Opsin evolution: key critters: Difference between revisions
Tomemerald (talk | contribs) No edit summary |
Tomemerald (talk | contribs) No edit summary |
||
| Line 53: | Line 53: | ||
The synteny circle surviving at this phylogenetic depth will be local (optimistically Pancrustacean). That is, the blue opsin of Daphnia might in synteny with Drosophila (ie establish orthology) but not to Platynereis ciliary opsin much less any vertebrate opsin (eg encephalopsin). This could be remedied to some extent by ancestral gene order reconstruction. The degree to which synteny can contribute to validating orthology relations within opsins is not currently known. | The synteny circle surviving at this phylogenetic depth will be local (optimistically Pancrustacean). That is, the blue opsin of Daphnia might in synteny with Drosophila (ie establish orthology) but not to Platynereis ciliary opsin much less any vertebrate opsin (eg encephalopsin). This could be remedied to some extent by ancestral gene order reconstruction. The degree to which synteny can contribute to validating orthology relations within opsins is not currently known. | ||
* <span style="color: #990099;">Carybdea marsupialis (jellyfish) cnidarian no opsins yet</span> | |||
Cnidarians are the earliest diverging invertebrates with multicellular light-detecting organs, called ocelli. Photodetectors include simple eyespots, pigment cups, complex pigment cups with lenses, and camera-type eyes with a cornea, lens, and retina. These remarkable eyes are located on sensory clubs called rhopalia with four lining the bell of Each houses six eyes: a pair of pit ocelli, a pair of slit ocelli, and two unpaired lens eyes with counterparts to cornea, cellular lens and retina of ciliated photoreceptors. Anatomically, ocelli have bipolar sensory photoreceptor cells interspersed among nonsensory pigment cells with the apical end making the light-receptor and the basal end forming an axon that synapses with second-order neurons to form what amounts to ocular nerves. | |||
The spectral sensitivity of neritic (near-shore) lens eyes of a box jellyfish, Tripedalia cystophora [http://jeb.biologists.org/cgi/content/full/209/19/3758#REF17 recently considered] by MM Coates et al was interpreted as a single vitamin A-1 based opsin with peak sensitivity near 500 nm (blue-green). However nothing was sequenced. | |||
The most striking jellyfish from the perspective of a complex set of eyes is Carybdea marsupialis, as [http://www.biology.appstate.edu/faculty/martinvj.htm reviewed by VJ Martin]. Antibody studies based on vertebrate cone/rod opsins are doubtful because of cross reactivity to generic GPCR proteins; again no opsins have been sequenced yet.. This would make a great genome to study provided the retroposon and base composition are not unwieldy. Nematostella and Hydra, whatever their other genomic merits, sits in the Anthozoa and Hydrozoa respectively, types of cnidarian lacking elaborate visual systems. | |||
Vision has roles in the reproduction and feeding of cubomedusae which can find each other and chase, catch, and eat teleost fish. A patch of Pelagia nocticula 10 square miles in extent and 35 feet deep recently destroyed a salmon farm off Northern Ireland. | |||
* <span style="color: #990099;">Amphimedon (sponge) metazoan no opsins | |||
Sponges lie at the base of multicellular animals. They are not noted for eyes. However demosponge larva do exhibit phototaxis (shadow seeking under coral rubble) but the action spectrum is [http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=11976887 reportedly] a better fit to a flavin or carotenoid chromophore. The genome of Amphimedon queenslandica has been available for years at the Trace Archives but never assembled. The species was formerly called Reniera spp. and it is still carried under that name at [http://www.jgi.doe.gov/sequencing/why/CSP2005/reniera.html JGI Genome]. Consequently tblastn of contigs is not available without do-it-yourself assembly. | |||
The Oakley group reported [http://www.plosone.org/article/info:doi/10.1371/journal.pone.0001054 searching] for sponge opsins but finding only "non-opsin, rhodopsin-class GPCR genes" from Amphimedon. Similarly, no opsins were located in the even earlier diverging placozoan Trichoplax, choanoflagellate Monosiga, and fungal genomes. This fits a picture of photoreceptor opsins first appearing subsequent to sponge in eumetazoa cnidarians. However these were not de novo genes but rather evolved out of the already-rich cauldron of GPCR gene copies. | |||
Some later diverging species such as the model organism C. elegans lost all of their opsin genes, making them useless in Urbilateran ancestor reconstruction. This argues for much more intensive genomic sampling so as to sidestep the widespread problem of gene loss in model organisms. | |||
<tt> -=-=-= coming real soon! -=-=- </tt> | <tt> -=-=-= coming real soon! -=-=- </tt> | ||
| Line 81: | Line 97: | ||
* <span style="color: #990099;">Hydra (hydra) cnidarian opsins</span> | * <span style="color: #990099;">Hydra (hydra) cnidarian opsins</span> | ||
* <span style="color: #990099;">Trichoplax (placozoan), Monosiga (choanoflagellate), and Fungi no opsins</span> | * <span style="color: #990099;">Trichoplax (placozoan), Monosiga (choanoflagellate), and Fungi no opsins</span> | ||
[[Category:Comparative Genomics]] | [[Category:Comparative Genomics]] | ||
Revision as of 21:21, 24 November 2007
See also: Opsin evolution
Imaging eyes developed in deuterostomes rather late relative to protostomes -- after body plan appearance and divergence echinoderms, acornworms, amphioxus, and tunicates but before divergence of lamprey (and the visually degenerated hagfish sister group) with the next node up cartilaginous fish. Consequently visual systems in these species that bracket the origin of eyes are vital to understand, so the biology and opsin availability for these key species are considered in depth below.
