENISLIEFENLAIDRVKLLKSVENLGVSYVKGTEQYQSKLESELRKLKFSYRENLEDEYEPRRRDHISHFILRLAYCQSEELRRWF IQQEMDLLRFRFSILPKDKIQDFLKDSQLQFEAISDEEKTLREQEIVASSPSLSGLKLGFESIYKIPFADALDLFRGRKVYLEDGFAYVPLKDIVAIILNEFRAKLSKALALTARSLPAV QSDERLQPLLNHLSHSYTGQDYSTQGNVGKISLDQIDLLSTKSFPPCMRQLHKALRENHHLRHGGRMQYGLFLKGIGLTLEQALQFWKQEFIKGKMDPDKFDKGYSYNIRHSFGKEGKRT DYTPFSCLKIILSNPPSQGDYHGCPFRHSDPELLKQKLQSYKISPGGISQILDLVKGTHYQVACQKYFEMIHNVDDCGFSLNHPNQFFCESQRILNGGKDIKKEPIQPETPQPKPSVQKT* KDASSALASLNSSLEMDMEGLEDYFSEDS* 0 >PRIM2_sacCer Saccharomyces cerevisiae (yeast) P20457 aka: PRI2_YEAST primase large subunit PDB|3LGB MFRQSKRRIASRKNFSSYDDIVKSELDVGNTNAANQIILSSSSSEEEKKLYARLYESKLSFYDLPPQGEITLEQFEIWAIDRLKILLEIESCLSRNKSIK EIETIIKPQFQKLLPFNTESLEDRKKDYYSHFILRLCFCRSKELREKFVRAETFLFKIRFNMLTSTDQTKFVQSLDLPLLQFISNEEKAELSHQLYQTVS ASLQFQLNLNEEHQRKQYFQQEKFIKLPFENVIELVGNRLVFLKDGYAYLPQFQQLNLLSNEFASKLNQELIKTYQYLPRLNEDDRLLPILNHLSSGYTI ADFNQQKANQFSENVDDEINAQSVWSEEISSNYPLCIKNLMEGLKKNHHLRYYGRQQLSLFLKGIGLSADEALKFWSEAFTRNGNMTMEKFNKEYRYSFR HNYGLEGNRINYKPWDCHTILSKPRPGRGDYHGCPFRDWSHERLSAELRSMKLTQAQIISVLDSCQKGEYTIACTKVFEMTHNSASADLEIGEQTHIAHP NLYFERSRQLQKKQQKLEKEKLFNNGNH* 0
Alignment-ready sequences
The sequence sets above can be aligned as-is but for most purposes -- especially comparing one orthology class to another -- it is preferable to trim off non-conserved variable length regions at the N- and C-termini and also to groom sequences by removing insertions limited to a single species especially in outlier taxa as these create gap columns that distract from the central evolutionary story.
Some assemblies have missing exons, so when the rate of divergence is low, the exon can be infilled (ie predicted) from the closest related species. For example, zebra finch is missing the first exon of CPD but that is available from other songbirds (which are more suitable than chicken or duck). While this loses true variation, it improves products such as logos and overlays on 3D structures.
However it is counterproductive to infill sequences with multiple missing exons. These are included in the sequence compilation primarily to establish the presence/absence of a gene in the given phylogenetic position. For narrowly targeted purposes, their exons can be informative. However the very fact of poor coverage speaks to the quality of sequencing project, meaning observed variation is not necessarily reliable. Thus the primary use is in confirming change (or not) implied by the complete sequences.
Finally, pseudogenes are not appropriate for groomed alignment inclusion because they introduce spurious amino acid variation not informative to structure or function. The issue here is not older pseudogenes riddled with frameshifts and internal stop codons, but rather newer pseudogenes that have not yet accrued grossly inappropriate features.
These newer pseudogenes can be recognized because their pattern of amino acid substitution does not respect the pattern of conservation within the protein -- only in a pseudogene is a residue strictly invariant for a billion years as likely to be substituted as an inconsequential loop residue. Operationally, the 'dot product' of observed substitutions of a subtle pseudogene with the conservation logo will score too high (ie outside the cluster defined by the non-pseudogene sequences). However very recent pseudogenes cannot be detected by any method. These contribute an effect similar to residual sequencing error and individual polymorphisms.