However a broader view is needed because rhabdomeric non-imaging opsins of vertebrates have a predominantly protostomal context (where they constitute the primary imaging opsins); conversely cilliary opsins of vertebrates have functionally obscure but definite antecendents in the ecdyzoa and lophotrochozoa wings. The third type of opsin which uses Go gustducin signalling must also have an important evolutionary history. Recall signaling GPCR alpha subunits experienced gene duplication and subfunctionalization in coordination with opsinlike genes -- for that reason opsins classes can be indicated either by localization (apical rhabdome, cilium membrane, retinal ganglial cell), monophyletic clade in gene tree, or type of signalling partner. This works because after a certain degree of co-evolution, an opsin is irrevocably located into a receptor type.
Some species such as drosophila have lost all ciliary opsins [ref] -- clearly they are not essential for a successful visually complex flying insect with 5-color vision and circadian rhythm. Bees, annelids, and mammals retain ciliary opsins so we know this must be the ancestral bilateran state state. This predicts ciliary opsins in cnidaria and indeed one was just found in cnidaria. One sees the importance of complete genomes here (versus transcripts or immunostained sections): absence of ciliary opsin evidence in a genome is truly evidence of ciliary opsin absence.
When the eye is reduced to a single pigment cell backing a single photoreceptor cell, the opsin of that species will be expressed only in one cell of the entire body. In this situation, the opsin may never show up in transcript collections, even with subtraction of common ones.
Vertebrates could never have evolved cilliary opsin vision had the bilateran ancestor possessed the limited opsin repertoire of fruit fly. Thus most pressing question is -- assuming rhabdomeric opsins were thoroughly entrenched in the earliest imaging eyes and photoreception systems -- what kept ciliary opsins around in early bilatera (and even cnidaria) so that they could later be co-opted for ciliary opsin-based vision? We could also ask why vertebrates did not stay on the rhabdomeric track of early deuterostomes but instead underwent this profound switch to the 'untested' ciliary track. It is not at all clear what advantages ciliary offers over rhabdomeric -- ever miss swatting a fly?
Fossil dna does not go back nearly this far back, the nearly non-existent fossil record of soft body parts is unhelpful, and transcripts plus genomes of living species are 450-550 million years removed from the crucial ancestral chain of events. For example, transcript-labelled thin sections of photoreceptor systems in modern amphioxus only speak to the current situation, as does extraction of opsin genes from the new assembly, and speak to the history only by inference. The situation today may be seriously different from the ancestor in terms of both innovations and losses. And that history is not necessarily the most parsimonius (even though we will often assume that).
It's worth expanding on this perpetual source of confusion by emphasizing here too that contemporary tunicates, lancelets, and lamprey are not ancient, ancestral, antiquated, archaic, character-retaining, dead-end, failed experiments, frozen in time, genetically stationary, living fossil, primitive, primordial, relic, or retro species. They're full modern -- the tree of life is right-justified. Indeed their genes, regulatory signalling systems, and enzymes may be more finely honed than mammals because of more rapid evolution attributable to larger effective population sizes, reproductive mode, generation time, and marine selective predatory pressures.