CRY7
The 14 CRY7 sequences become quite conserved after a ragged amino terminal start. The groomed and gapped sequence set from initial to final regions of conservation extends for 774 amino acids, even after removing an internal spacer region. The sequences are in phylogenetic order relative to Xenopus, the only tetrapod to retain the gene.
Following the curated sequences, the first alignment is conventional (full sequences for all species), allowing motifs of interest to be located by web browser text search. The second alignment, made with Multalin2 is a difference alignment set to display an amino acid position (alignment column) only when 91% or more residues in an alignment column agree. The figure 91% is chosen so 50% of the total amino acids are displayed.
CRY7 is highly conserved despite its restricted phylogenetic distribution. The sequences here omit two fragmentary genes from mollusk. The erratically evolving connector domain following the UIM motif has been trimmed out (3 spaces show its location) to allow for easier comparisons to other members of the gene family. Short non-conserved regions have also been removed from the N- and C- termini.
The best options for modeling the spacial locations of the downstream ultra-conserved residues are Drosophila CRY64 or an Arabidopsis CRY1A. Since the percent identities are fairly low, the two fold domains will be recapitulated but the precise orientations of sidechains will remain problematic. For example it will not prove feasible to dock potential antenna molecules to that domain. Nonetheless, comparisons can be made to conserved residues in other members of the gene family, notably tryptophans. The novel upstream domain cannot be modeled at all for lack of template, though the UIM ubiquitin domain is known to be a plain alpha helix.
CRY7_xenTr ultra-conserved amino acid positions in frog: CRY7_xenTro ....G......F.....S.LG...T...F......L..........L......YF...............L.. ..........RR.R.K...----..........PVL.W.RRDLRL.DNPAL...L..G.PVIP.F.W...EE.G...T.A.GGA.KYWLH.AL......L...GS..... CRY7_xenTro ...-------.....L..L...TGA.T....A.YEPWL..RD......L...GV.....HSYCL..P..V.T.GVGLRGIGSVSHF..CC..N.....G..L..P..LP.P..WP....L..L.L..MP.RKDGT..DWA..IR..WDFSE.GA...L..FL.DGV..YEKES.RAD.P.TS..SP CRY7_xenTro YLHFGQ.S.R.....A........KF.RKLAWRDLAYW...LFP..P.E..RP.YK..RWS.D..HL.AWQ.G.TGYPLVDAAMR.LW.TGWM.NY.RHVVASFL.AYLH..W..GYRWFQDTL.DADVAI.AMMWQNGGM.GLDHWNFVMHPVD.A.TCDP.G.YVRKWCPEL..LPD..IHKPW CRY7_xenTro .C..S.LRRAGV..G..YP.RI..DLEERR..SL.DV..VR.......D..SGCD....P..L....LG..............FLLPVITR.EFK.....P...--.NPY..VLKGYVSR.RDE..A......FTAS...E...R.ER.....R..EGLP..........RT..-.D..S..P... >CRY7_xenTr ultra-conserved amino acids in frog for display on CRY64_droMel 1U3C 34% identity QLESGSVQADEFLCLVLSILGSSRTYSQFPAILQSLSRKEPAMYRELMDLHAEYFRKEPADLETLGYETDLEL RPRESKAKHSRRSRKKKKS APSRGLVAMKPVLVWFRRDLRLHDNPALISALEHGVPVIPVFLWCINEETGQNFTLATGGATKYWLHHALLKLNQSLQRFGSHIIFR VAR SCEEELVSLVHETGADTIIINAVYEPWLKERDDLISETLRRHGVELKKHHSYCLYEPDSVSTEGVGLRGIGSVSHFMSCCKRNNSAPIGMPLDAPRCLPAPCNWPESDHLDTLELGKMPHRKDGTLIDWAVTIRESWDFSEDGAYTCLANFLQDGVKHYEKESGRADKPYTSHISP YLHFGQISPRTVLHEAYFTKKNVPKFLRKLAWRDLAYWLLILFPDMPSEPVRPAYKSQRWSSDLNHLRAWQKGLTGYPLVDAAMRELWLTGWMCNYSRHVVASFLVAYLHIHWVHGYRWFQDTLLDADVAINAMMWQNGGMSGLDHWNFVMHPVDSALTCDPYGSYVRKWCPELAGLPDEYIHKPW KCAPSQLRRAGVILGRNYPHRIVLDLEERREQSLKDVVEVRKKHLEYLDEVSGCDMVQIPDQLLALTLGTSGEDEVVRNRTGSFLLPVITRKEFKYKTLQPDTK DNPYNTVLKGYVSRKRDETIAYMNERHFTASTINEGAQRHERIERTNRLMEGLPAPSDAKNKSRRTPK KDPFSIIPPSY
CRY64
The 27 available CRY64 sequences do not need significant trimming at either the N- or C-terminus. The latter is somewhat peculiar in that the penultimate exon is quite ragged at its end (though preserving its phase 12 splice donor) but the last exon is quite conserved in character (many basic residues) at least in vertebrates. Since the alignment from lizard to sponge is very gappy here, the alignment is splice-driven (that is, forced to agree at the start of the final exon. This exon corresponds to the third from the end in CRY1; in other words, CRY64 lacks any specialized extension as befits its straightforward DNA repair role.
CRY64 has a moderate level of conservation overall but certain interior regions are quite invariant, even at the 90% level over the entirety of metazoan evolution (sponge to lizard). That localization of conservation can be seen thematically in the first image below -- it is predominantly concentrated in the catalytic FAD domain. Vertebrate CRY64 are easily modeled using one of eight Drosophila structural determinations; these have 57% identity over 91% of the length (7 IHWFR...KAAYA 488). This template does not cover the terminal basic exon of vertebrate protein. The groomed and gapped sequence set file also contains an alignment colored by conservation and a difference alignment relative to the CRY64 consensus sequence.
Ultra-conserved residues of CRY64 displayed within lizard sequence for purposes of alignment with paralogs: CRY64_anoCar M IHWFR LRLHDNPAL A PIFILD VG NR RFL L DLD L N RLFVVRG E! L W V LT E D EPY RD V V V HTLYD D I N G PLTY P P CRY64_anoCar Y P L EL F GGE EAL RL WV FEKP PN PSTTVLSPY FGCLS R F EV S QL WREFFYT PNF M GN Q WD N E L W E TGYPFIDA M QL EGWIH CRY64_anoCar HLARH VACFLTRGDLWISWEEG VFEE LLD D LNA NW WLS S FF Q FRVYSP FGKKTD G YI KY P L KFP YIYEPW AP Q AGCI G DYP IV H N M AY >CRY64_conSer Ultra-conserved residues of CRY64 for display on CRY64_droMel 3CVV 59% identity MMAHVSIHWFRKGLRLHDNPALLAAMKNSAEIYPIFILDPWFPKNMQVSINRWRFLIESLKDLDESLKKLNSRLFVVRGRPAEVFPELFTKWKVTRLAFEVDTEPYAR-RDAEVVRLAAEHGVQVIQKVSHTLYDTERIIVENSGKAPLTYTRLQTLVASLGPPKQPVPAPKLEDMK DCCTPVKEDHDLEYGTPSYEELGQDPKTAGPHLYPGGETEALARLDLHMKRTSWVCNFKKPETHPNSLTPSTTVLSPYVKFGCLSVRMFWWKLAEVYQGRKHSDPPVSLHGQLLWREFFYTAGAGIPNFDRMENNPVCVQVDWDNNQEYLRAWREGQTGYPFIDAIMTQLRTEGWIH HLARHAVACFLTRGDLWISWEEGQKVFEELLLDADWSLNAANWQWLSASAFFHQFFRVYSPVTFGKKTDKNGEYIKKYLPFLRKFSNDYIYEPWKAPRSLQERAGCIIGQDYPKPIVEHEKVYKRNLERMKAAYARRSPNLVIQAKDKVSQKK-GVNRKRPEAPTKAKVQAKKVRK
DASH
DASH is a relatively isolated sequence class with an uneven phylogenetic distribution that implies multiple gene loss events, both old and new. The 35 sequences provided here range from sponge to amniotes; conservation is moderate relative to other members of its gene family. The groomed and gapped sequence set shows rather stricter conservation of the catalytic FAD domain. The final region of the protein requires manual alignment utilizing the last intron to insure homological register in a very gappy region. The final RGIDFYF motif can then be aligned but some arbitrary gap placement is still necessary even in closely related species, demonstrating the lack of selective pressure outside of the splice junction and motif.