However we can hope that ancestral character traits will still be reflected to some extent in these earlier diverging species and that with enough complete opsin repertoires from taxonomically appropriate species, the ancestral genes and even visual systems can be reconstructed at key nodes on the phylogenetic tree. The story describing the evolution of the human eye then amounts to describing the status at these successive nodes and perhaps interpolating between them. There are definitely limits to knowledge here as extant metazoans provide only 35 nodes between sponge and human -- gaps between nodes may average 30 million years but can seriously exceed that. This is offset by the occasional proposal of new deuterostome branches (Xenoturbella, Convoluta).
The ideal set of genomes needed to study the evolution of the metazoan eye is only partly completed, underway, or even proposed. In some cases, the genome size of clade representatives is so large (eg lungfish at 25x human) the species may never be sequenced, though opsin transcripts could still be obtained. In others, the rate of evolution has been so fast so long that very little information about photoreception at ancestral nodes can remain (eg Oikopleura). Hagfish opsins, which would conveniently break up the crucial lamprey long branch, are not available at GenBank but here the animal has adopted a deep water (dark) habitat, meaning that its cone opsin genomic repertoire will be highly reduced, if not gone entirely, in its markedly degenerated eyes. (Its other opsins could still be informative.)
Model organism choices do not always coincide with genome sequenceability, transcriptome projects, nor (worst of all) with slow-evolving less derived species. Finally, most sequencing speaks to narrow anthropocentric interests, whereas the more broadly conceived sequencing need is greatest farther back (to break up branches). The evolution of the eye needs a rather different portfolio of genomes than a typical disease gene because of the earlier intrinsic timing of the innovative events. In fact, one product of the investigation here is to spell out these needed genomes. Of course one obvious genome choice is the cubomedusan jellyfish Carybdea marsupialis with its 24 eyes of 6 types.
It's worth reviewing genome status and recent experimental literature on key species. While abstracts are readily available at PubMed, access to free full text is unpredictable, so those links are collected when available. It suffice to reference only recent articles because they cite the earlier literature and citation in turn of their paper are collected by Google Scholar (or AbstractsPlus at PubMed). Most opsin sequences in the Opsin evolution reference sequence collection have a PubMed accession as a field their fasta header database; those can simply be compiled to an active link that opens all of them in one PubMed window.
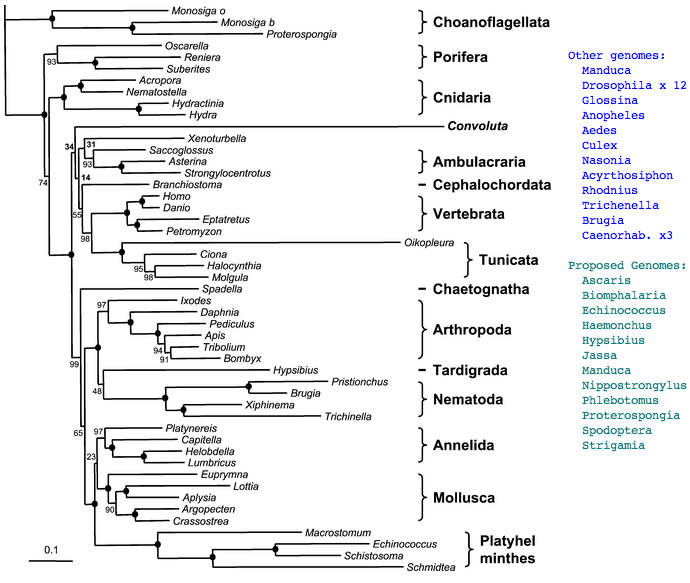
Figure adapted from: Acoel Flatworms Are Not Platyhelminthes: Evidence from Phylogenomics Hervé Philippe, Henner Brinkmann, Pedro Martinez, Marta Riutort, Jaume Baguñà PLoS ONE. 2007 Aug 8
- Callorhinchus (elephantshark) chondrichthyes opsins
Five ray-finned fish genomes are available but these have major lineage-specific expansions and are quite derived. Some sequences are available from lobed-finned fish and coelocanth genome has been proposed. This makes the preliminary genome assembly of the much earlier diverging Callorhinchus (oft-misspelled) and skate transcipts very special because it is the "last stop" before lamprey.