If a residue is displayed only when 21 or more of its 24 column mates agree (87.5% conservation), the figure at left -- which excludes the terminal exon -- shows the uneven distribution of the 276 best conserved residues.
>DASH_conSeq ultra-conserved consensus sequence residues for display on DASH_araTha 2VTB 53% identity MSMSSSRTVICLLRNDLRLHDNEVLHWAQRNADHIVPLYCFDPRHYVGTHHYNFPKTGPHRLRFLLESVKDLRNTLKKKGSNLVVRRGKPEEVVADLIKQLGSVSAVAFHEEVTKEELDVEKALKQVCAQHGVKVHTFWGSTLYHRDDLPFRHISRLPDVYTQFRKAVESQSRVRPT FPMPDQLKPLPPGLEEGSIPTAEDLGQKDPATDPRSAFPCSGGETQALARLKHYFWDTNLVASYKETRNGLIGMDYSTKFSPWLALGCISPRYIYEQIKKYEKERTANQSTYWVIFELLWRDYFRFVALKYGNRIFYLRGLQDKSVPWKKDMKLFDAWKEGRTGVPFVDANMRELAATGFMS NRGRQNVASFLTKDLGLDWRMGAEWFEYLLVDHDVCSNYGNWLYSAGIGNDPRENRKFNMIKQGLDYDGNGDYVRLWVPELQGIKGGDVHTPWTLSSAALSQAGVSLGETYPNPIVTAPEWSRHINKKPSGSGPSPRGRKGPSHTPKQHKDRGIDFYFSRSKLS
CPD
The 33 CPD sequences are unalignably N-terminally (iMet wander) but are otherwise quite orderly in their conservation. The groomed and gapped sequence set from initial to final regions of conservation runs to 473 amino acids. The sequences are in phylogenetic order relative to marsupial.
The first alignment is conventional (full sequences for all species), allowing regions of interest to be located by web browser text search. For example, Oryza sativa (rice) has a determined CPD structure (3UMV) marked up in its GenBank entry for alpha helices and beta sheets. Methanosarcina mazei (euryarchaeota) also has a relevant structure 2XRY that is still 49% identical to frog that is relevant for comparative purposes.
The second alignment, made with Multalin2 is a difference alignment set to display an amino acid position (alignment column) only when 77% or more residues agree. The figure 77% is chosen so 50% of the total amino acids are displayed.
This is an important evolutionary conservation parameter that differs for each ortholog class of cryptochrome and photolyase (though comparisons are inappropriate unless matched for phylogenetic depth).
The graph shows how this parameter changes as stringency required of conservation is varied. Here the X axis is percentage column conservation required and the Y axis is percentage of amino acids conserved of the 437 residues x 33 sequences = 15,642 aa. CPD is falling off quite slowly compared to an average protein, indicating a large core of residues that cannot be changed if the structure and function are to be maintained.
CPD is remarkably well conserved. That can be seen by requiring a conservation level of 96% (which allows 1 sequence of the 33 to vary, allowing some degree of data error). Of the 473 residues, 145 (30.5%) are invariant. These residues are key to mapping conservation onto 3D location and to reliable cross-alignment of CPD to other diverged homologs. These ultra-conserved residues are shown below for a theran mammal:
CPD_monDom ultra-conserved amino acid positions in marsupial opossum: YWM.RD.R..DNWA.......A.....PL...F......L....R...F...GL.............F..L.G....................D..PL................-.....QVDAHN.VP.W..S.K.E.. A.T.R.K........L..FP................-.W.................W...G.......L..F...R.......RN.P.....S.LSP..H.G....QR..L............V....EE...RREL..N FC.Y...YD...G...WA...L..H..D.R...Y....LE...T.D.LWN..Q......GK.HGF.RMYWAKKILEWT..P...L.....LN.....DG.DP.GYVGC.WS..G.HD.GW.ER..FGK.R.MNY.GC.RKF >CPD_conSer ultra-conserved residues for display on CPD_orySat 3UMV 100% identity YWMLRDQRLADNWALLHAAGLAAASASPLAVAFALFPRLLSARRRQLGFLLRGLRRLAADAAARHLPFFLFTGGPAE-IPALVQRLGASTLVADFSPLRPVREALDAVVGRREAPGVAVHQVDAHNVVPVWTASAKMEYS AKTFRGKVSKVMDEYLVEFPELPAVVPWDREQPEGLVDWDALIARVCSEENVPEIDWCEPGEEAAIEALDGFLTKRIKSYETDRNDPTKRALSGLSPYLHFGHISAQRCALEAKKCRHLSPKSVDAFLEELVVRRELADN FCYYQPQYDSLSGAWEWARKTLMDHAADKREHIYTREQLENAKTHDPLWNASQLEMVHHGKMHGFMRMYWAKKILEWTSGPEEALSTAIYLNDKYEIDGRDPSGYVGCMWSICGLHDQGWKERPVFGKIRYMNYAGCKRKF
When the ultra conserved residues defined by alignment of 33 eukaryotic sequences are displayed (yellow, at left) on the crystallographic structure for rice CPD, a rather striking result emerges: the conserved residues predominantly cluster at the interface between the two domains and also about the catalytic FAD site and DNA binding groove. The structure and function of CPD cannot be said to be understood until an explanation is at hand for all observed conserved residues. Some of this conservation is mundane internal packing but other conserved surface residues are likely involved in protein-protein regulatory interactions while a conserved antenna pocket impies an antenna molecule, and yet other conserved aromatic residues are critical to the light sensing reaction or key to DNA binding and conformational change.
Many different displays of conservation on structure are feasible, for example ultra-conserved tryptophans, a small subset of which presumably participate in electron flow. Those of CPD could be compared (homologically aligned) with those of other orthology classes. While snapshots have some value, an interactive 3D resource is ultimately much better. Whether a viewer can be embedded in this wiki for the diverse computer platforms of visitors needs further exploration. If so, then the focus can shift from static images into providing comparative genomics resources that properly initiate the interactive viewer.
Article authorship and data usage policy
I researched this article in its entirety in the winter of 2012, not paying much attention to previous studies which are excellent on reaction mechanisms and regulatory cycles but completely clueless on comparative genomics ten years into that era. Cryptochromes are a moderately difficult topic as vertebrate genes go because the timing of gene duplications largely falls between the cracks of phylogenetic coverage and because 'model' organisms did not retain a representative set of orthologs.
I plan to greatly expand the treatment of 3D structural aspects of comparative genomics during the summer of 2012. My interests are primarily in the long range evolutionary acquisition and divergence of function-enabling structure, starting from primases and 4Fe-4S cluster photolyases and ending with circadian, magneto-sensing and couplings with opsins. However comparative genomics has major applications to rapid hypothesis testing in all aspects of cryptochrome and photolyase research, the main point being the strong coupling between sequence conservation (ie selective pressure) and functional importance. This means conservation never persists without a reason, and conversely that non-conserved features are not important.
Although copyrighted, all the information here is in the public domain and can be used by anyone without additional permissions if properly sourced; however if data, figures or original observations are taken wholesale for a peer-reviewed scientific publication, it might be appropriate (after consultation early on) to include me among secondary co-authors.
Rather than make article edits yourself, please contact me by email with clarifications, corrections or additions to the content so I can make edits while maintaining a consistent approach. For broader disagreements or different interests, a better option is to simply register at the UCSC genomeWiki site and create your own page within the comparative genomics category.
This is just a scientific research article on a vertebrate gene family, not a counseling resource for personal genomics, recommendation for or against melatonin supplementation, or medical advice on insomnia and metabolic disorder -- thanks in advance for not sending inappropriate email. Technical terms from genetics and molecular biology are not explained in the article when keywords have a satisfactory treatment at wikipedia or in undergraduate genetics texts; because of good keywords, the scientific literature is easily searched at PubMed so not duplicated here.
My last dozen published research papers in PNAS, Nature, Science etc can be found here. Watch for 4 additional comparative genomics paper to appear in 2012. I've also written over a thousand pages of comparative genomics for other human genes, authored the original user manual to the UCSC human genome browser and in 1999 an advanced tutorial on metazoan genome annotation still widely available online. I thank the UCSC Genomics Group (Hiram Clawson, Brian Raney, Maximilian Haeussler) for software, manuscript and literature resources, Evim Foundation for logistical support, and the Sperling Foundation for financial support under project grant 2012.GNTCS.006.