This large-eyed cartilaginous fish lives to depths to 200m on the continental shelf of southern Australia and New Zealand but migrates into coastal estuaries to lay egg cases (lower image) in sand and muddy substrates. The distinctively-shaped egg cases are sometimes found washed ashore after storms. They are up to 25cm long, 10cm wide, and take up to eight months to hatch. The one studied member of the genus has a vitamin A1-based photopigment with maximum absorbance at 499 nm presumably adapted to its overall photic environment.
I made an exhaustive search of the WGS and Trace divisions of GenBank on 5 Nov 2007, recovering many complete exons but most fragmentary genes. The opsin classifier can easily place these fragments. Overall, Callorhinchus appears to have a full complement of vertebrate opsin genes. The exceptions are RHO2, SWS1, SWS2 (oddly also missing in skate and dogfish ESTs) apparently leaving elephantshark with only RHO1 and LWS rod/cone pigments. Parietopsin was also missing. Two encephalopsin- and two melanopsin-class opsins were found. The RGR, peropsin, and neuropsin genes will prove important in better determining their overall gene tree placement (which an October 2007 opsin phylogeny paper placed deeply within rhabdopsins).
- Ixodes (tick)
The genome project was completed long ago but has experienced a multi-year bottleneck in assembly release and publication. However contigs built from 19.4 million traces should become available to tblastn of the GenBank "wgs" division by late 2007. Ixodes has a very conservative genome (regretably 2.1 gbp in size), seemingly far less derived than drosophilids in matters such as intron, gene retention, and protein sequence conservation. This, in conjunction with the helpful phylogenetic position of chelicerate outgroup to the many insect genomes, has improved prospects for reconstructing the ancestal opsin repertoire of Arthropoda and eventually Protostomia and UrBilatera.
A large collection of annotated Ixodes ESTs is available at the DFCI Gene Index of which 3 are marked up (2 wrongly) as opsins. Using the Opsin Classifier, I recovered the full length gene for the first of these TC19272 on 24 Nov 07, intronated the transcript at the Trace Archives (4 introns, superb coverage), and added it to the classifier fasta collection as RHAB1_ixoSca. It classifies with rhabdomeric opsins (ie with deuterostome melanopsins) with a very respectable 57% maximal percent protein identity. The second and third intron have classical ancestral position (following GWSR and LAK) and phase (2 and 0). Synteny awaits assembly of large contigs -- adjacent exons are not spanned by single traces. An apparent ciliary opsin fragment in Ixodes was located using that of Platynereis dumerilii as probe, it is stored as CILI_ixoSca but needs further analysis.
- Daphnia (water flea)
An 8.7x genome assembly was released in July 2007 at JGI with further support at wFleaBase. This crustacean provides a potentially important outgroup to insects (together forming Pancrustacea). However the opsin story, summarized in a meeting abstract is an embarrassment of riches, not conducive to deducing ancestral arthropod genome content. The total number of opsin genes came in at 37, comprised of 22 rhabdomeric opsins (mostly long wavelength), 7 ciliary opsins (pteropsins), and 8 in a novel family without close affiliates. This seems excessive but Daphnia has ommatidia (compound eyes), circadian rhythms, and a need to assess water turbidity and depth. Planned in situ hybridization studies may illuminate biological roles of these opsins. The pteropsins are probably of most interest here.
Gene models have not been submitted yet to GenBank but are likely extractable by text query at wFleaBase. What is needed here however is not the clutter of 37 sequences but their collapse into UV, blue, long, pteropsin, and novel ancestral representatives. This would remove 'noise' from lineage-specific expansions. The intron structure could provide very important support to classification schemes.
The expansions may have arisen through retroprocessing (rather than segmental duplication) of a few master exonic genes, which would then be the orthologs to other arthropod opsins. Indeed the intronation pattern -- typically far more deeply conserved than protein sequence -- could link pteropsins more convincingly to lophotrochoan and deuterostome opsins than alignments with percent identities in the 20's.
This blast twilight zone is especially dangerous for photoreceptor opsins because they are embedded in much larger gene family of generic rhodopsin and GPCR which share many structural and signaling properties. A slowly evolving generic rhodopsin might well score higher than fast evolving photoreceptor opsins. Gene expansions are noted for markedly enhanced rates as copies neo- or subfunctionalize. The generic rhodopsin might also share diagnostic residues through convergence at least at the level of statistical signficance ambiguity. Consequently intron location/phase and synteny can provide important backup.
The synteny circle surviving at this phylogenetic depth will be local (optimistically Pancrustacean). That is, the blue opsin of Daphnia might in synteny with Drosophila (ie establish orthology) but not to Platynereis ciliary opsin much less any vertebrate opsin (eg encephalopsin). This could be remedied to some extent by ancestral gene order reconstruction. The degree to which synteny can contribute to validating orthology relations within opsins is not currently known.
- Carybdea marsupialis (jellyfish) cnidarian no opsins yet
Cnidarians are the earliest diverging invertebrates with multicellular light-detecting organs, called ocelli. Photodetectors include simple eyespots, pigment cups, complex pigment cups with lenses, and camera-type eyes with a cornea, lens, and retina. These remarkable eyes are located on sensory clubs called rhopalia with four lining the bell of Each houses six eyes: a pair of pit ocelli, a pair of slit ocelli, and two unpaired lens eyes with counterparts to cornea, cellular lens and retina of ciliated photoreceptors. Anatomically, ocelli have bipolar sensory photoreceptor cells interspersed among nonsensory pigment cells with the apical end making the light-receptor and the basal end forming an axon that synapses with second-order neurons to form what amounts to ocular nerves.
The spectral sensitivity of neritic (near-shore) lens eyes of a box jellyfish, Tripedalia cystophora recently considered by MM Coates et al was interpreted as a single vitamin A-1 based opsin with peak sensitivity near 500 nm (blue-green). However nothing was sequenced.
The most striking jellyfish from the perspective of a complex set of eyes is Carybdea marsupialis, as reviewed by VJ Martin. Antibody studies based on vertebrate cone/rod opsins are doubtful because of cross reactivity to generic GPCR proteins; again no opsins have been sequenced yet.. This would make a great genome to study provided the retroposon and base composition are not unwieldy. Nematostella and Hydra, whatever their other genomic merits, sits in the Anthozoa and Hydrozoa respectively, types of cnidarian lacking elaborate visual systems.
Vision has roles in the reproduction and feeding of cubomedusae which can find each other and chase, catch, and eat teleost fish. A patch of Pelagia nocticula 10 square miles in extent and 35 feet deep recently destroyed a salmon farm off Northern Ireland.
- Amphimedon (sponge) metazoan no opsins
Sponges lie at the base of multicellular animals. They are not noted for eyes. However demosponge larva do exhibit phototaxis (shadow seeking under coral rubble) but the action spectrum is reportedly a better fit to a flavin or carotenoid chromophore. The genome of Amphimedon queenslandica has been available for years at the Trace Archives but never assembled. The species was formerly called Reniera spp. and it is still carried under that name at JGI Genome. Consequently tblastn of contigs is not available without do-it-yourself assembly.
The Oakley group reported searching for sponge opsins but finding only "non-opsin, rhodopsin-class GPCR genes" from Amphimedon. Similarly, no opsins were located in the even earlier diverging placozoan Trichoplax, choanoflagellate Monosiga, and fungal genomes. This fits a picture of photoreceptor opsins first appearing subsequent to sponge in eumetazoa cnidarians. However these were not de novo genes but rather evolved out of the already-rich cauldron of GPCR gene copies.
Some later diverging species such as the model organism C. elegans lost all of their opsin genes, making them useless in Urbilateran ancestor reconstruction. This argues for much more intensive genomic sampling so as to sidestep the widespread problem of gene loss in model organisms.
-=-=-= coming real soon! -=-=-
- Petromyzon (lamprey) and Eptatretus (hagfish) agnathan opsins
- Ciona (tunicate) urochordate opsins
- Branchiostoma (amphioxus) cephalochordate opsins
- Stronglyocentrotus (sea urchin) echinoderm opsins
- Saccoglossus (acorn worm) hemichordate opsins
- Xenoturbella plus Convoluta no opsins yet
- Schmidtea (planaria) opsins
- Hypsibius (water bear) tardigrade no opsins yet
- Platynereis (polychaete) lophotrochozoan opsins
- Capitella (annelid) lophotrochozoan opsins
- Lottia and Aplysia lophotrochozoan opsins
- Nematostella (anemone) cnidarian opsins
- Hydra (hydra) cnidarian opsins
- Trichoplax (placozoan), Monosiga (choanoflagellate), and Fungi no opsins